Aug 13, 2025
Zika NS2B-3-4A Protein Construct cleavage pattern assay
- Adeeba Dhalech1,2
- 1Stanford University;
- 2ASAP Discovery Consortium
- ASAP Discovery

Protocol Citation: Adeeba Dhalech 2025. Zika NS2B-3-4A Protein Construct cleavage pattern assay. protocols.io https://dx.doi.org/10.17504/protocols.io.yxmvm9dw9l3p/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: March 10, 2025
Last Modified: August 13, 2025
Protocol Integer ID: 124112
Keywords: Polyprotein cleavage, In vitro translation, RRL Extract, NS2B3, NS2B, NS4A, cis cleavage pattern by viral protease, viral protease, cleavage of structural protein vp1, enteroviral 2a protease, vp12a by the enteroviral 2a protease, 4a protein construct cleavage pattern, flavivirus, structural protein vp1, enterovirus, dengue virus, antiviral compound, protein synthesis, helicase domains of ns3, expression plasmid pt7cfe1, cis cleavage pattern, zika, transmembrane domain, protease, cofactor domain of ns2b, 4a cleavage site
Funders Acknowledgements:
National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID)
Grant ID: U19AI171399
Disclaimer
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Abstract
Cis cleavage pattern by viral protease is analyzed using an expression plasmid pT7CFE1-NHA-Chis-target of interest. For flaviviruses (ZIka/DENV) the plasmid has been designed to include the cofactor domain of NS2B, NS2B-3 cleavage site, protease and helicase domains of NS3, and first 49 residues of NS4A to include the NS3-4A cleavage site (1). The plasmid lacks transmembrane domains. For enteroviruses (EVD68/EVA71), the plasmid includes the cleavage of structural protein VP1 from VP12A by the enteroviral 2A protease (2). Protein synthesis is carried out using a Rabbit reticulocyte lysate (TNT‱ Quick Coupled Transcription/Translation Systems; Promega) and 1 µg/50 µL of plasmid template via a coupled transcription/translation system in presence of S-35 labeled L-Methionine (EasyTag; PerkinElmer) (3) and output is quantified using SDS gels.
Zika plasmid used for IVT assays
- D.A. Constant, R. Mateo, C.M. Nagamine, & K. Kirkegaard, Targeting intramolecular proteinase NS2B/3 cleavages for trans-dominant inhibition of dengue virus, Proc. Natl. Acad. Sci. U.S.A. 115 (40) 10136-10141, https://doi.org/10.1073/pnas.1805195115 (2018).
- Doherty JS, Kirkegaard K. 2023. Differential inhibition of intra- and inter-molecular protease cleavages by antiviral compounds. J Virol 97:e00928-23.https://doi.org/10.1128/jvi.00928-23
Guidelines
ASAP Compounds are diluted in DMSO and adjusted to obtain the desired final concentration in the assay.
| Compound | Stock Conc (0.5 uL) | Final Conc (in 10 uL) | |
| ASAP-00XXXXX | 200 nM | 10 nM | |
| 600 nM | 30 nM | ||
| 2 uM | 100 nM | ||
| 6 uM | 300 nM | ||
| 20 uM | 1 uM | ||
| 60 uM | 3 uM | ||
| 200 uM | 10 uM | ||
| 600 uM | 30 uM | ||
| 2 mM | 100 uM |
Materials
TNT T7 Quick Master Mix, [35S]methionine (1,000 Ci/mmol at 10 mCi/ml), Plasmid DNA Template (0.5 μg)/Controls, PCR Enhancer, Nuclease-Free Water, Compound Stock Conc (0.5 µL), Final Conc (in 10 µL)
Troubleshooting
Safety warnings
Always wear appropriate PPE for this protocol
Refer to Material Safety Data Sheets for additional safety and handling information
Assay Prep
Prepare Master Mix for the number of reactions based on Assay Reagents.
Assay Plate Setup
Have 0.5 ml tubes/96 well plates ready for each drug and desired concentration. Add equal amount of DMSO to control tubes.
| Assay Reagent | Standard Reaction (uL) | Scaled Down Reaction (uL) | |
| TNT T7 Quick Master Mix | 25 | 8 | |
| [35S]methionine (1,000Ci/mmol at 10mCi/ml) | 2 | 0.4 | |
| Plasmid DNA Template (0.5μg)/Controls | 2 | 0.4 | |
| PCR Enhancer | 1 | 0.2 | |
| Nuclease-Free Water to a final volume of | 47.5 | 9.5 |
Assay Setup
Aliquot 9.5 µL of the Master mix to each tube and pipet gently to mix.
Incubate the reaction
For enteroviruses (EVD68 and EVA 71) we run the assay for 30 mins at 30 °C.
For flaviviruses (Zika and Dengue), we run it for 60 mins at 30 °C.
Remove from heat and add 10 µL (or equal volume) of 2X Sample buffer to each tube to stop the reaction.
Heat at 65°C for 5 mins.
Run the samples on 12 or 15% SDS Page gel.
30 mins at 70 volts followed by 100 V until the dye front reaches the bottom of the gel.
Remove gels from cassette and transfer to blotting paper.
Cover with Serum wrap and expose the gel overnight on phosphor screen.
In the table, for zika/dengue we use pT7NS2B-3-4A DNA template, and study 3 to 4 cleavage patterns. NS3 S135A is the catalytically inactive protease. For EVD68 and EVA71, we use pT7VP1-2A as the enterovirus template and focus on one cleavage between Vp1 and 2A. C107R is the EVD68 catalytically inactive protease and C110A mutation makes the EVA71 protease catalytically inactive.
Results
Read the exposed gel using a Typhoon Biomolecular Imager (or any machine that reads autoradiography). We use the Typhoon FLA 9500 Biomolecular Imager by GE and read gels at 200 and 50 uM.
Analysis
Gel analysis is carried using LI-COR Studio to optimize visualization of protein bands.
Quantify band intensity using ImageJ. This is the protocol we follow to quantify the bands:
1)Open the file in ImageJ as16-bit
2)Each lane is individually boxed to encompass the entire lane using the rectangle tool.
3)Plot lanes based on rectangular selection using Gel Analyser.
4)Normalize the peak density of each band against background.
5)Record the area of peak intensity of each band using the wand tool.
6)Plot data depending on output on Prism.
Plot data on Graphpad Prism.
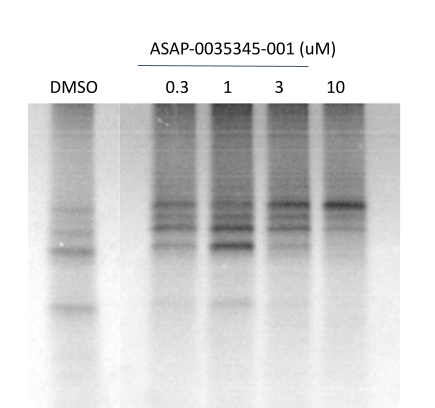
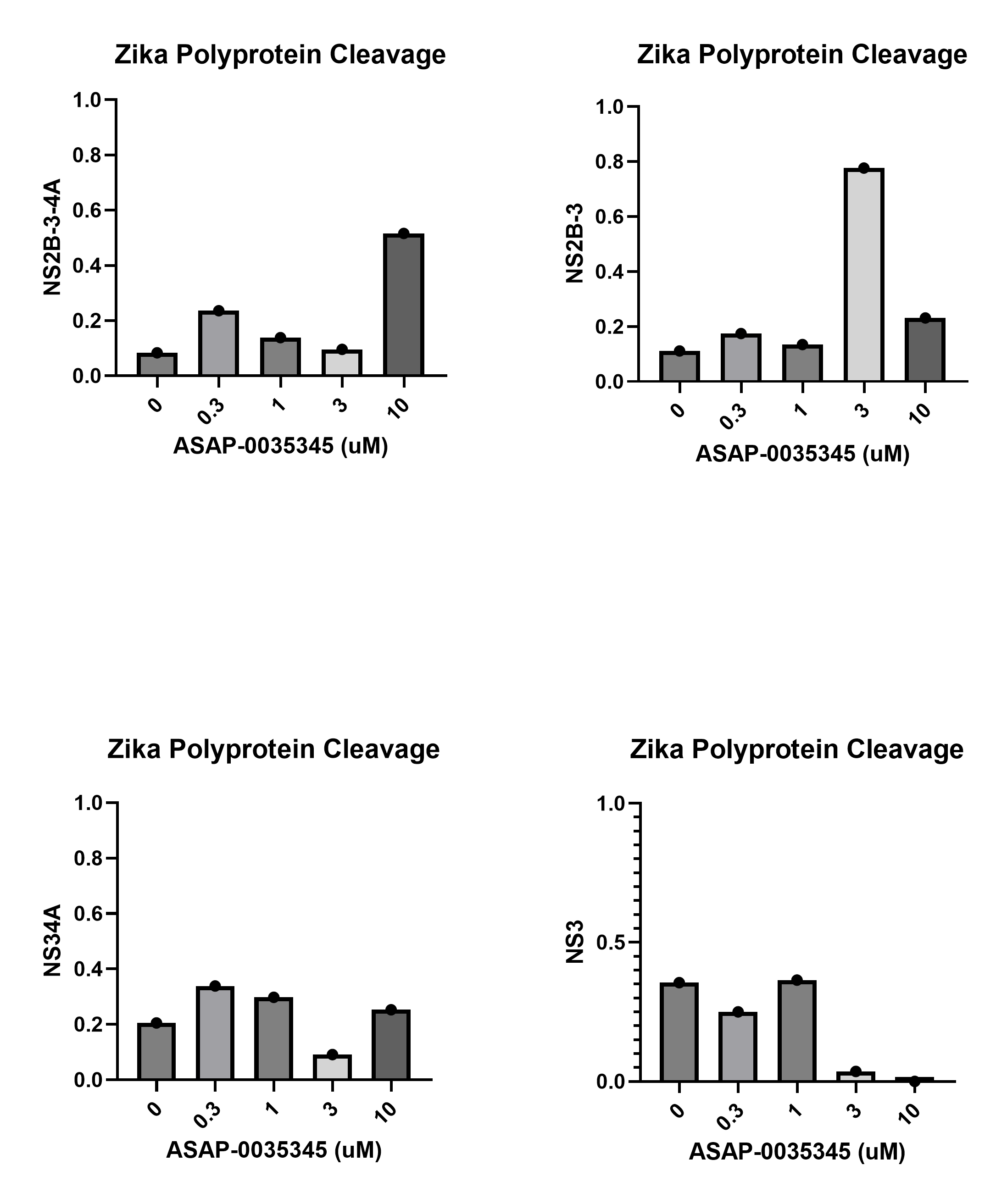
Gel Output
Data from above Gel
Template showing cleavage patterns
Flavivirus (Dengue/Zika) NS3 Protease cleavage and products.
Enterovirus (EVA-71/EVD-68) 2A Protease cleavage and products.
Protocol references
- D.A. Constant, R. Mateo, C.M. Nagamine, & K. Kirkegaard, Targeting intramolecular proteinase NS2B/3 cleavages for trans-dominant inhibition of dengue virus, Proc. Natl. Acad. Sci. U.S.A. 115 (40) 10136-10141, https://doi.org/10.1073/pnas.1805195115 (2018).
- Doherty JS, Kirkegaard K. 2023. Differential inhibition of intra- and inter-molecular protease cleavages by antiviral compounds. J Virol 97:e00928-23.https://doi.org/10.1128/jvi.00928-23