Feb 29, 2024
Wholemount immunolabeling and stereological quantification of autophagosomes in Arabidopsis thaliana root epidermal cells
- Michal Danek1,
- Kateřina Eliášová1,
- Daniela Kocourková1
- 1Institute of Experimental Botany of the Czech Academy of Sciences, Rozvojová 263, 165 00 Prague, Czech Republic

Protocol Citation: Michal Danek, Kateřina Eliášová, Daniela Kocourková 2024. Wholemount immunolabeling and stereological quantification of autophagosomes in Arabidopsis thaliana root epidermal cells. protocols.io https://dx.doi.org/10.17504/protocols.io.bp2l6x5yklqe/v1
Manuscript citation:
Danek M, Kocourkova D, Korec Podmanicka T, Eliasova K, Nesvadbova K, Krupar
P, Martinec J (2024): A novel workflow for unbiased quantification of
autophagosomes in 3D in Arabidopsis thaliana roots, Journal of
Experimental Botany, DOI: 10.1093/jxb/erae084
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: February 07, 2024
Last Modified: February 29, 2024
Protocol Integer ID: 94824
Keywords: autophagy, autophagosome, Cavalieri principle, confocal microscopy, disector, immunolabeling, immunofluorescence, quantification, stereology, stereological quantification of autophagosome, immunolabeling of an autophagosome marker atg8, reproducible quantification of autophagosome, quantification of autophagosome, autophagosome marker atg8, counting of the immunolabeled autophagosome, immunolabeled autophagosome, epidermal cells the workflow, epidermal cell, confocal microscope, immunolabeled sample, arabidopsis thaliana, tissue with the cavalieri principle, stereological quantification, atg8 marker, wholemount immunolabeling, examined tissue, subsequent stereological quantification, cell
Funders Acknowledgements:
Horizon 2020 COST action TRANSAUTOPHAGY
Grant ID: LTC17084
Ministry of Education, Youth and Sports of the Czech Republic
Grant ID: LM2023050 Czech-BioImaging
Disclaimer
DISCLAIMER – FOR INFORMATIONAL PURPOSES ONLY; USE AT YOUR OWN RISK
The protocol content here is for informational purposes only and does not constitute legal, medical, clinical, or safety advice, or otherwise; content added to protocols.io is not peer reviewed and may not have undergone a formal approval of any kind. Information presented in this protocol should not substitute for independent professional judgment, advice, diagnosis, or treatment. Any action you take or refrain from taking using or relying upon the information presented here is strictly at your own risk. You agree that neither the Company nor any of the authors, contributors, administrators, or anyone else associated with protocols.io, can be held responsible for your use of the information contained in or linked to this protocol or any of our Sites/Apps and Services.
Abstract
The workflow enables the quantification of autophagosomes in Arabidopsis thaliana root epidermal cells in 3D. It combines immunolabeling of an autophagosome marker ATG8 with commercially available anti-ATG antibody and the subsequent stereological quantification of the immunolabeled particles. The immunolabeled samples are imaged with a confocal microscope and Z-stacks are acquired. The stereological methods involve counting of the immunolabeled autophagosomes using the optical disector and the determination of volume of the examined tissue with the Cavalieri principle and point counting. The image analysis is performed entirely in Fiji and requires no further special software. The method provides a powerful toolkit for unbiased and reproducible quantification of autophagosomes and offers a convenient alternative to the standard of live imaging with FP-ATG8 markers.
Materials
Consumables:
- Microscope slides and coverslips (e.g. Marienfeld 1000000 and 0101192)
- Parafilm
- Incubation baskets (Intavis 12.440)
- 0.22 µm syringe filter (Merck SLGS033SS)
- 24-well cell culture plates
Equipments:
- Dissecting scissors
- Fine tweezers
- Magnetic stir bars
- Heating magnetic stirrer
- Automated pipetting station (Intavis InSituPro VS)
- Dissecting microscope
- Confocal microscope
Reagents:
- anti-ATG8 primary antibody (Agrisera; AS142769)
- anti-rabbit-DyLight488 secondary antibody (Agrisera; AS09633)
- BSA (Albumin fraction V (from bovine serum); Merck 112018)
- CalcoFluor White (Fluorescent Brightener 28 disodium salt; Sigma 910090)
- DMSO (dimethyl sulfoxide; WWR 85396.290E)
- EGTA (3,12-Bis(carboxymethyl)-6,9-dioxa-3,12-diazatetradecane-1,14-dioic acid; Sigma 03777)
- Glycerol (MP Biomedicals 800688)
- Igepal CA-630 (octylphenoxy poly(ethyleneoxy)ethanol; Sigma I8896)
- MgSO4.7H2O (magnesium sulphate heptahydrate; Merck 105886)
- Pectolyase (pectolyase Y-23 from Aspergillus japonicus; Duchefa Biochemie P8004)
- PFA (paraformaldehyde; Sigma P6148)
- PIPES (2,2′-(piperazine-1,4-diyl)di(ethane-1-sulfonic acid); Sigma P6753)
- Triton X-100 (2-[4-(2,4,4-trimethylpentan-2-yl)phenoxy]ethanol; MP Biomedicals 807426)
- ddH2O or Milli-Q H2O
Troubleshooting
Safety warnings
Autophagy induction can be triggered by relatively small alterations in the conditions under which plants occur. This may contribute to an increase in autophagy already at the beginning of an experiment or under control conditions. Such an increase can then reduce the difference in autophagy intensity between treatments/variants. Avoid exposing plants to additional stress that may occur when samples are collected prior to the fixation step. Plants can induce autophagy when brought into the lab from controlled conditions in a growth chamber and when left on the lab bench for extended periods of time. Try to handle the plant material as quickly and as gently as possible.
Ethics statement
not applicable
Before start
Prepare a sufficient number of slides by washing them thoroughly in distilled water with detergent and then in pure ethanol before use.
Carry out a control treatment and an autophagy-inducing treatment of your choice.
Install the Sampling Window plugin [4] in Fiji.
Microtubule-stabilizing buffer preparation
40m
Microtubule-stabilizing buffer (MTSB) [1] is used throughout the procedure.
Composition: 50 mM PIPES, 5 mM EGTA, 5 mM MgSO4.7H2O, pH = 6.8 (5M KOH).
Note: PIPES will not dissolve in water unless the solution is alkalized.
30m
Add Triton X-100 to 0.1 % final concentration (v/v) to obtain MTSB-T. Mix well using a magnetic stirrer.
10m
Root tissue fixation
2h 35m
PFA [2] is depolymerized in MTSB-T by heating the solution to 75 °C in a covered container in a fume hood to obtain a final concentration of 4 % (w/v). It is let cool down to the laboratory temperature. If the solution volume is reduced due to evaporation during heating, the original volume is topped up with water.
The solution is filtered through 0.22 µm syringe filter to remove traces of undepolymerized PFA.
1h
1 mL of fixation solution is pipetted in wells of the 24-well plate. The incubation baskets are placed in the wells.
10m
Treated seedlings (whole seedlings can be used up to 4 days of age) or detached root tips of approximately 2 cm (in the case of older seedlings) are put in the incubation baskets. Six whole seedlings or 8-10 detached roots can fit into one basket.
Note: The required time is estimated for preparation of two variants, each comprising ca.15 - 20 roots.
15m
Fixation is carried out for 1 hour at the room temperature. To terminate the fixation the baskets are moved to wells of the Intavis specimen holder (equipment of the InSituPro VS station), each containing 600 µL of MTSB-T.
1h 10m
Immunolabeling in the automated station
1d 1h 40m
The specimen holder is mounted in the automated station and the steps 3 – 9 are carried out in the station. The volume of each solution pipetted per one incubation basket is 600 µL. The following steps are set up in the pipetting protocol editor:
20m
Initial rinse and equilibration
- 15 mins with MTSB-T, room temperature; repeat 5 times
- 15 mins with 0.1 % Triton X-100 in water, room temperature; repeat 5 times
2h 30m
Cell wall digestion (+ wash)
- 30 mins with 0.05 % pectolyase in MTSB-T (w/v), 37 °C
- 15 mins with MTSB-T, room temperature; repeat 5 times
1h 45m
Membrane permeabilization (+ wash)
- 30 mins with 10 % DMSO + 3 % Igepal CA-630 in MTSB-T (v/v), room temperature; repeat twice
- 15 mins with MTSB-T, room temperature; repeat 5 times
2h 15m
Blocking
- 1 hour with 2 % BSA in MTSB-T (w/v), room temperature
1h
Primary antibody incubation (+ wash)
- 6 hours with anti-ATG8 diluted 1:1000 in 2 % BSA in MTSB-T (filtered through 0.22 µm syringe filter to remove aggregates), 37 °C
- 15 mins with MTSB-T, room temperature; repeat 8 times
Note: The primary antibody solution can be kept and re-used a few times.
8h
Secondary antibody incubation (+ wash)
- 5 hours with anti-rabbit-DyLight488 diluted 1:1000 in 2 % BSA in MTSB-T (filtered through 0.22 µm syringe filter to remove aggregates), 37 °C
- 15 mins with MTSB-T, room temperature; repeat 5 times
- 15 mins with water, room temperature; repeat 5 times
7h 30m
CalcoFluor White counterstaining (+ final wash)
- 20 mins with 0.002 % CalcoFluor White in water, room temperature
- 15 mins with water, room temperature; repeat 8 times
2h 20m
Specimen preparation
3h 30m
Samples in baskets are removed from the pipetting station and put in wells of a 24-well plate each containing 1 mL 50 % glycerol (v/v).
10m
Strips of one layer of Parafilm are pressed onto microscope slides in order to prevent direct contact between the slide and coverslip and to avoid mechanical damage to the roots.
20m
Immunolabeled roots are carefully removed from the incubation baskets using tweezers and mounted on slides. A dissecting microscope may be necessary to locate the roots in the baskets. 50 % glycerol (v/v) is used as the mounting medium. Sodium azide (0.02 %) can be added to the mounting medium for long-term storage of specimens (several months) in the cold.
Note: The required time is estimated for the preparation of two variants, each comprising ca. 15 - 20 roots.
3h
Microscope observation and Z-stack acquisition
4h
A spinning disk confocal microscope or scanning confocal microscope is used for observation. A 40x objective provides sufficiently detailed view.
Z-stacks are recorded in the meristermatic and/or differentiation zone of the root. The Z-step is set to 500 nm. The first slide of the Z-stack is placed above the surface of the root. Calcofluor White signal is used to determine the correct position. The number of Z-steps within a Z-stack is set to exceed the minimum number of Z-steps required to construct the disector with the desired Z-dimension. This enables the correct placement of the disector upon visual assessment of the surrounding tissue. If the Z-dimension of the disector is 7.5 µm, the minimum number of Z-steps is 21.
Note 1: Other disector dimensions can be chosen. The number of Z-steps is then adjusted accordingly.
Note 2: The required time is estimated for the imaging of two variants, each comprising ca. 15 - 20 roots.
4h
Stereological quantification
10h 30m
The quantification is run in Fiji [3]. Sampling Window plugin [4], enabling the construction of a disector [5], needs to be installed.
Disector construction
(i) A Z-stack is opened in Fiji.
(ii) Disector plugin is launched: Plugins>Disector.
(iii) Disector (x; y; z) dimensions are set to (50; 50; 7.5) µm in the meristematic zone, and to (150; 150; 7.5) µm in the differentiation zone. The X and Y dimensions are set as "Width" and "Height" in the Disector dialog window. The Z dimesnion is set as the number of slides with "First slide" and "Last slide" box.
(iv) Disector position in the X and Y direction within a Z-stack is adjusted with "Horizontal offset" and "Vertical offset" box. Disector position in the Z direction is adjusted with "First slide" and "Last slide" box.
Note: Other disector dimensions can be set. Accordingly, several smaller disectors can be placed within one Z-stack.
1h
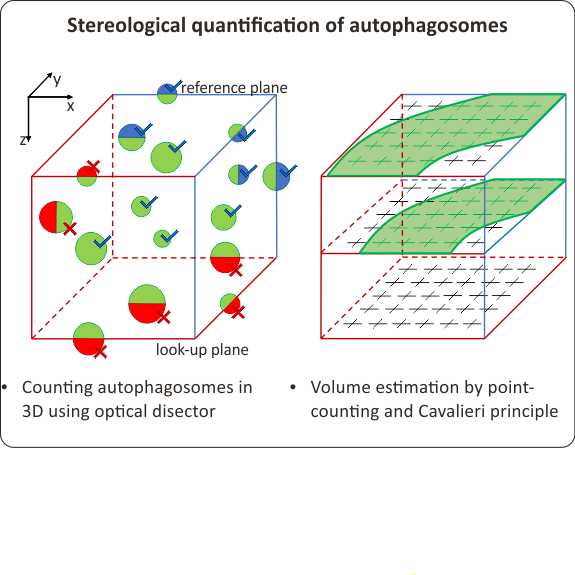
Optical disector principle
An appropriately set disector appears on the first selected slide and remains present until the last selected slide and on all slides in between. Disector projection on each slide has a shape of a square delimited with two exclusion (solid) lines and two inclusion (dashed) lines. Similarly, two planes delimit a disector in the Z direction. The first and the last slide set up in the previous step represent the reference (inclusion) and look-up (exclusion) plane, respectively [6].
Counting autophagosomes using optical disector
A disector delineates the region of interest within a Z-stack in which the autophagosomes are counted. An individual particle appearing anywhere in the disector volume starting from the reference plane (i.e. the first slide) is included in the autophagosome count if both of the following conditions are met:
(i) a particle appears within the disector volume and disapears again before the look-up plane (i.e. the last slide) is reached.
(ii) a particle that lies at the disector boundaries is included if it is intersected by the inclusion (dashed) line.
Particles intesected with the exclusion (solid) lines and particles present at the look-up plane (the last slide) are not included in the number of autophagosomes [6].
The disector volume is observed by scrolling up and down within a given Z-stack. Each particle theat appears within the disector volume is visually tracked and it is counted by clicking on it using e.g. Multi poins selection tool or Cell counter tool of Fiji.
Note: The required time is estimated for the analysis of two variants, each comprising ca. 15 - 20 roots.
6h
Assessment of tissue volume
The volume of the disector is calculated by multiplying its (x; y; z) dimensions. If the entire space of the disector is fully filled with the examined tissue, the disector volume is equal to the tissue volume and the number of autophagosomes is directly related to the disector volume:
E. g., if the disector (x; y; z) dimensions are equal to (50; 50; 7.5) µm in the meristematic zone, the volume of the disector is:
If only a part of the examined tissue lies within the disector, the volume of such an intersection is counted by combination of the Cavalieri principle and the point-counting method.
Assessment of tissue volume by the Cavalieri principle and point counting
The volume of the intersection of the examined tissue and the disector is estimated by counting the number of regularly distributed points in selected Z-slides [6].
(i) After autophagosomes have been counted in step 20, the opened Z-stack with apllied disector must not be closed. Subsequent point-counting procedure is performed on it.
(ii) A 2D grid is generated and superimposed over the opened image using the Grid tool in Fiji: Analyze>Tools>Grid. The parametr "Area per point" is set to 300 µm2 in the dialog box.
(iii) The points of the grid superimposed over the examinated tissue within the disector are counted using e.g. Multi poins selection tool. The points of the grid that do not lie in the tissue are excluded. Points are counted in individual slides within the disector but it is not necessary to use all the slides. Counting points at every third slide (i. e. with 1.5 µm distance between two neighboring slides where the points are counted) should provide a sufficiently accurate estimate of the volume. At least 200 points in total should be counted to get a good estimate. The position of the first slide where the points are counted is randomly selected from the first to the third slide within the disector.
(iv) The estimate of the volume of the examined tissue within the disector is calculated as [6]:
where estV is the estimator of volume in µm3, T is the distance between two neighboring optical sections where the points have been counted, a is the area in µm2 represented by one testing point of the testing grid, and Pj is the number of testing points hitting the j-th (j = 1, 2, …, n) optical section. If the above indicated values are set, then:
Note: The required time is estimated for analysis of two variants, each comprising ca. 15 - 20 roots.
2h
Estimation of the number of autophagosomes per volume of tissue
The number of autophagosomes counted in step 20 is finally related to the volume of the tissue determined in step 21 according to:
where N is the number of autophagosomes and V is the volume of the disector in µm3 . If the disector is not completely filled with the examined tissue, the estimate of volume estV determined by the Cavalieri principle and point counting is used in the equation instead:
30m
Statistical evaluation
To assess putative differences in observed data, appropriate statistical anylysis is performed. The t-test or Wilcoxon test is used to detect differences between two treatments or genotypes, and the ANOVA or Kruskal–Wallis test is applied when more than two variants are compared. 10 - 15 samples are usually sufficient for a statistical analysis.
1h
Protocol references
1. Marhava P, Aliaga Fandino AC, Koh SWH, Jelinkova A, Kolb M, Janacek DP, et al. Plasma Membrane Domain Patterning and Self-Reinforcing Polarity in Arabidopsis. Developmental Cell 2020; 52:223-35 e5.
2. Pasternak T, Tietz O, Rapp K, Begheldo M, Nitschke R, Ruperti B, et al. Protocol: an improved and universal procedure for whole-mount immunolocalization in plants. Plant Methods 2015; 11:50.
3. Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nature Methods 2012; 9:676-82.
5. Sterio DC. The unbiased estimation of number and sizes of arbitrary particles using the disector. Journal of Microscopy 1984; 134:127-36.
6. Kubinova Z, Janacek J, Lhotakova Z, Kubinova L, Albrechtova J. Unbiased estimation of chloroplast number in mesophyll cells: advantage of a genuine three-dimensional approach. Journal of Experimental Botany 2014; 65:609-20.
