Nov 18, 2025
Wheat protoplast preparation and transformation in a 96-well plate
- Eric C Pereira1,
- Danish Ilyas Baig1,
- Benjamin Schwessinger1
- 1Australian National University

Protocol Citation: Eric C Pereira, Danish Ilyas Baig, Benjamin Schwessinger 2025. Wheat protoplast preparation and transformation in a 96-well plate. protocols.io https://dx.doi.org/10.17504/protocols.io.5qpvod1ndg4o/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: October 28, 2025
Last Modified: November 18, 2025
Protocol Integer ID: 231016
Keywords: wheat protoplast preparation, wheat leaf protoplast, cell normalisation, well plate this protocol detail, peg exposure
Abstract
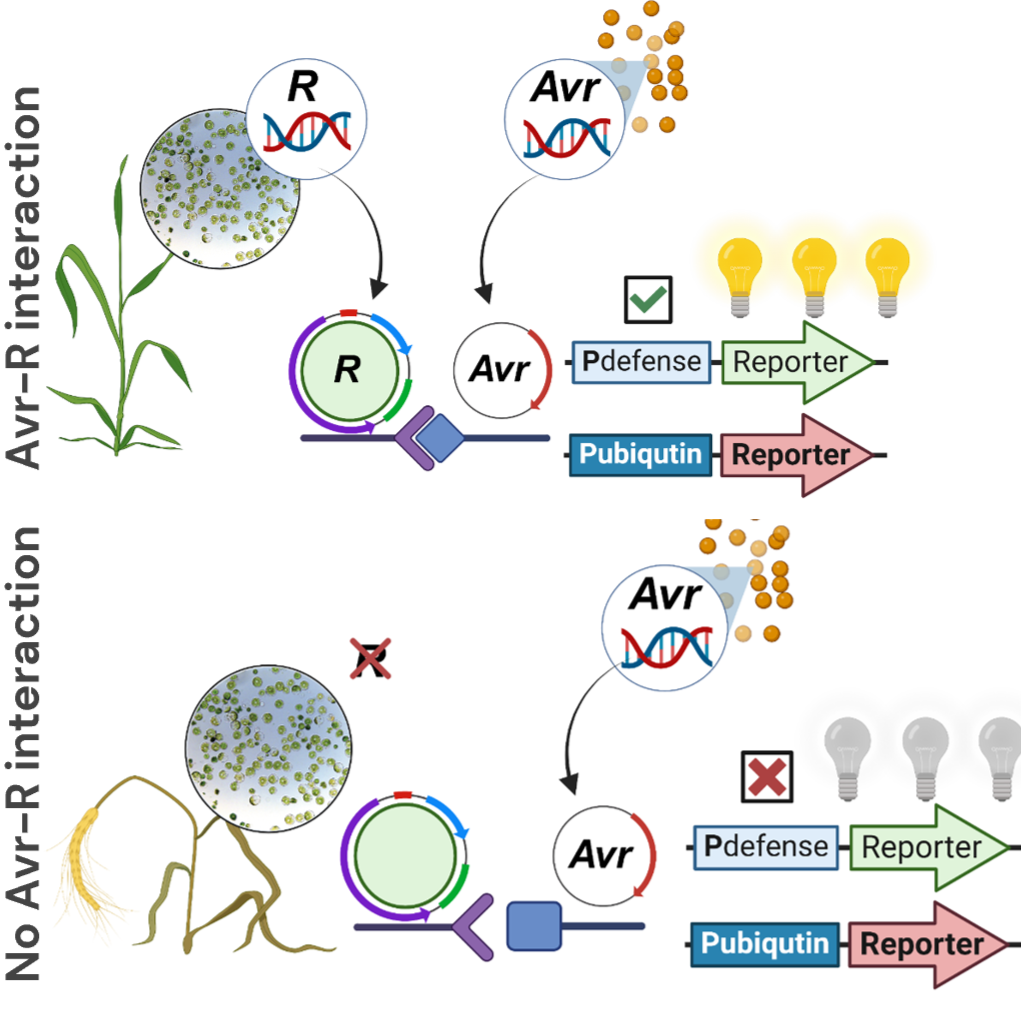
This protocol details a robust, 96-well plate–based workflow for the isolation and PEG–Ca²⁺ transfection of wheat leaf protoplasts, enabling high-throughput screening. The miniaturised format standardises cell normalisation, PEG exposure, and plate handling, and is compatible with luciferase reporters for ~16 h readouts
Materials
0.45 μm filter sterilise and store at RT:
- MES-KOH 0.2 M (Sigma M8250) adjusted to pH 5.7 with KOH
- Mannitol, 0.8 M (Sigma M8250)
- Mannitol 0.6 M (Sigma M8250)
- MgCl2, 2 M (Sigma M2670)
- CaCl2, 2 M (Sigma C7902)
- KCI, 2 M (Sigma P9541)
- NaCl, 2 M (Univar 465)
Store at 4ºC:
- Cellulase 'Onozuka' R-10 (Yakult)
- Macerozyme R - 10 (Yakult)
- Purified plasmid DNA which will express reporte
Store at RT:
- PEG 4000 (Sigma 81240)
- Wheat seeds, potting mix and fertiliser
Filter sterilise aliquots of the required quantity and store at -20ºC
- BSA 10% w/v (Calbiochem 126609)
Equipment:
- Syringe sterilisation filters 0.45 μm
- Cell strainer 100 μm
- 30 mL round-bottom tubes (Sarstedt 55.517)
- Hemocytometer
- V-bottom 96-well plates
- 96-well white opaque plates (for luminescence readings)
W5
| A | B | C | |
| Volume | Final concentration | ||
| MES 0.2M | 0.5 mL | 2 mM | |
| KCl 2M | 0.125 mL | 5 Mm | |
| CaCl2 2M | 3.125 mL | 125 mM | |
| NaCl 2M | 3.850 mL | 154 mM | |
| dH20 | To 50 mL |
MMG
| A | B | C | |
| Volume | Final concentration | ||
| MES 0.2M | 0.8 mL | 4 mM | |
| Mannitol 0.8M | 20 mL | 0.4 Mm | |
| MgCl 2M | 0.3 mL | 15 mM | |
| dH20 | To 50 mL |
Troubleshooting
Plant Growth
- Label. Label each pot with wheat cultivar/line and sowing date.
- Fill pots. ¾ Fill a 10cm diameter pot with soil (Martins mix / seed raising mix)
- Sow. Sprinkle 10 to 15 seeds on top, then fill the pots to just below the rim.
- Place in trays. Place pots in the shallow black tray and dampen the soil with water mixed with complete soluble fertiliser (Hortico brand, complete mix – 1 scoop/1L).
- Environment. Move trays to the growth space (16day/8night hrs; 22°C; 200umol light intensity).
- Plants are ready to use after 7-9 days (2nd leaf just emerging). Some wheat cultivars may emerge more slowly, so adjust accordingly (e.g., Avocet lines).
Enzyme Solution
Prepare fresh on the day of use
| A | B | C | D | |
| Volume | Final concentration | Directions | ||
| MES 0.2M pH 5.7 | 0.5 mL | 2 mM | ||
| Mannitol 0.8M | 3.125 mL | 125 Mm | ||
| KCl 2M | 0.125 mL | 5 mM | ||
| 55 °C for 5 min | ||||
| Cellulase ‘Onozuka’ R-10 | 75mg | 1% w/v | ||
| Macerozyme R-10 | 37.5mg | 0.75% w/v | ||
| 55 °C for 10 min | ||||
| Cool to RT | ||||
| CaCl 2M | 25 ul | 10 Mm | ||
| BSA (10% W/V) | 50 uL | 0.1% | ||
| dH20 | upto 5 mL | mix gently |
Protoplast isolation
A - Tissue prep & enzymatic release
- Osmotic pre-equilibration. Add 5mL of 0.6 M Mannitol to the small petri plate. - Small leaves + mannitol pre-plasmolysis enriches the mesophyll and minimises bulk tissue.
- Leaf selection & peel. From 3–4 small leaves (randomly chosen from different seedlings), make a shallow longitudinal nick in the abaxial or adaxial surface with a blunt razor. Use tweezers to gently peel a thin epidermal strip (~3–5 mm wide). - Aim for multiple short peels rather than one long sheet.
- Short mannitol soak. Place peels, peeled-side down, into the mannitol for ≤10 min to equilibrate (do not exceed 10 min).
- Enzymatic digestion. Add 5 mL freshly made enzyme solution to a clean petri plate. Transfer the mannitol peels to the enzyme solution using tweezers. Wrap the plate in foil (dark) and incubate at RT with gentle orbital shaking for 3 h. - Leaf pieces become glassy/transparent; the suspension turns slightly opalescent with free protoplasts.
Tissue preparation
B - Recovery & washing
- Leaf removal & final release. After 3 h, remove the peels with tweezers, then lift and agitate in the enzyme solution to release trapped protoplasts; discard the peels.
- Pre-wet filter. Place a cell strainer (100um) over a small Petri dish and wet it with 1 mL W5.
- Filter. Pour the enzyme suspension (now with protoplasts) through the strainer.
- Rinse well. Rinse the original petri plate with 1 mL W5 to collect residual protoplasts, then pass the mixture through the strainer.
- Collect. Transfer the filtrate to a 14 mL round-bottom tube and bring the volume to ~10 mL with W5.
- Pellet. Centrifuge at 100 × g for 3 min (RT).
- Resuspend. Carefully remove supernatant without disturbing the loose pellet.
- Wash/stabilise. Add 10 mL W5, gently resuspend.
- Incubation. Place in cold (preferably 4 °C) in the dark for up to 40 min, or until protoplasts have settled.
Note: Use cut tips or wide bored tips for handling fragile protoplasts
C - Counting
- Count. Load 10 µL of protoplast suspension onto a haemocytometer. Count cells in 4 big squares without grids (exclude debris; count only intact, round protoplasts).
- Calculate. total number of cells = (average cells of 4 squares counted) x 10,000 x initial Volume (10 mL) / 350,000
- Adjust. Remove supernatant after chilling (if settled), then resuspend in MMG to 3.5 × 10⁵ protoplasts mL⁻¹.
Protoplast suspension onto a haemocytomete
Protoplasts are now ready for the transformation step (they should be used ASAP)
PEG preparation - during protoplast incubation
- Prepare fresh PEG-Ca solution (shake vigorously to dissolve PEG). Don't reuse
| A | B | C | |
| Volume | Final concentration | ||
| PEG 4000 | 1200 mg | 40 % w/v | |
| Mannitol 0.8M | 750 ul | 5 Mm | |
| CaCl2 (2M) | 150 uL | 100 mM | |
| dH20 | to 3 mL |
Plasmid DNA preparation for transfection
- Prepare setup. Prepare a table containing information about the transfections, including date, cultivar, plasmids, and repetitions.
- Unify lots if needed. If a construct is present in multiple preps, combine aliquots into a single master lot to avoid batch-to-batch variation. Mix by gentle inversion.
- Dispense into the plate. Using fresh barrier tips for every well, dispense the required volume of DNA into the bottom of each assigned V-bottom well per the setup sheet.
- Storage (if not used immediately). Short term: 4 °C up to 24 h.
Transfection
- Verify PEG. Confirm the PEG solution is fully dissolved, clear, and at RT.
- Pre-load PEG. Pre-load a pipette tip with 50 µL PEG for immediate addition after cells.
- Re-suspend cells. Gently swirl the protoplast tube to homogenise (they settle fast). Mix just before every dispense.
- Add cells to DNA. Dispense 50 µL protoplast suspension directly into the DNA in the assigned V-bottom well. Touch off at the bottom; avoid splashing.
- Immediate PEG addition. Add the pre-loaded 50 µL PEG to the same well. Gently pipette up–down 5× to homogenise.
- Incubate. Repeat for the remaining wells, working in a consistent order. Incubate 15 min at RT.
- Stop reaction. Add 200 µL of W5 slowly down the wall of each well to stop the reaction. Optional: mix gently by pipetting once.
- Incubation. Incubate ~16 h in light at RT (≈22–24 °C) on a level surface.
Luminescence measurement
- Remove supernatant. During the overnight incubation, the protoplasts settled at the bottom. Carefully aspirate the supernatant from each well without touching the pellet (took out 125ul twice)
- Lyse. Add 50 µL of 1× lysis buffer to each well. Gently pipette up and down 5× to mix (avoid bubbles).
- Incubate. 15 min at RT to complete lysis.
- Prepare substrate. During lysis, load a PCR strip with Steady-Glo substrate (luciferin) for multichannel dispensing.
- Transfer lysate. Transfer 50 µL lysate from each well into a white, opaque flat-bottom 96-well plate.
Go to the plate reader for the final addition of substrate
- Read. Measure the background luminescence with the plate reader.
- Add substrate. In the plate reader, add 50 µL Steady-Glo to each well with a multichannel pipette, then pipette up and down 2× gently to mix.
- Read. Measure luminescence with the plate reader (Tecan, Mannedorf, Switzerland; Infinite 200Pro, 1000 ms integration, 0 ms settle): Luminescence (no filter) and Integration time: 1000 ms (0-ms settle; auto-gain or fixed as validated). Get individual luminescence using the installed Lumi Green band-pass and Red NB long-pass filters, with the same acquisition settings.
Data processing
Perform deconvolution calculations using measurements from red and green filters, using spreadsheet template.
Protocol references
Wilson, S., Dagvadorj, B., Tam, R., Schwessinger, B. (n.d.). Wheat protoplast preparation and transformation
(Version 1). protocols.io. https://dx.doi.org/10.17504/protocols.io.q26g7r3zkvwz/v1
Wilson, S., Dagvadorj, B., Tam, R., Murphy, L., Schulz-Kroenert, S., Heng, N., Crean, E., Greenwood, J., Rathjen, J.P., Schwessinger, B. 2024. Multiplexed effector screening for recognition by endogenous resistance genes using positive defense reporters in wheat protoplasts. New Phytol, 241: 2621-2636. https://doi.org/10.1111/nph.19555
Khan, A., Lloyd, J.P.B., Kidd, B., Lister, R. 2022. Protoplast isolation and transfection in a 96-well plate. protocols.io. http://dx.doi.org/10.17504/protocols.io.yxmvmn1d6g3p/v1
