Nov 13, 2025
Updated Eastern Hemlock Tissue Collection for DNA
- Karl Fetter1
- 1University of Connecticut, Department of Ecology and Evolutionary Biology
- PlantCompGenomics

Protocol Citation: Karl Fetter 2025. Updated Eastern Hemlock Tissue Collection for DNA. protocols.io https://dx.doi.org/10.17504/protocols.io.dm6gpmkrdgzp/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: November 12, 2025
Last Modified: November 13, 2025
Protocol Integer ID: 232185
Keywords: eastern hemlock tissue collection for dna step, eastern hemlock tissue collection, tissue from eastern hemlock, plant computational genomics lab, landscape genomics study of climate adaptation, landscape genomics study, metadata in treesnap, eastern hemlock, treesnap, dna analysis, dna step, tissue sample, dna, climate adaptation, updated eastern hemlock tissue collection
Funders Acknowledgements:
USDA - NIFA
Grant ID: 1030624
Disclaimer
DISCLAIMER – FOR INFORMATIONAL PURPOSES ONLY; USE AT YOUR OWN RISK
The protocol content here is for informational purposes only and does not constitute legal, medical, clinical, or safety advice, or otherwise; content added to protocols.io is not peer reviewed and may not have undergone a formal approval of any kind. Information presented in this protocol should not substitute for independent professional judgment, advice, diagnosis, or treatment. Any action you take or refrain from taking using or relying upon the information presented here is strictly at your own risk. You agree that neither the Company nor any of the authors, contributors, administrators, or anyone else associated with protocols.io, can be held responsible for your use of the information contained in or linked to this protocol or any of our Sites/Apps and Services.
Abstract
Steps for collecting tissue from Eastern Hemlocks for DNA analysis. The Plant Computational Genomics lab is conducting a landscape genomics study of climate adaptation in the Eastern Hemlock. This protocol is a step-by-step guide to collect a tissue sample. The protocol records metadata in TreeSnap (https://treesnap.org) and is designed to allow revisiting a collection site to collect more data by future researchers.
Troubleshooting
Introduction
This protocol is intended for field collection of leaf tissue for the Eastern Hemlock conservation genomics project sponsored by the Plant Computation Genomics lab at the University of Connecticut. The goal of the project is to identify climate adapted genomic variation for seed banking and potential use in future breeding programs or for assisted migration.
A complete data collection will include the sample, metadata of the tree, a photograph, and a tagged tree. In some managed lands, tagging the tree is not possible and this may be considered optional. However if possible, we ask collectors to tag their trees to allow new data collection from the tree, increasing the value of the DNA sequence data.
Mary Rutter
C/O Rachel O'Neill
University of Connecticut
67 North Eagleville Rd
U-3197, ESB Room 209
Storrs, CT 06269
Collection Materials
You will be provided a collection kit with
- plastic bag (Ziploc) with a barcode
**- paper towels
- silica gel
**Please moisten the paper towels with water.
**- This protocol was recommended for smaller trips that was conducted by the PCG lab at UConn.
If you have silica gel in your collection kit, this is to help reduce the dependency in the field for water to moisten paper towel.
Please pour a bit of the loose silica dessicant into the ziploc bag.
Instead of using moist paper towels you will be placing hemlock cuttings into the ziploc bag with silica gel.
Cutting shears or knife, a smart phone, hammer, and GPS are required, but not supplied.
Download and install TreeSnap onto your smartphone. TreeSnap is a light weight app for collecting metadata for tree studies.
Going into the field
A few things to remember when you are ready to collect.
1. Always collect with permission. For managed lands, a permit or permission from the land manager is required. For collecting on private lands, always ask the land owner before collecting. Most people love hemlocks and will give permission to make a small collection.
2. Bring along your sampling kit
3. Bring a buddy and have fun collecting!
Finding a site
Road side botany is great fun, but be sure you aren't collecting a planted tree. Trees of wild provenance will only be used in downstream analyses.
Choose a site based on these characteristics:
- You have permission to collect
- Sites must be at least 1 km (0.62 miles) distant from each other
- Site is accessible by car or a short walk/hike
- The site will make a good TreeSnap entry
Please note that it is preferable to collect two samples of the same tree OR a neighboring tree. This is to increase the chances of a successful DNA extraction.
Good photographs for TreeSnap will include landmarks like telephone poles, signs, or other features that aid revisiting the site. When you can't tag a tree, be sure there are good landmarks in the photo, or point to the tree in the photo!
Identifying a tree to collect
- Eastern Hemlock trees should be adults or established juveniles (>4" DBH).
- Regenerating trees should be sampled only if no adult trees can be found.
- Trees should ideally be accessible for tissue collection by hand or with a pole pruner near a trail head or road.
- Collect trees of wild provenance. In an rural or urban setting, iIt's impossible to know if a tree near a house was planted. Avoid collecting in someone's yard.
There is no minimum sample size for a collector, but 10 samples from a collector is appreciated.
Collecting tissue
Collect approximately 1 to 2 grams of leaf tissue for each sample. 1 gram is about 5 segments of terminal branches (see images below). Needles will be extracted and very woody samples should be avoided if possible. Freshly expanded green leaves or buds are great to include.
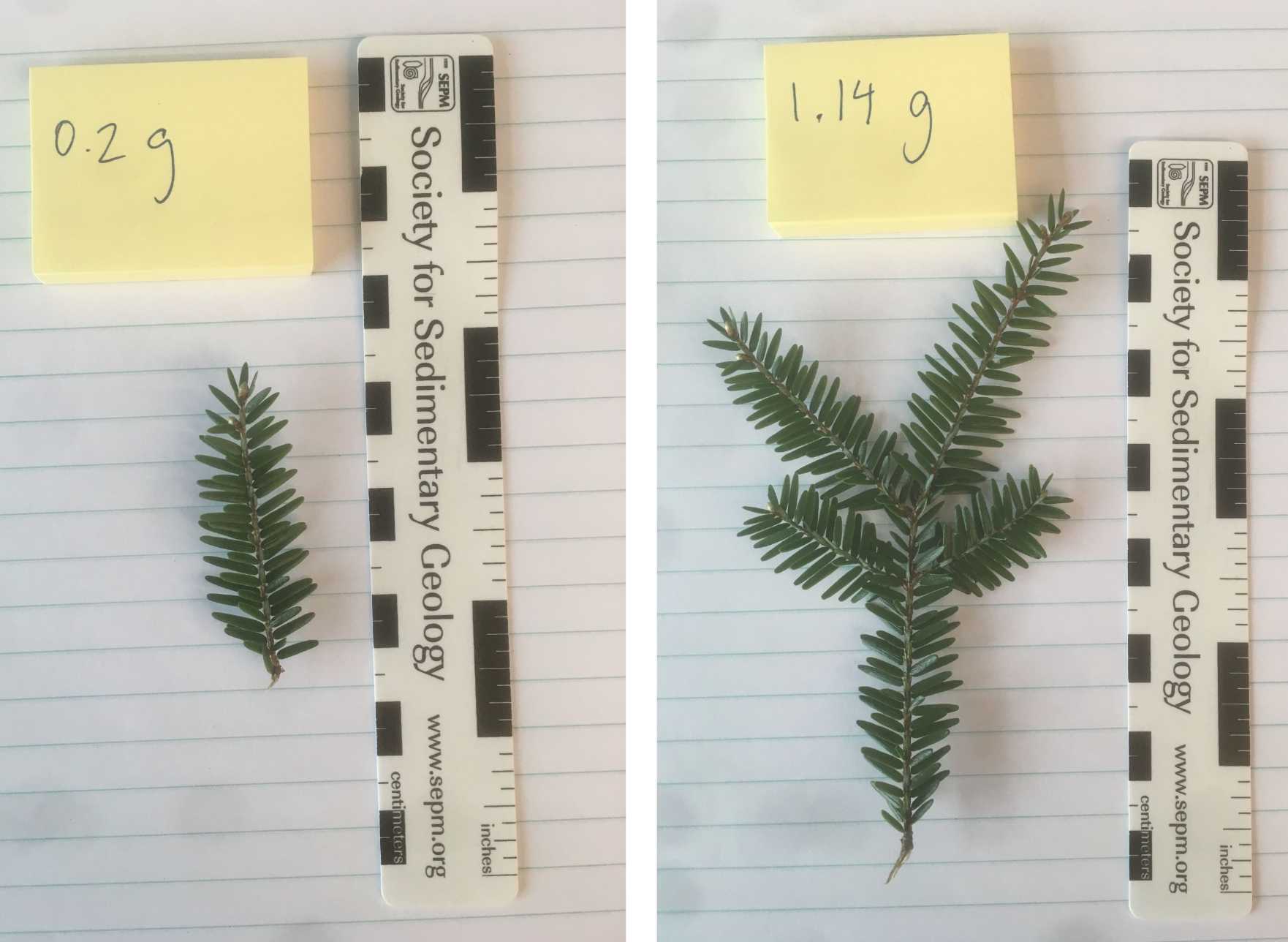

Make the collection by cutting the stem with a knife, shears, or snapping the branch.
- Place the sample into a moist paper towel
- Deposit the collection into the ziploc.
- Label the envelope with "your intials"-"MonthDateYear"-"CollectionNumber". For example, my first collection (July 19th, 2023) was KF-071923-01.
- Write the latitude and longitude of the tree on the ziploc.
- Collect metadata through TreeSnap

Example of text that would be written on the Ziploc bag
It is preferred to write the sample ID and the Lat.-Longs on the envelope. If TreeSnap fails, the written data is the backup.
Metadata
Metadata should be collected which allows 1) revisiting the tree, 2) general health characteristics, and 3) a general description of the site. A photograph should be taken of each tree. Use TreeSnap to collect these data. LeafSnap can be downloaded from the App Store or Google Play Store. After installing the app and registering, click on the Hemlock button. If you are without cell service in the field, when you connect to service again, open the app and click the notification to upload data.
Open TreeSnap and click on the hemlock button.
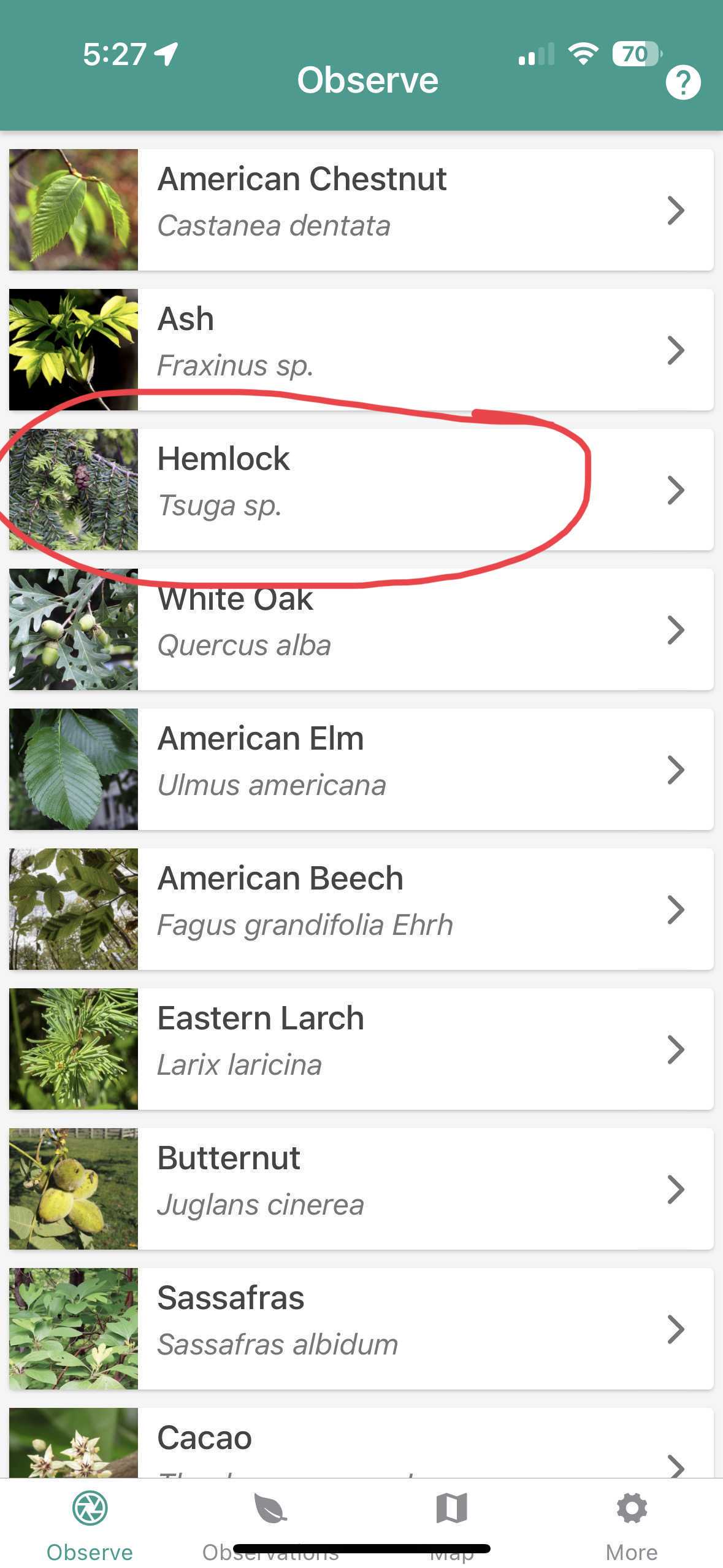
Landing page of TreeSnap.
Before adding pictures of the trees and entering metadata, choose the appropriate collection purpose:
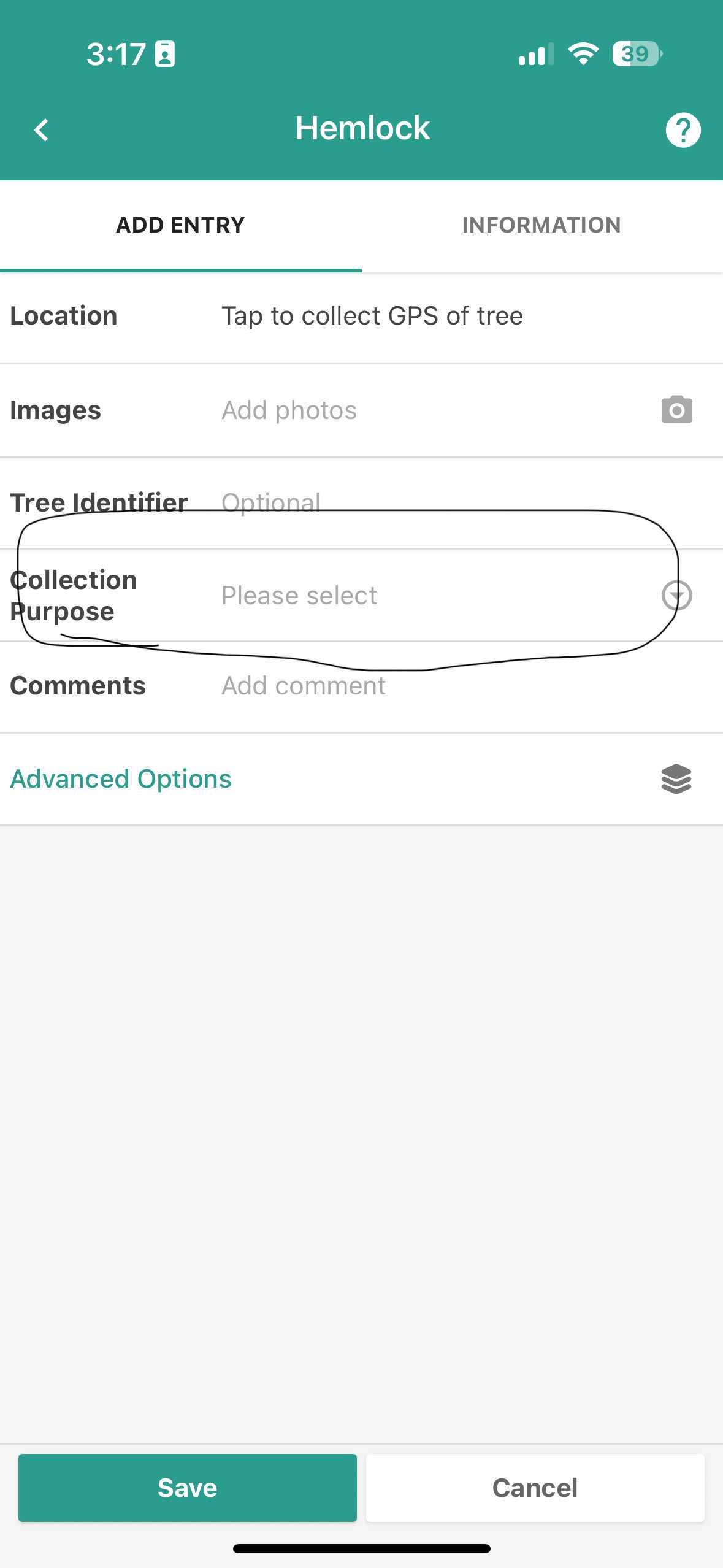
Screenshot of TreeSnap page after selecting species
Please select 'Landscape Genomics project with the University of Connecticut'
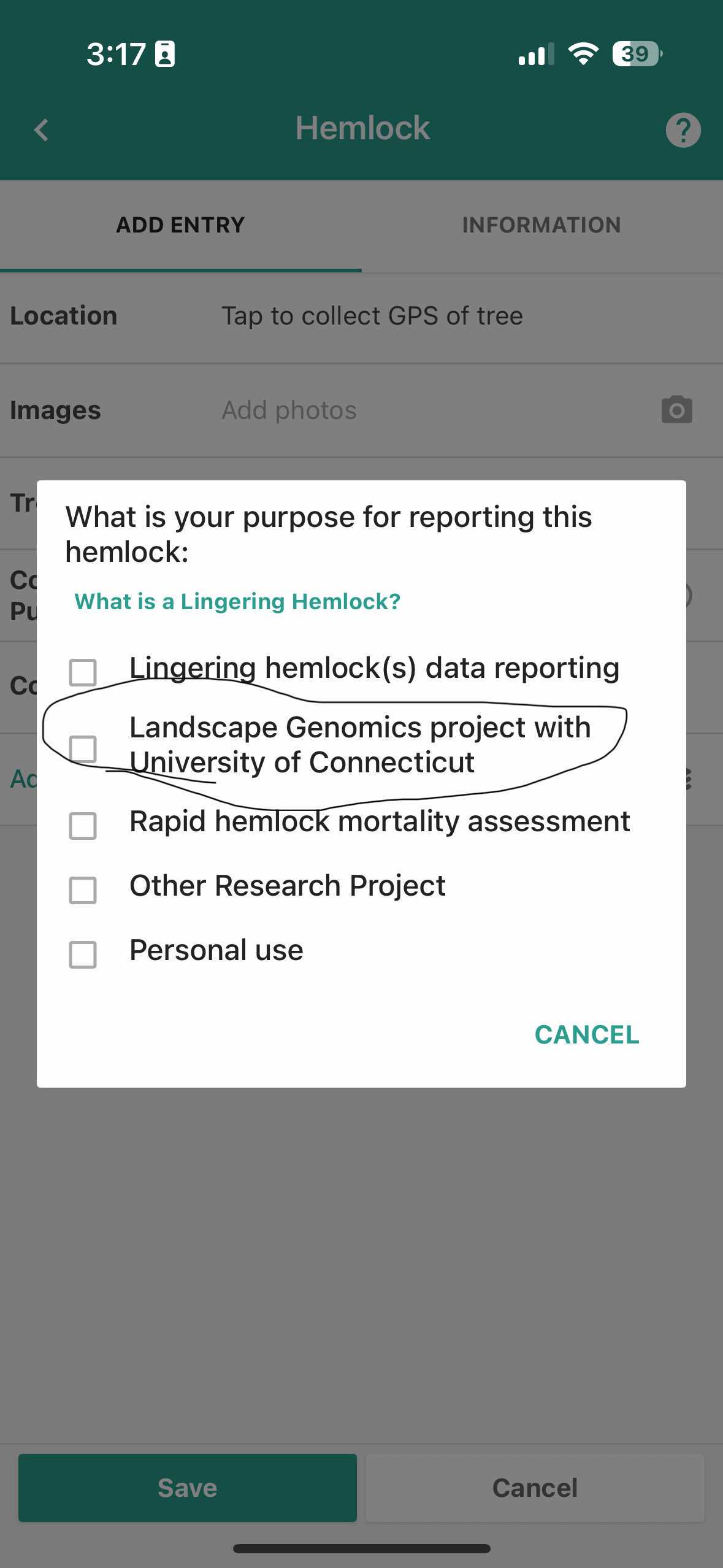
Screenshot of choices of collection purpose
Once selected, the app opens up more categories of metadata including details of stressors, HWA and EHS presence, and the presence of cones.
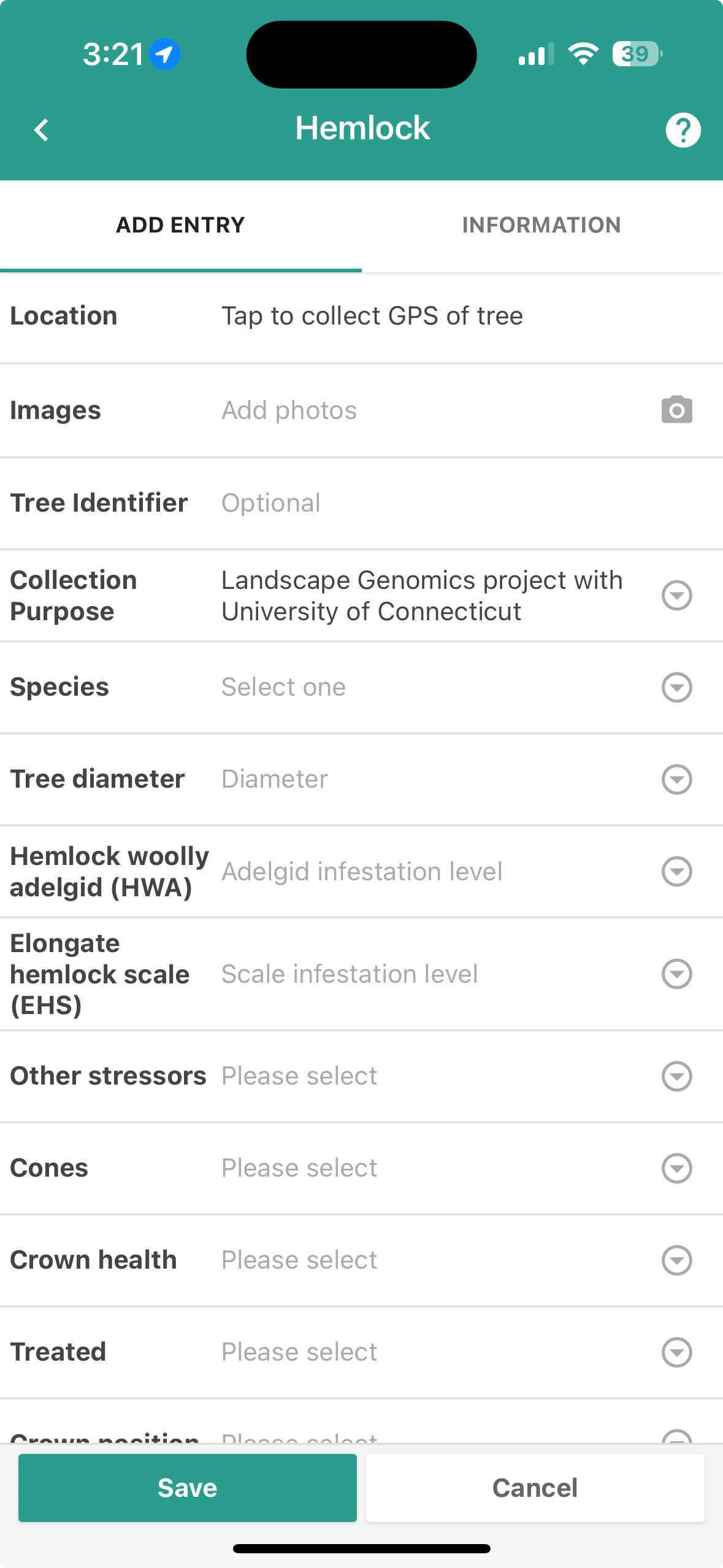
Enter information into each field as applicable.
- In the 'Tree Identifier' field, name each collection with "your initials"-"MonthDateYear"-"CollectionNumber". For example, my first collection for today (7/19/23) would be KF-071923-01.
- Collect or estimate the diameter at breast height (DBH) using inches.
- In the comments field, add a short description of the site. For example, "Tree located at the Happy Logger trail head", or "Near intersection of Pond St and VT 116".
The lat-longs of the tree are automatically saved when you create the entry (not when you save it!). So be sure to create the entry next to the tree. If you have a GPS that is more precise than your phone and you wish to submit those points instead, enter them in the comments field.
Don't worry if you don't have cell phone reception in the field. The data is temporarily saved on your phone. After you connect to cellular service or wifi again, open the app and select the yellow pop-up box to upload data to TreeSnap.
Save the entry and move on the next tree.
Tree Tagging (Optional)
A tagged tree is incredibly valuable, as it allows for a confirmed recollection of tissue or germplasm. When tagging your tree, angle the galvinized nail at a 30 - 45 degree angle and allow the tag to freely dangle. Hammer the nail approximately an inch into the tree.
If tagging a tree is not an option, then take descriptive photographs that have landmarks.
After you've collected
After you've made your collections. Be sure your samples are in the resealable plastic bag. Put the bag containing samples in the mailer with an ice pack and reseal it. Using the provided postage, mail the samples to:
Mary Rutter
C/O Rachel O'Neill
University of Connecticut
67 North Eagleville Rd
U-3197, ESB Room 209
Storrs, CT 06269
If you don't have an ice pack, please let us know, and we will provide one to you.
