Jun 13, 2025
UMN-TMCs: CosMx Spatial Molecular Imager for FFPE Tissue – RNA 1K-Plex (v2)
- Samuel Peters1,
- Laura Niedernhofer1,
- Paul Robbins1
- 1University of Minnesota, Minneapolis, MN
- Cellular Senescence Network (SenNet) Method Development Community

Protocol Citation: Samuel Peters, Laura Niedernhofer, Paul Robbins 2025. UMN-TMCs: CosMx Spatial Molecular Imager for FFPE Tissue – RNA 1K-Plex (v2). protocols.io https://dx.doi.org/10.17504/protocols.io.3byl41p7zlo5/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: June 12, 2025
Last Modified: June 13, 2025
Protocol Integer ID: 220089
Keywords: SenNet, CosMx, Spatial transcriptomics, cosmx spatial molecular imager for ffpe tissue, university of minnesota tissue mapping center, plex cosmx rna data, cosmx spatial molecular imager, minnesota tissue mapping center, bruker spatial biology, property of bruker spatial biology, export data from the atomx spatial informatics platform, atomx spatial informatics platform, ffpe tissue, rna, individual rna transcript, rna 1k, plex in situ visualization, nm spatial resolution, protein stain, imaging, tissue, spatial resolution, situ visualization, transcript
Funders Acknowledgements:
NIH
Grant ID: 5U54AG076041-04
NIH
Grant ID: 5U54AG079754-03
Disclaimer
DISCLAIMER – FOR INFORMATIONAL PURPOSES ONLY; USE AT YOUR OWN RISK
The protocol content here is for informational purposes only and does not constitute legal, medical, clinical, or safety advice, or otherwise; content added to protocols.io is not peer reviewed and may not have undergone a formal approval of any kind. Information presented in this protocol should not substitute for independent professional judgment, advice, diagnosis, or treatment. Any action you take or refrain from taking using or relying upon the information presented here is strictly at your own risk. You agree that neither the Company nor any of the authors, contributors, administrators, or anyone else associated with protocols.io, can be held responsible for your use of the information contained in or linked to this protocol or any of our Sites/Apps and Services.
Abstract
The CosMx Spatial Molecular Imager (SMI) enables high-plex in situ visualization of individual RNA transcripts and/or protein stain overlays with 120 nm spatial resolution. This protocol outlines basic criteria for sample preparation or selection; tissue slide processing; and imaging to generate 1,000-plex CosMx RNA data and export data from the AtoMx Spatial Informatics Platform (SIP). The steps herein – based largely on materials developed by and property of Bruker Spatial Biology, Inc. – are specific procedures followed by the University of Minnesota Tissue Mapping Centers (TMCs) as part of the NIH Common Fund's SenNet Consortium and are not officially condoned by Bruker Spatial Biology.
Troubleshooting
Sample Requirements
Prospective or retrospective samples embedded in paraffin blocks should meet the following criteria:
From the time of excision*, tissue samples should be or should have been placed in 10% neutral-buffered formalin within 1 hour to prevent RNA degradation.
*For tissues collected prospectively for CosMx SMI, consider perfusing with cold fixative (e.g. 4% paraformaldehyde), particularly for tissues >5 mm in width on any axis. This will help preserve RNA deeper in the tissue.
Tissues should be or should have been fixed in 10% neutral-buffered formalin for 18-24 hours.
Following fixation, tissues should be or should have been transferred to ethanol (e.g. 70%) or passed through ethanol gradients, then embedded in paraffin blocks.
Following embedding, blocks are stable for long-term storage at room temperature. However, blocks older than 10 years should be used with caution.
It may be prudent to determine DV200 values for samples intended for CosMx, particularly older samples. It has been demonstrated that the assay may still perform well on samples with degraded RNA (DV200 values of approximately 20%). However, in many cases, more intact RNA (DV200 >50%) produces higher quality results for other such in situ hybridization assays.
Formalin-fixed paraffin-embedded (FFPE) blocks should be sectioned using a calibrated microtome with a clean blade using the following general guidelines:
Trim the block face at least 20-30 μm before cutting sections for CosMx analysis.
Cut sections to 5 μm thickness. Sections may cut up to 10 μm; however, only the 5 μm closest to the slide will be imaged by the CosMx instrument.
Carefully mount the sections on a 75 mm x 25 mm glass slide within the CosMx imageable area. VWR SuperFrost Plus Micro, Leica BOND PLUS, or other charged slides are recommended.
Some tissue outside the imageable area is okay, but will later need to be trimmed to allow proper adhesion of the flow cell coverslip.
Ensure each mounted section is free of folds, cracks, wrinkles, bubbles, or other anomalies.
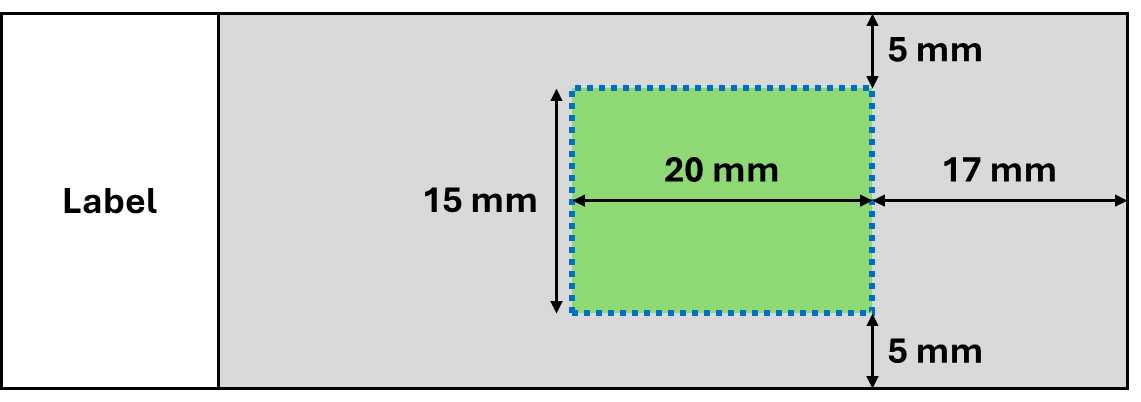
Only tissue within the imageable area (green) can be captured during acquisition.
Optionally, slides may be baked at 37ºC for 2 hours to improve tissue adherence.
Dry the slides at room temperature overnight.
Slides may be stored in a desiccator at room temperature or 4ºC for up to 2 weeks.
Deparaffinization
Incubate slides vertically in a slide rack at 60ºC overnight.
Prior to removing slides from the 60ºC incubation, prepare the following items:
Prepare NBF Stop Buffer:
| Reagent | Total volume | Component | Amount | Vendor | Cat. number | |
| NBF Stop Buffer (0.1 M) | 500 mL | Tris Base | 6.06 g | Roche | 10708976001 | |
| Glycine | 3.75 g | Invitrogen | 15527-013 | |||
| DEPC-treated water | To 500 mL | Invitrogen | AM9922 |
NBF Stop Buffer may be prepared up to two weeks in advance.
Equilibrate sulfo-NHS acetate powder (e.g. Thermo Fisher Scientific #26777) to room temperature prior to opening. Once equilibrated, aliquot approximately 15 mg (for 2 slide runs) or 25 mg (for 4 slide runs) into separate microcentrifuge tubes, annotating the exact mass on each tube. If sulfo-NHS-acetate has already been aliquoted, skip this step. Store in -20°C until the end of "Fiducial Application".
Fill staining jars with following solutions and label them accordingly:
- 2 jars of reagent-grade xylenes: Xylenes 1 & Xylenes 2
- 2 jars of reagent-grade 200-proof (100%) ethanol: Ethanol 1 & Ethanol 2
Label 1 slide staining jar Target Retrieval. Prepare 1X Target Retrieval Solution in this staining jar by combining 1 part 10X Target Retrieval Solution with 9 parts DEPC-treated water.
| Reagent | Total volume | Component | Volume | Vendor | Cat. number | |
| 1X Target Retrieval Solution | e.g. 100 mL | 10X Target Retrieval Solution | 10 mL | NanoString Technologies | 121500006 (Box 1) | |
| DEPC-treated water | 90 mL | e.g. Invitrogen | e.g. AM9922 |
Volumes for a staining jar with 100 mL capacity,
Place the jar labeled Target Retrieval, containing 1X Target Retrieval Solution, in a laboratory-grade tissue pressure cooker (e.g. Bio SB TintoRetriever). Fill the reservoir with water to a level about halfway up the height of a slide staining jar, well below the jar lid. Ensuring the lid is not air-tight, equilibrate the pressure cooker to 100ºC.
Remove slides from the oven and briefly allow to cool, about 5 minutes, keeping the oven on and set to 60ºC. Then deparaffinize the tissue:
Transfer the slides to Xylenes 1 and incubate for 5 minutes at room temperature.
Transfer the slides to Xylenes 2 and incubate for 5 minutes at room temperature.
Transfer the slides to Ethanol 1 and incubate for 2 minutes at room temperature.
Transfer the slides to Ethanol 2 and incubate for 2 minutes at room temperature.
Remove the slides from Ethanol 2. Dry the tissues at 60ºC for 5 minutes. Dried slides may remain at room temperature briefly until ready to proceed.
Target Retrieval
Initiate target retrieval:
Once the pressure cooker has equilibrated to temperature, cancel the incubation. If not already done, open the lid valve to release all pressure from the cooker prior to opening.
Carefully remove the lid and, using heat-resistant gloves, take out the heated 1X Target Retrieval Solution (Target Retrieval).
Places the tissue slides into Target Retrieval and return the jar to the pressure cooker, ensuring that the jar is covered but not completely sealed.
Replace and secure the lid on the pressure cooker and return to 100ºC. Ensure the pressure release valve is open. Once the temperature has re-equilibrated to 100°C, begin timing target retrieval. The UMN-TMCs have used the following retrieval time(s) on the indicated tissues:
| Tissue type | Retrieval time (min) | |
| Human Liver (FFPE) | 15 | |
| Mouse Liver (FFPE) | 15 | |
| Mouse Brain (FFPE) | 15 |
During target retrieval, fill staining jars with following solutions and label them accordingly:
- 1 slide staining jar filled with fresh DEPC-treated water: DEPC 1
- 1 slide staining jar filled with fresh 200-proof (100%) ethanol: Ethanol 3
When target retrieval is finished, cancel the incubation. Carefully remove the lid and take out the heated 1X Target Retrieval Solution.
Immediately transfer tissue slides to DEPC 1. Immerse and remove slides repeatedly for 15 seconds to wash.
Transfer slides to Ethanol 3 and incubate for 3 minutes at room temperature.
Remove slides from Ethanol 3 and lay flat to dry in a clean, dust-free area for 30 minutes to 1 hour.
Tissue Digestion
While slides are drying, prepare Diluted Proteinase K. 1X phosphate-buffered saline (PBS) should be prepared from DEPC-treated water. Pipette mix and centrifuge briefly.
| Reagent | Total volume | Component | Volume | Vendor | Cat. number | |
| Diluted Proteinase K (200 μg/mL) | 200 μL | Proteinase K (20 mg/mL) | 2 μL | NanoString Technologies | 121500006 | |
| 1X PBS | 198 μL | Various | Various |
Create a serial dilution of Proteinase K (Digestion Buffer) for tissue application. Pipette mix and centrifuge briefly, then maintain on ice until ready to use. The concentration of Proteinase K in the Digestion Buffer may vary by tissue. The UMN-TMCs have used the following concentration(s) on the indicated tissues:
| Tissue type | Proteinase K concentration | |
| Human Liver (FFPE) | 3 μg/mL | |
| Mouse Liver (FFPE) | 3 μg/mL | |
| Mouse Brain (FFPE) | 3 μg/mL |
| Reagent | Total volume | Component | Volume | Vendor | Cat. number | |
| Digestion Buffer | 880 μL | Diluted Proteinase K (200 μg/mL) | 13.2 μL | Prepared above | Prepared above | |
| 1X PBS | 866.8 μL | Various | Various |
Volumes needed to create Digestion Buffer with 3 μg/mL Proteinase K for 2 slides. Multiply values by 2 for 4-slide runs.
Prepare a clean hybridization tray. This should be RNAse-AWAY treated (Thermo-Fisher Scientific #7002), as it can be a potent source of oligonucleotide contamination.
Line the tray bottom with clean laboratory wipes dampened with DEPC-treated water or nuclease-free 2X SSC. Set the slide rack over the wipes.
Insert the hybridization tray into a compatible hybridization oven (e.g. Boekel Scientific RapidFISH Slide Hybridizer) and pre-heat to 40°C. If the hybridization tray does not have a water-tight seal, place it in a zip-lock bag and seal before inserting into the oven.
Place an incubation frame, located in NanoString Technologies 121500006 (Box 1), onto each slide around the tissue and imageable area of the slide:
Ensure that the slide area that will contact the Incubation Frame adhesive is clean and dry. If necessary, use a clean scalpel blade to trim the tissue near the borders to fit entirely within the Incubation Frame.
Remove the thin polyester sheet from the Incubation Frames to expose the adhesive. Ensure the thicker polyester backing (with center square removed) remains attached.
On a flat surface, center the incubation frame over the tissue with the frame borders flush to the slide edges on the short end. The incubation frame should be aligned to the imageable area of the slide. Lightly press the border to ensure the incubation frame is well-adhered to the slide.
Using a sharp clean blade, trim excess plastic from the incubation frame to ensure there are no overhangs on the slide edges.
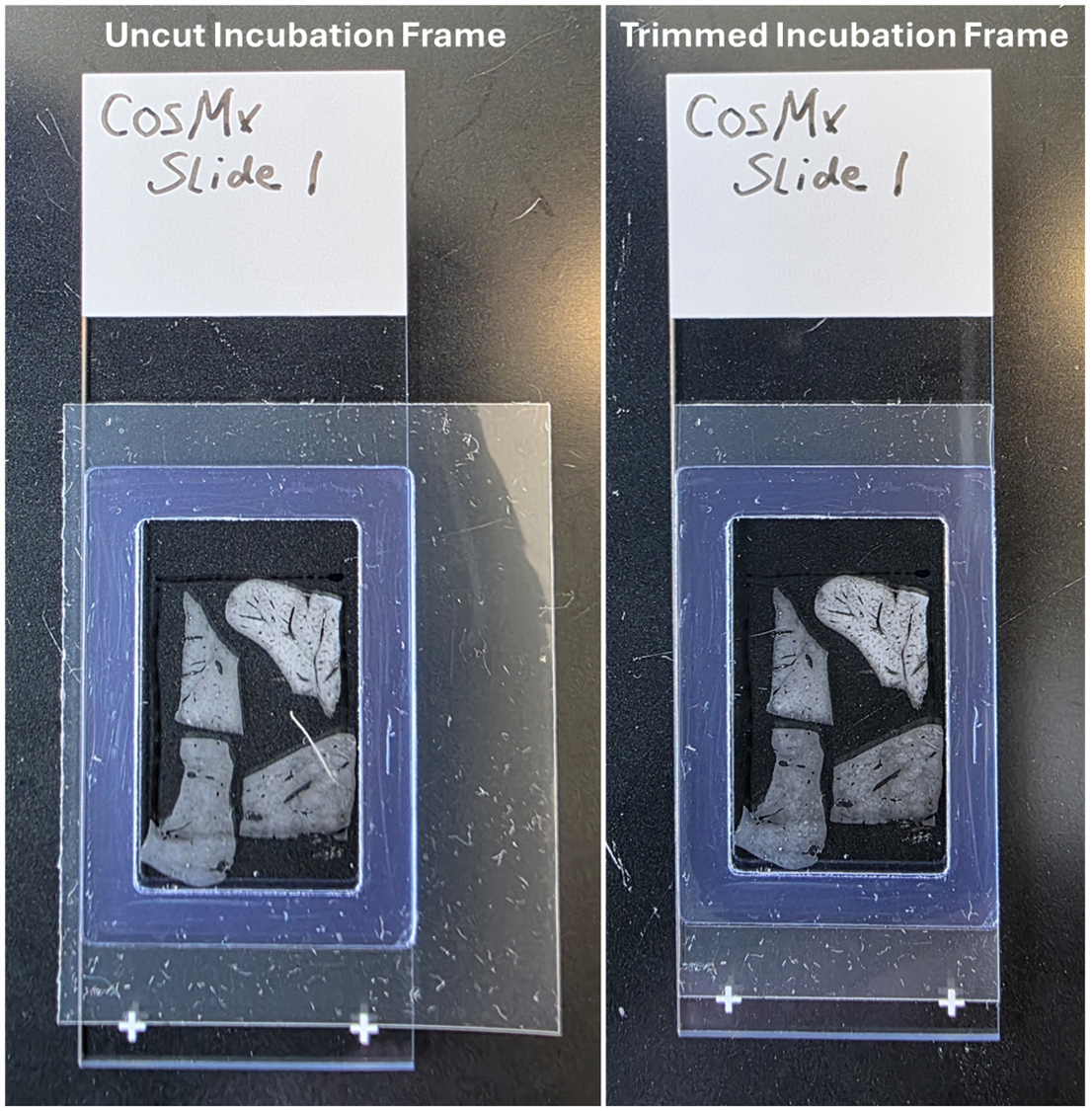

At this stage, the incubation frame should completely surround the tissue and imageable area; should be trimmed flush with the slide edges; and the plastic backing with the center hole should still be attached.
Fill two slide staining jars with fresh DEPC-treated water. Label the jars DEPC 2 and DEPC 3.
Initiate tissue digestion:
Remove the Digestion Buffer from ice and warm to 37°C (e.g. by hand) for approximately 3 minutes.
Remove the pre-heated hybridization tray from the hybridization oven. Set each slide, with incubation frames applied, tissue side-up into the slide rack insert of the tray.
Slowly add 400 µL of Digestion Buffer to each slide. If needed, gently tap the slide to spread the liquid to cover the entire tissue, then return each slide to a slot in the rack of the hybridization tray.
Replace the hybridization tray lid – seal the chamber, if necessary – and re-insert the tray into the hybridization oven, still set to 40°C. Begin timing the tissue digestion step. The UMN-TMCs have used the following tissue digestion time(s) on the indicated tissues:
| Tissue type | Digestion time (min) | |
| Human Liver (FFPE) | 30 | |
| Mouse Liver (FFPE) | 30 | |
| Mouse Brain (FFPE) | 30 |
During tissue digestion, remove the CosMx fiducials and 2X SSC-T from 4°C and equilibrate to room temperature protected from light for at least 10 minutes. These are located within NanoString Technologies 121500006 (Box 1).
When tissue digestion is complete, remove the hybridization tray from the oven and, one slide at a time, gently tap off the Digestion Buffer onto a clean laboratory wipe. Transfer the slides to DEPC 2 and move the slides up and down 3-5 times to wash.
Transfer slides to DEPC 3 and move the slides up and down 3-5 times to wash once more. Slides may stay submerged in DEPC 3 briefly until Diluted Fiducials are ready to apply.
Fiducial Application
Prepare a clean light-impermeable slide staining tray. Partially fill the basin with DEPC-treated water of nuclease-free 2X SSC. Lining with laboratory wipes can help prevent splashing of liquid.
Once the CosMx fiducials have equilibrated to room temperature for at least 10 minutes, prepare them as below:
- Vortex for 1 minute;
- Sonicate in an ultrasonic water bath for 2 minutes;
- Vortex for 1 minute;
- Sonicate in an ultrasonic water bath for 2 minutes;
- Vortex for 1 minute.
Immediately prepare Diluted Fiducials. 0.001% (1:100) is recommended unless otherwise advised.
| Reagent | Total volume | Component | Volume | Vendor | Cat. number | |
| Diluted Fiducials (0.001%) | 1200 µL | CosMx Fiducials | 12 µL | NanoString Technologies | ||
| 2X SSC-T | 1188 µL |
Volumes for both 2-slide and 4-slide runs. Only 250 µL per slide is needed, but it is recommended that no less than 1 mL of Diluted Fiducials be prepared at a time due to the tendency of fiducials to aggregate.
Remove slides from DEPC 3 and gently tap off excess liquid onto a clean laboratory wipe. Lay each slide tissue-side up in the staining tray.
Vortex Diluted Fiducials for 1 minute before applying up to 250 µL to each slide. Vortex Diluted Fiducials again for 30 seconds again between slides. Gently tap the slides as needed to ensure solution covers the entire area – tissue and glass – within the incubation frame.
With Diluted Fiducials applied, cover the slide staining tray and incubate slides at room temperature for 5 minutes.
During the incubation, fill staining jars with following solutions and label them accordingly:
- 2 jars of nuclease-free 1X PBS (prepared with DEPC-treated water): 1X PBS 1 & 1X PBS 2
- 1 jar of 10% neutral-buffered formalin (e.g. StatLab Medical Products 28600-1): 10% NBF
- 2 jars of NBF stop buffer (prepared earlier): NBF Stop 1 & NBF Stop 2
When incubation is complete, gently tap off Diluted Fiducials solution onto a clean laboratory wip
Transfer slides into 1X PBS 1 and wash for 1 minute at room temperature.
Post-fix the tissues:
Transfer the slides to 10% NBF and incubate for 1 minute at room temperature.
Transfer the slides to NBF Stop 1 and incubate for 5 minutes at room temperature.
Transfer the slides to NBF Stop 2 and incubate for 5 minutes at room temperature.
Transfer the slides to 1X PBS 2 and incubate for 5 minutes at room temperature. Slides may be kept in this jar briefly until read to proceed to 'NHS Acetate Blocking'.
NHS Acetate Blocking
Complete the following preparation steps before proceeding:
Remove aliquoted (15 mg or 25 mg) sulfo-NHS acetate powder from -20 °C and equilibrate to room temperature.
Remove Buffer R and NHS Acetate Buffer from 4°C and equilibrate to room temperature. These are within NanoString Technologies 121500006 (Box 1).
Remove CosMx RNase Inhibitor (121500004) from -20°C and put on ice.
Remove RNA probes from -20°C and put on ice. This includes base RNA panels, add-on panels, and RNA-based segmentation markers (if applicable).
Fill 2 slide staining jars with 2X SSC. Label these jars 2X SSC 1 and 2X SSC 2.
Pre-heat a hybridization oven to 37°C.
Add 38.5 µL NHS Acetate Buffer per mg to aliquoted sulfo-NHS acetate poweder (e.g. 577 µL of buffer would be added to 15.0 mg of powder). Pipette mix, vortex, and centrifuge briefly to ensure all powder is dissolved. This makes a 100 mM NHS Acetate solution.
Remove slides from 1X PBS 2 and gently tap off excess liquid onto a clean laboratory wipe. Lay each slide tissue-side up in the slide staining tray.
To each slide, apply 200-250 µL of 100 mM NHS Acetate, ensuring the entire tissue is covered.
Incubate protected from light at room temperature for 15 minutes.
Following incubation, gently tap off 100 mM NHS Acetate onto a clean laboratory wipe.
Wash the slides:
Transfer the slides to 2X SSC 1 and incubate for 5 minutes at room temperature.
Transfer the slides to 2X SSC 2 and incubate for 5 minutes at room temperature. After this final wash step, slides may be kept in in this jar for up to 1 hour at room temperature, or up to 6 hours at 4°C.
Probe Hybridization
Complete the following preparation steps before proceeding:
Pre-heat a thermal cycler and lid to 95°C.
For each slide, remove 1 incubation frame cover, located in NanoString Technologies 121500006 (Box 1), and clean with ethanol. Dry with a clean laboratory wipe and ensure all dust is removed. Set aside incubation frame covers on a clean laboratory wipe until use.
Prepare a clean hybridization tray. This should be RNAse-AWAY treated (Thermo-Fisher Scientific #7002), as it can be a potent source of oligonucleotide contamination. Line the tray bottom with clean laboratory wipes dampened with DEPC-treated water or nuclease-free 2X SSC. Set the slide rack over the wipes. Insert the hybridization tray into the pre-heated 37°C hybridization oven. If the hybridization tray does not have a water-tight seal, place it in a zip-lock bag and seal before inserting into the oven.
Once RNA probes (and RNA segmentation markers, if applicable) are thawed, flick to mix and centrifuge. Do not vortex.
Transfer an aliquot of the exact volume of each RNA probe/RNA segmentation marker to be used into a separate 0.2-mL PCR tubes. Do not combine aliquots yet and only use what is needed for the assay (indicated in step 38.8).
Heat aliquoted RNA probes / RNA segmentation markers at 95°C on the thermal cycler for 2 minutes with the lid closed.
Crash-cool the aliquoted RNA probes/RNA segmentation markers on ice for 1 minute immediately following 95°C incubation. Leave RNA probes / RNA segmentation markers on ice until use.
Prepare the Hybridization Mix as indicated in the appropriate table. After combining reagents, gently pipette mix and centrifuge. Do not vortex.
| Reagent | Total volume | Component | Volume | Vendor | Cat. number | |
| Hybridization Mix | 320 µL | Buffer R | 256 µL | NanoString Technologies | 121500006 (Box 1) | |
| RNase inhibitor | 3.2 µL | NanoString Technologies | 121500004 / 121500005 | |||
| 950-plex base RNA panel | 32 µL | NanoString Technologies | Various | |||
| 7-50 plex add-on panel | 16 µL | NanoString Technologies | Various | |||
| RNA segmentation markers (if applicable) | 12.8 µL | NanoString Technologies | 121500024 | |||
| DEPC-treated water | Up to 320 µL (if needed) | e.g. Invitrogen | e.g. AM9922 |
Volumes for 2-slide runs (with or without RNA segmentation markers). Multiply values by 2 for 4-slide runs.
Complete the following steps one slide at a time:
Remove each slide from 2X SSC 2 and gently tap off excess liquid onto a clean laboratory wipe.
Remove the remaining thick polyester layer from the incubation frame to expose the adhesive layer of the top.
Lay the slide flat and apply up to 150 µL of Hybridization Mix to the slide, ensuring the entire tissue is covered.
Carefully apply the incubation frame cover, starting at the end of the incubation frame closest to the slide label and gradually lowering the cover at an angle. Avoid introducing bubbles.
Place the covered slide into the slide rack insert of the hybridization chamber. Repeat for each slide in the set until all slides are in the hybridization chamber.
Cover the hybridization tray – seal, if necessary – and insert into the pre-heated 37°C hybridization oven. Incubate for 16 to 18 hours. Do not exceed 18 hours. Record exact hybridization times to the minute.
Remove lyophilized catalase and pyranose oxidase (P20X) from 4°C and centrifuge.
Reconstitute enzymes in 250 µL (2 slide runs) or 400 µL (4 slide runs) of DEPC-treated water. Pipette mix, vortex, then centrifuge. Ensure that all lyophilized enzyme is dissolved, as clumps can clog the instrument fluidics systems.
Add 250 µL (2 slide runs) or 400 µL (4 slide runs) of Reconstituted Catalase and 125 µL (2 slide runs) or 200 µL (4 slide runs) of Reconstituted Pyranose Oxidase to CosMx Buffer 4, in NanoString Technologies 1004080 (Box 1). Indicate on the bottle that enzymes have been added.
Stringent Washes
Complete the following preparation steps before proceeding:
Set a water bath to 37°C.
Remove nuclear stain and cell segmentation kits from -80°C. Thaw the nuclear stain at room temperature and the cell segmentation markers on ice. Any RNA segmentation markers should have been used in the Hybridization Mix on Day 1.
Warm 100% deionized formamide (e.g. Invitrogen AM9344) to 37°C in the water bath for at least 30 minutes prior to opening. It can be helpful to prepare single-use aliquots in advance.
If the time is approaching the 18-hour mark, transfer slides to a jar containing 2X SSC while reagents and equipment are being prepared (see steps 39.7, 40.1, & 40.2).
Create Stringent Wash Buffer by combining 1 part 4X SSC with 1 part 100% warmed deionized formamide.
Fill two staining jars with Stringent Wash Buffer and label them Stringent Wash 1 and Stringent Wash 2. Heat these staining jars in the 37°C water bath, with the water level well below the lid, for at least 30 minutes.
Fill three staining jars with fresh 2X SSC. Label these jars Pre-Wash 2X SSC, 2X SSC 3, and 2X SSC 4.
Complete the following steps one slide at a time:
Remove each slide from the hybridization chamber and use a clean forceps to remove the incubation frame cover. If the cover cannot be removed without removing the incubation frame, remove the incubation frame. A new one can be applied at a later time (see 42.3). If needed, periodically dip this slide into 2X SSC to prevent the tissue from drying until the incubation frame cover is removed.
Place the slide into the jar labeled Pre-Wash 2X SSC, clean the forceps with ethanol, and repeat the steps above until all slides are in this jar.
Wash the slides:
Transfer the slides to Stringent Wash 1 and incubate for 25 minutes at 37°C.
Transfer the slides to Stringent Wash 2 and incubate for 25 minutes at 37°C.
Transfer the slides to 2X SSC 3 and incubate for 2 minutes at room temperature.
Transfer the slides to 2X SSC 4 and incubate for 2 minutes at room temperature. Slides may remain in this jar briefly while nuclear stain reagents are being prepared.
Nuclear Stain & Blocking
Complete the following preparation steps before proceeding:
Fill three staining jars with fresh 1X PBS. Label these jars 1X PBS 3, 1X PBS 4, 1X PBS 5, and 1X PBS 6.
Prepare a clean light-impermeable slide staining tray. Partially fill the basin with DEPC-treated water of nuclease-free 2X SSC. Using laboratory wipes to soak up liquid can help prevent splashing of liquid.
Reapply a new incubation frame to each slide, if needed (see step 15).
Prepare the Nuclear Stain Buffer as indicated.
| Reagent | Total volume | Component | Volume | Vendor | Cat. number | |
| Nuclear Stain Buffer | 440 µL | CosMx DAPI Nuclear Stain | 11 µL | NanoString Technologies | 121500020 | |
| CosMx RNA Blocking Buffer | 429 µL | NanoString Technologies | 121500006 (Box1) |
Volumes for 2-slide runs. Multiply values by 2 for 4-slide runs:
Complete the following steps one slide at a time:
Remove the slide from 2X SSC 4 and gently tap off excess liquid onto a clean laboratory wipe. Wick away excess liquid from the corners of the Incubation Frame using a clean laboratory wipe.
Lay each slide horizontally in the slide staining tray with the tissue side facing up.
Apply up to 200 µL of Nuclear Stain Buffer to each slide within the incubation frame. Gently tilt the slide back and forth to ensure the entire tissue is covered with buffer. Repeat these steps until all slides are covered in Nuclear Stain Buffer.
Cover the slide staining tray with a light-impermeable lid and incubate for 15 minutes at room temperature.
After incubation, remove slides from the tray and gently tap off Nuclear Stain Buffer onto a clean laboratory wipe.
Wash slides for 5 minutes 1X PBS 3. Slides may remain in this jar briefly while Segmentation Mix reagents are being prepared.
Segmentation Marker Stain
Prepare the Segmentation Mix. Reagent volumes will depend on the selected kit.
| Reagent | Total volume | Component | Volume | Vendor | Cat. number | |
| Segmentation Mix | 400 µL | CosMx CD298/B2M Marker, Ch2 | 8 µL | NanoString Technologies | 121500020 / 121500042 | |
| CosMx PanCK/CD45 Marker Mix, Ch3/Ch4 | 16 µL* | NanoString Technologies | 121500020 / 121500042 | |||
| CosMx A La Carte Marker, Ch5 | 16 µL* | NanoString Technologies | Various | |||
| CosMx RNA Blocking Buffer | Up to 400 µL | NanoString Technologies | 121500006 (Box 1) |
[Universal Cell Characterization Markers] Volumes for 2-slide runs. Multiply values by 2 for 4-slide runs. Listed catalog numbers are for Human / Mouse Universal Cell Characterization RNA segmentation kits, respectively.
*These markers are optional. Only DAPI (Ch1) and CD298/B2M (Ch2) are essential for segmentation.
| Reagent | Total volume | Component | Volume | Vendor | Cat. number | |
| Segmentation Mix | 400 µL | CosMx Neuro Histone Marker, Ch3 | 16 µL | NanoString Technologies | 121500024 | |
| CosMx Neuro GFAP Marker, Ch4 | 16 µL | NanoString Technologies | 121500024 | |||
| CosMx RNA Blocking Buffer | 368 µL | NanoString Technologies | 121500006 (Box 1) |
[Neuroscience Markers] Volumes for 2-slide runs. Multiply values by 2 for 4-slide runs. Listed catalog numbers are for the Mouse Neuroscience RNA segmentation kit.
Complete the following steps one slide at a time:
Remove each slide from 1X PBS 3 and gently tap off excess onto a clean laboratory wipe.
Lay each slide horizontally in the slide staining tray with the tissue side facing up.
Apply up to 200 µL of Segmentation Mix to each slide within the Incubation Frame. Gently tilt the slide back and forth to ensure the entire tissue is covered. Repeat these steps until all slides are covered in Segmentation Mix.
Cover the slide staining tray with a light-impermeable lid and incubate for 1 hour at room temperature.
While slides are staining, remove the CosMx RNA imaging tray from 4°C and equilibrate to room temperature for at least 1 hour.
1000-plex, 2-slides: NanoString Technologies 122000156
1000-plex, 4-slides: NanoString Technologies 122000157
After incubation, remove slides from the tray and gently tap off Segmentation Mix onto a clean laboratory wipe.
Wash the slides:
Transfer the slides to 1X PBS 4 and incubate for 5 minutes at room temperature.
Transfer the slides to 1X PBS 5 and incubate for 5 minutes at room temperature.
Transfer the slides to 1X PBS 6 and incubate for 5 minutes at room temperature.
After the last wash step, use a clean forceps to carefully remove the incubation frame from all slides one at a time. After the incubation frame has been removed, keep each slide in 1X PBS to avoid drying. Slides may be assembled into flow cells immediately, or transferred to 2X SSC and stored overnight at 4°C.
Flow Cell Assembly
Complete the following preparation steps before proceeding:
Clean the benchtop and workspace with RNase AWAY (Thermo-Fisher Scientific #7002) or 70% ethanol.
Clean the flow cell assembly tool (NanoString Technologies 903703) with ethanol or isopropanol. Use a pressurized air cannister to blow off dust.
Inspect flow cell coverslips (NanoString Technologies 122000061) for cracks or chips. Do not use damaged flow cell coverslips.
Remove slides from buffer one at a time and dry the back of the slide. Using the slide template on the flow cell assembly tool, carefully dry the area of the slide where the flow cell coverslip will adhere. Do not touch the area that was within the incubation frame: the glass around the tissue will contain fiducials. If the slide has an adhesive slide label, ensure that it is not more than 295 μm thick and not folded over on itself. If necessary, trim the slide label with a clean razor blade.
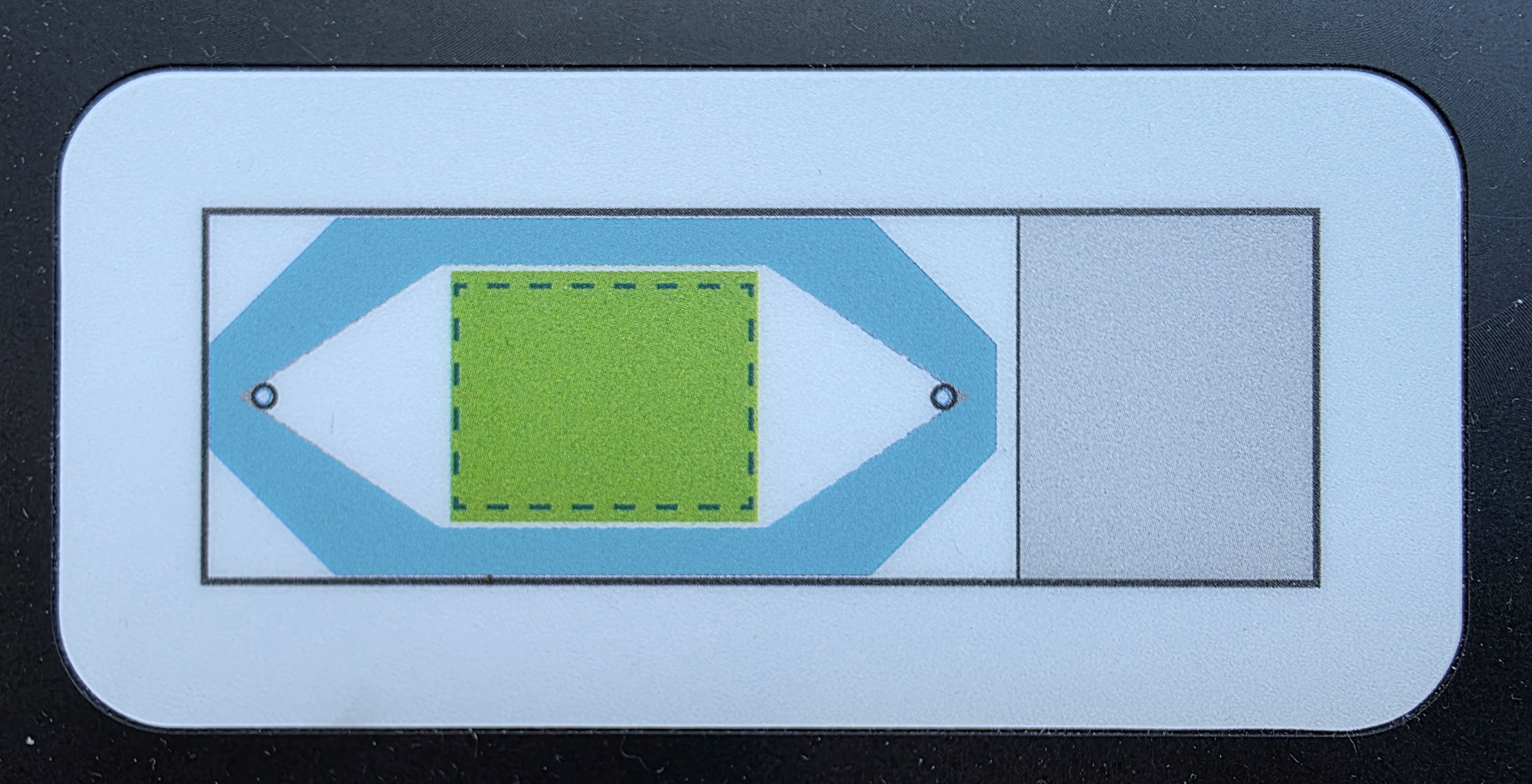

The area in blue surrounding the green box (indicating the imageable area) should be dry to properly adhere the flow cell coverslip.
Use the flow cell assembly tool to apply the flow cell coverslip to the tissue slide:
With the arm of the flow cell assembly tool open, lower the tailgate (1). With the tissue-side up, fully insert the slide into the bottom opening of the slide stage with the label on the tailgate side. Ensure the unlabeled end of the slide contacts the back of the slide stage (2), then raise the tailgate up (3) to secure the slide.
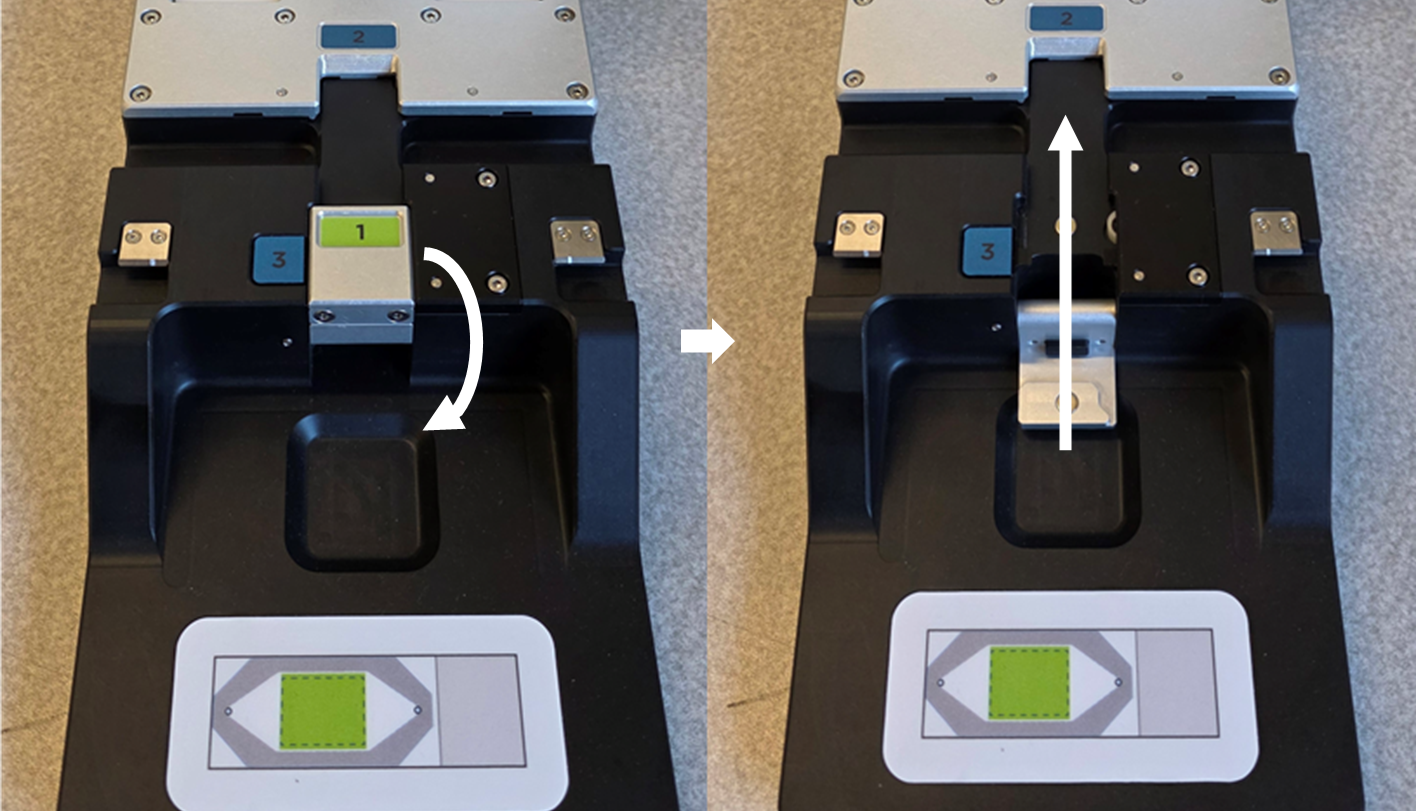

With the arm of the flow cell assembly tool raised and the slide tailgate (1) lowered, the slide can be inserted into the stage area with the unlabeled side toward position 2 (right).
Carefully remove the adhesive backing from the flow cell coverslip using a gloved hand or clean forceps.
Adhesive side down, carefully place the flow cell coverslip onto the slide. One edge should line up with the edge of the unlabeled side of the slide at position 2.

The barcode side of the flow cell coverslip may be placed at either end of the tissue slide. Line up the end of the flow cell coverslip, held at an angle, with the groove by position 2 on the flow cell assembly tool. The corner of the flow cell coverslip should contact the tissue slide. Without allow the slide to catch on any surface of the flow cell assembly tool, carefully place it on top of the slide between position 2 and the raised tailgate (1).
Lightly tap the four corners of the flow cell coverslip to ensure it is not misaligned or catching anywhere on the flow cell assembly tool. Dark patches should form where the flow cell coverslip has begun to adhere to the slide.
Bring the arm of the flow cell assembly tool down and apply light pressure until both latches of the tool have engaged. You should hear a click and the release buttons will pop out.
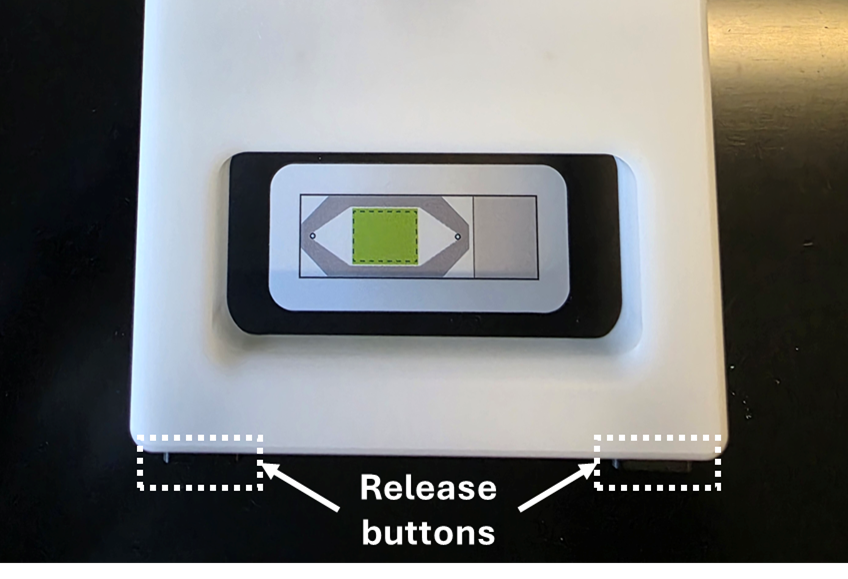

The release buttons will click and pop out when the latches have engaged. To re-open the flow cell assembly tool, push in the release buttons and raise the arm.
Push the release buttons to disengage the latches of the flow cell assembly tool and raise the arm. Inspect the flow cell coverslip for cracks, chips, or damage. The flow cell coverslip should be fully adhered to the slide.
Lower the tailgate (1) of the flow cell assembly tool and remove the flow cell constructed from the slide and flow cell coverslip.
Flow in 200 μL of storage buffer (2X SSC) by carefully placing a 200 µL pipette tip directly on one of the fluidics ports of the flow cell, without applying too much much pressure, and slowly pressing the pipette plunger to allow buffer to slowly fill the chamber until it flows out of the other port. Alternatively, the flow cell may be dipping vertically into a 50-mL conical tube filled with fresh 2X SSC, allowing the buffer to slowly fill the chamber through capillary action. With either route, avoid generating bubbles or air pockets.
Once the tissue is covered with buffer inside the flow cell, use a clean laboratory wipe to carefully wick away excess buffer from around the flow cell ports without contacting the port.
Place the prepared flow cells into a light-protected slide staining chamber until ready to load onto the CosMx SMI Instrument. Record the flow cell ID, the six-digit number located on the flow cell coverslip, for each slide.
Run Configuration & Loading
Complete the following preparation steps before proceeding:
Ensure the CosMx RNA imaging tray has equilibrated to room temperature for at least 1 hour.
Retrieve RNase Inhibitor from -20°C. On the CosMx RNA imaging tray, pre-puncture the foil over the indicated well (A11, marked with a ⊕ symbol) with a clean pipette tip. Using a new pipette tip, add 4 µL (2-slide runs) or 8 µL (4-slide runs) of CosMx RNase inhibitor (NanoString Technologies 121500004 / 121500005) to the bottom of this well.
Log in to the CosMx SMI Control Center using valid Okta credentials.
Select "New Acquisition" to set up a new run.
Complete the following steps for each flow cell:
From the flow cell configuration screen, select "Create Flow Cell".
Select a pre-bleaching profile to quench tissue autofluorescence. The UMN-TMCs have used the following pre-bleaching profile(s) on the indicated tissues:
| Tissue type | Pre-Bleaching Profile | |
| Human Liver (FFPE) | Configuration B | |
| Mouse Liver (FFPE) | Configuration B | |
| Mouse Brain (FFPE) | Configuration B |
Select a cell segmentation profile. The UMN-TMCs have used the following cell segmentation profile(s) on the indicated tissues:
| Tissue type | Cell Segmentation Profile | |
| Human Liver (FFPE) | Configuration E | |
| Mouse Liver (FFPE) | Configuration E | |
| Mouse Brain (FFPE) | Configuration B |
Using the drop-down lists, select the cell segmentation kit, supplemental marker kit, and the a la carte marker (if applicable). Note: it is not recommended to change the default exposure times for image channels. The UMN-TMCs have used the following default exposure times on the indicated tissues and morphology markers:
| Tissue type | Ch1 (DAPI) | Ch2 (CD298/B2M) | Ch3 (PanCK) | Ch4 (CD45) | Ch5 (Empty) | |
| Human Liver (FFPE) | 24 ms | 100 ms | 33 ms | 50 ms | N/A |
| Tissue type | Ch1 (DAPI) | Ch2 (CD298/B2M) | Ch3 (PanCK) | Ch4 (CD45) | Ch5 (CD68) | |
| Mouse Liver (FFPE) | 24 ms | 100 ms | 33 ms | 50 ms | 25 ms |
| Tissue type | Ch1 (DAPI) | Ch2 (rRNA) | Ch3 (Histone) | Ch4 (GFAP) | Ch5 (Emtpy) | |
| Mouse Brain (FFPE) | 24 ms | 100 ms | 33 ms | 50 ms | N/A |
Select the RNA panel(s) used in the reagent configuration drop-down list.
Fill in the tissue information with section number, position in block, and section thickness.
After all fields have been completed, select "Save" to return to the flow cell configuration screen. Repeat each step for all flow cells.
After all flow cell records have been made, select "Replace Reagent Tray" to continue. Verify all flow cell information is complete and select "Proceed".
When prompted, open the imaging tray bay door and remove the used blue cleaning tray.
Carefully load in the new imaging tray, label-side first, until fully in the imaging tray bay. Ensure proper alignment, then close the imaging tray bay door.
When prompted, open the upper door of the instrument and complete the following steps:
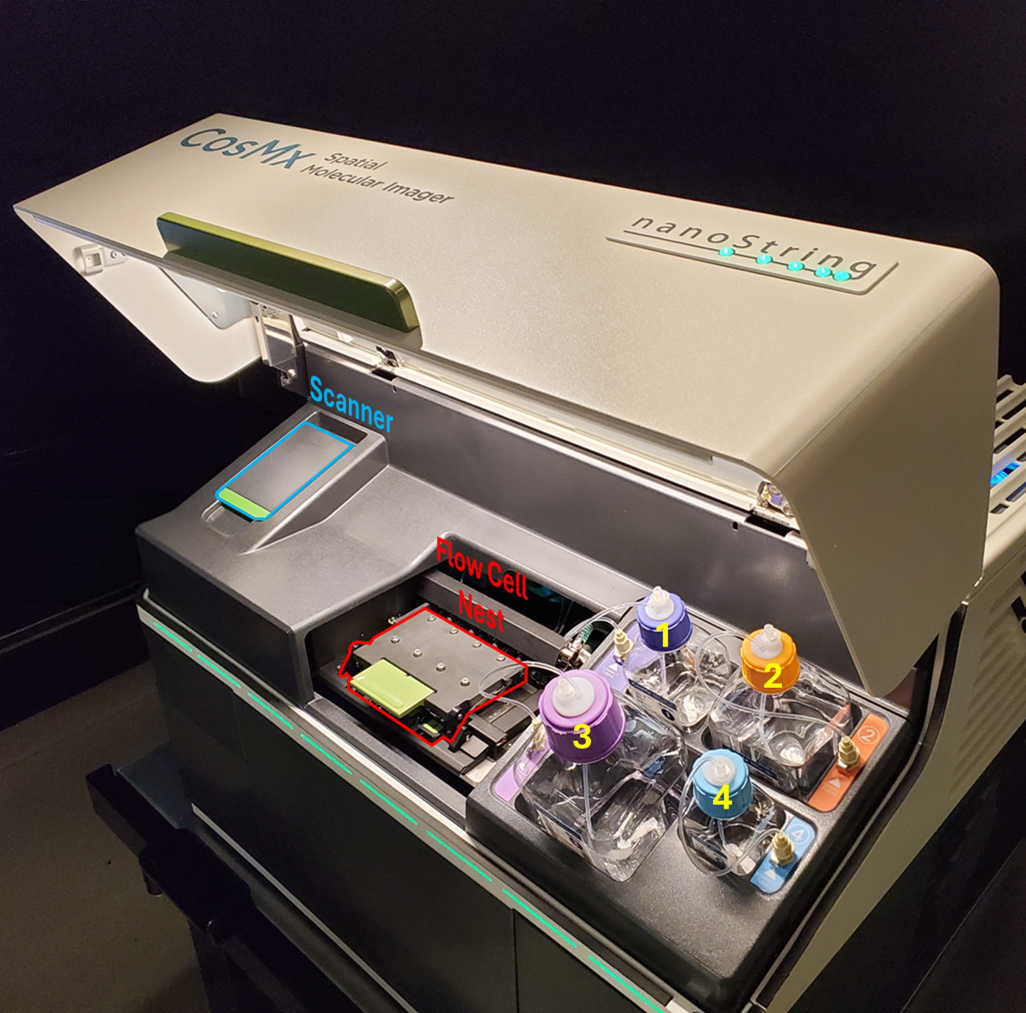

The upper door of the CosMx instrument must be opened to access the barcode scanner bay (blue), the flow cell nest (red), and the instrument bulk reagents (1-4, yellow).
Open the flow cell nest lid. Remove and discard or store any used flow cells.
When prompted, open the scanner bay door on the left side of the instrument deck. One slide at a time, scan the flow cell barcode and place tissue-side down into any available slot in the flow cell nest. Ensure fluidics gaskets are properly aligned to flow cell reagent ports. Repeat for all flow cells in the run until all are loaded, then close the flow cell nest lid and scanner bay door.
When prompted, remove all used bulk reagent bottles (1, 2, 3, and 4) by disconnecting the cap fittings at the side attached to the bay. Exchange the caps on fresh bulk reagent bottles with the corresponding fluidics cap fittings. Ensure enzymes have been added to bottle 4. Place all fresh bottles with the label facing forward into their appropriate bay and reconnect the cap fitting to the instrument. After all 4 bottles have been exchanged correctly, the on-screen bottle status cards will show as ready.
Following on-screen prompts, ensure all interior lids and doors are closed, then close the upper door.
Confirm the run parameters and select "Empty and Replace Waste", followed by "Proceed" when asked to mark flow cells as used. When prompted, disconnect the waste container cap fitting from the instrument and remove the container from the bay to discard waste. Replace the waste container and reconnect the cap fitting to the instrument. Allow the system to verify connections and waste container status and the instrument will automatically begin deck validation.
If prompted after deck validation, verify and approve the scan area to acquire the preview scan. This step may be omitted from newer software versions.
After the preview scan has been acquired, you may adjust the channel settings (e.g. contrast limits) of resulting morphology marker stains to guide placement of 0.5 mm by 0.5 mm square fields of view (FOVs) where data will be acquired. Instrument run time scales with the number of FOVs placed between all slides in a run. It is recommended to limit the number of FOVs to keep imaging time under 14 days.
Once all FOVs have been placed and approved across all flow cells, select "Begin Cycling".
Post-Run Cleaning
Within 48 hours of cycling completion, follow the on-screen prompts to open the imaging tray bay door and remove the imaging tray.
Insert a new blue pre-filled cleaning tray (NanoString Technologies #122000152) and close the imaging tray bay door. Cleaning should begin automatically
When the cleaning cycle is finished (~1 hour), follow the on-screen prompts to empty and replace the waste container as instructed.
Follow the on-screen prompts to remove used flow cells.
Data Export & Processing
Initial image processing, target decoding, and segmentation are completed automatically within the AtoMx Spatial Informatics Platform (SIP, v1.3.2) based on run configuration parameters indicated in the previous section.
Flat and raw data files are exported to an AWS S3 bucket using the export module after AtoMx study creation.
OME-TIFF images are generated for each field of view from the five-channel Morphology2D *.TIFF images using the bfconvert command in Bio-Formats (v 8.1.1). 4-level pyramids (tile size = 512 by 512 pixels) were created with 2x tiered down-sampling and lossless LZW compression.
Protocol references
Bruker Spatial Biology, Inc. (2025). CosMx SMI Methods Template [Digital File].
https://university.nanostring.com/cosmx-smi-methods-template
Melissa Linkert, Curtis T. Rueden, Chris Allan, Jean-Marie Burel, Will Moore, Andrew Patterson, Brian Loranger, Josh Moore, Carlos Neves, Donald MacDonald, Aleksandra Tarkowska, Caitlin Sticco, Emma Hill, Mike Rossner, Kevin W. Eliceiri, and Jason R. Swedlow (2010). Metadata matters: access to image data in the real world. The Journal of Cell Biology 189(5), 777-782. doi: 10.1083/jcb.201004104
