Jan 05, 2026
Tomogram reconstruction and pre-processing using standardized workflow in Scipion-EM
- Benoit Gallet1,
- Guy Schoehn1
- 1Univ. Grenoble Alpes, CEA, CNRS, IBS, F-38000 Grenoble

Protocol Citation: Benoit Gallet, Guy Schoehn 2026. Tomogram reconstruction and pre-processing using standardized workflow in Scipion-EM. protocols.io https://dx.doi.org/10.17504/protocols.io.36wgqpo45vk5/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: September 11, 2025
Last Modified: January 05, 2026
Protocol Integer ID: 227018
Keywords: Cryo-electron tomography, scipion-em, image processing, segmentation, Cryo-tomogram, GPU processing, traceable workflows for electron microscopy, modular workflow for tomogram reconstruction, electron microscopy, em electron tomography, tomogram reconstruction, microscopy, architecture of biological specimen, standardized workflow in scipion, nanometer resolution, source image processing platform, biological specimen, cell biology, subcellular structure, traceable workflow, specimen, implementing standardized pipeline
Disclaimer
DISCLAIMER – FOR INFORMATIONAL PURPOSES ONLY; USE AT YOUR OWN RISK
The protocol content here is for informational purposes only and does not constitute legal, medical, clinical, or safety advice, or otherwise; content added to protocols.io is not peer reviewed and may not have undergone a formal approval of any kind. Information presented in this protocol should not substitute for independent professional judgment, advice, diagnosis, or treatment. Any action you take or refrain from taking using or relying upon the information presented here is strictly at your own risk. You agree that neither the Company nor any of the authors, contributors, administrators, or anyone else associated with protocols.io, can be held responsible for your use of the information contained in or linked to this protocol or any of our Sites/Apps and Services.
Abstract
Electron tomography (ET) is a powerful technique for determining the three-dimensional (3D) architecture of biological specimens at nanometer resolution. It is widely used in structural biology and cell biology to analyze subcellular structures in near-native conditions. However, tomogram reconstruction is a complex, multi-step process involving data pre-processing, alignment, reconstruction, and post-processing. The diversity of tools and the flexibility of workflows can lead to inconsistent results, making reproducibility and traceability a significant challenge. To address this, we present a standardized and modular workflow for tomogram reconstruction using the Scipion-EM framework. Scipion-EM is an open-source image processing platform that integrates multiple software packages within a unified environment, enabling reproducible and traceable workflows for electron microscopy (EM) data processing. The platform supports automation, provenance tracking, and user-friendly graphical interfaces, making it particularly suitable for implementing standardized pipelines.
Guidelines
This protocol outlines a standardized tomogram reconstruction workflow within Scipion-EM, with an emphasis on:
- Reproducibility and traceability of results;
- Interoperability between software packages (e.g., IMOD, Dynamo, RELION, Warp/M, etc.);
- Flexibility for customization based on project-specific requirements;
- Best practices for data management and metadata handling.
The goal is to enable both novice and experienced users to execute robust tomographic reconstructions, reduce methodological variability, and facilitate the sharing and publication of processing pipelines.
This protocol is intended for:
- Researchers and technicians engaged in electron tomography data processing;
- Cryo-EM core facility personnel developing training resources;
- Developers interested in expanding or benchmarking workflows within Scipion-EM;
- Collaborators seeking to replicate or audit published results using standardized methods.
While some knowledge with tomographic concepts and EM software is prerequised, this guide is designed to be accessible to users new to Scipion-EM.
Materials
Software/tools referenced on these pages: Scipion-EM GUI, AreTomo (ProtAreTomoAlignRecon), IMOD/3dmod (tomogram viewing, dose filtering and apply transformation), MotionCorr/ProtTsMotionCorr, CistemProtTsCtffind (CTF estimation), CTFEstimationTomoViewer, ProtImodDoseFilter (IMOD - Dose filter), ProtImodApplyTransformationMatrix (IMOD - Apply transformation), ProtImodTomoReconstruction (IMOD - Tomo reconstruction), CryoCARE (ProtCryoCARETraining and ProtCryoCAREPrediction/CryoCARE Prediction), Warp (ProtWarpDeconvTomo - warp deconvolve tomograms), tomo3D (ProtTomo3dProtDenoiseTomogram - denoise tomogram, Edge Enhancing Diffusion (EED) option), MemBrain (ProtMemBrainSeg - membrain - tomogram membrane segmentation), ChimeraX (3D rendering).
Hardware Requirements
While Scipion-EM is platform-independent, the following hardware is recommended for efficient tomographic reconstruction:
- CPU: Multi-core processor (8 cores or more recommended)
- GPU: CUDA-compatible GPU (for GPU-accelerated tools like MotionCor2, AreTomo, Membrain-seg, CryoCARE or RELION)
- RAM: Minimum 64 GB or more preferred for large datasets
- Storage: SSD or high-speed disk with sufficient capacity for intermediate files and reconstructed tomograms (8Tb minimum) and SATA discs for storage (100Tb is a good start)
Troubleshooting
Safety warnings
Ensure the following are installed and properly configured:
Scipion-EM (v3.0 or higher) with tomography-related plugins:
- scipion-em-xmipp
- scipion-em-xmipptomo
- scipion-em-imod
- scipion-em-motioncor
- scipion-em-aretomo
- scipion-em-relion (for CTF estimation or subtomogram averaging)
- scipion-em-dynamo (for subtomogram averaging workflows)
- scipion-em-tomo
- scipion-em-warp
- scipion-em-tomo3d
- scipion-em-membrain
- scipion-em-cryocare
Python: Included with Scipion; no separate installation needed
CUDA Toolkit (if using GPU-accelerated tools)
Environment variables set appropriately (e.g., for IMOD’s IMOD_DIR, CUDA paths)
Before start
To ensure consistency and reproducibility:
- Organize your data: Maintain a consistent folder structure (e.g., one folder per tilt-series with raw data and metadata)
- Backup raw data: Before starting any processing, create a read-only copy of your original files
- Version control: Record software versions and parameters, especially if you are working in a collaborative environment
- Check plugin updates: Scipion plugins evolve rapidly; always check for the latest stable versions with bug fixes or new features
- GPU vs CPU paths: Be aware of which steps benefit from GPU acceleration; plan your workflow accordingly if using a shared cluster
Organize your data in the following:
i. Tilt-Series Data
- Format: MRC, TIFF, or other supported image stacks
- Acquisition Metadata: Must include tilt angles, pixel size, defocus (if applicable), and microscope parameters
- Fiducials: Gold fiducials (10–20 nm) are recommended for robust alignment unless using fiducial-less methods
ii. Microscope and Acquisition Parameters (in .mdoc file)
- Voltage (e.g., 300 kV)
- Spherical aberration (Cs, 2.7 for Krios)
- Detector type (e.g., K3, Falcon 4)
- Exposure time and dose per tilt (Exposure dose)
- Acquisition scheme (dose-symmetric, bidirectional, etc.)
- Number of frames per movie
- Tilt angle (~85°)
- Pixel size (Å/pix2)
Accurate metadata is critical for CTF estimation, tilt alignment, and dose compensation steps, all these informations are available in the .mdoc file.
Start Scipion
Go to Document and create a new folder with the name of your experiment, copy your raw data inside (.mrc or .tiff/.eer AND .mdoc files).n the same folder, in which the movies are, you need the .gain file of the day of data collection.
Start scipion3 in a terminal.
Create new project
Create new project and give the same name that your data folder
On the left panel select the tomography tree for availability of adapted protocols
Workflow import and setup
Go to import workflow
Go to ScipionUserData/Worflows_template
Select the Workflow adapted to your project and data and import it
NB: you can always start a blank workflow and add each step as a new protocol from the list.
Import data
What to do before: open a .mdoc file and identify the mandatory information: kV, Cs, pixel size, movies frames, electron dose.
In the case of .mrc stack files
Protocol to use: tomo - import tilt-series
You have to specify:
- Folder with your data
- Pattern of the .mdoc file containing information of the data (*.mdoc or Position*.mdoc or TS*.doc)
In the case of individual tilt movies
Protocol to use: tomo - import tilt-series movies
Correct motion
In the case of .eer or .tiff movies of individual tilt, motion needs to be corrected using motion correction.
Protocol to use: motioncorr - align tilt-series movies
NB: In the case of using CryoCARE afterwards, it is mandatory to split odd/even frames!
Open the results:
Align tilt series and evaluate the quality of the data
AreTomo, IMOD patch, IMOD fiducials
Select the imported data or the motion corrected data
- Skip alignment: No
- Reconstruct the tomogram: Yes
- Reconstruct Odd/Even: No
NB: In the case of the use of CryoCARE later on, you need to always activate the Odd/Even option, in all the steps.
- Binning: 8
- Volume height for alignment: try between 200 and 400
- Tomogram thickness unbinned : 1200
- Refine tilt angle: No
- Refine tilt axis angle: Refine and use the refine value for the entire “stack”
- Generate extra IMOD output: No
- Estimate the CTF: Yes
- Do phase shift estimation: No
Select “Extra Options” Tab:
Select the SIRT method to have more contrast
- You can add the Patch tracking by selecting the option :
- Local motion correction - Well distributed 5 5
- Additional parameter: TiltCor 1
Check the result: (open output: Tomograms)
Open each, one by one, by double clicking on it: (you may need to transfer the files locally).
Check the tomogram and if the alignment is good, keep those parameters, otherwise restart this step trying to optimize it: increase/decrease the volume for alignment, use Patch (Local motion correction),...
Estimate CTF
Plug-in to use: cistem - tilt-series ctffind
Open result with CTFEstimationTomoViewer:
You can remove the bad tilt-series, or at least the bad images
Dose weighting
Protocol to use: Imod - Dose filtering
If the dose was correctly set in the import step you can select the Scipion import as Input dose source. Otherwise you can set the dose as fixed and enter the dose for each image (tilt) in electron / angstrom².
In the case of reconstructing the even and odd tomograms (for the use of CryoCARE denoising) you can apply this dose filtering to the halves, or not.
Tomogram reconstruction
Alignement
Binning 2, 4, 8 and purpose
The recommendations for the binning are the following:
- Start with Binning 8 because it’s faster, give more contrast, is useful for quality check of the data and the alignments/reconstructions parameters
- Then do Binning 4 and Binning 2 (and Bin1 for highest resolution reconstruction)
- Bin 4 for denoising and segmentation of membranes, for morphological studies
- Bin 2 for high resolution interpretation of the data, particle picking and STA (Sub-Tomogram Averaging)
Binning effects on reconstruction contrast and resolution
Binning effects on reconstruction contrast and resolution
Reconstruction
Plung-in: mod - Tomo reconstruction
Denoising
We propose three different ways for denoising: CryoCARE, Warp and tomo3D.
1) CryoCARE is a training-based approach, it is very long for the training but the result is of very high quality.
2) Warp is a CTF based filtering approach that is very fast and gives very good results
3) tomo3D is more a filtering method that can use either Anisotropic Nonlinear Diffusion (EED method) or TomoBFlow, a nonlinear filtering method, it is also a fast method for denoising.
To be able to use CryoCARE you need to have even and odd reconstructions of your tomograms. This is done at the “motion correction” step level and is also in option in all the subsequent steps by always selecting “Apply to odd/even”.
1- Training:
Protocol to use: cryocare - CryoCARE training
2- Predicition:
In case you want to denoise a full set of tomograms from a single data collection session, you can train the CryoCARE NN with one representative tomogram and then use the trained NN to predict all the other tomograms.
3- Results:
CryoCARE denoising
Warp
Protocol to use : warp - deconvolve tomograms
Input: the reconstructed tomogram in Bin4 (or Bin8)
CTF tomo series: the CTF estimation from AreTomo or gctf
Plug-in: warp - deconvolve tomogram
Results:
Warp denoising
Protocol to use: tomo3d - denoise tomogram
If you want to explore the best settings you can play either with the Denoising method or with the parameters.
Results:
tomo3d denoising
Segmentation
Membrain-seg
Protocol to use : membrain - tomogram membrane segmentation
Membrain is an automated method to segment membranes in denoised cryo-tomograms. It uses a neural network that has been trained on Chlamydomonas data mainly. The result will be a .mrc file with the segmented membranes, it can be grey scaled according to connected components
"Input" tab:
Select the data: denoised tomograms (provide denoised tomograms as input in the Input tab of the ProtMemBrainSeg protocol).
“Connected components analysis” tab:
You can either choose to label connected component, or not, depending on the software you will use in a second step.
”Test-time augmentation” tab: always “Yes”
Open the result to check, with 3dmod or with chimera (as volume)
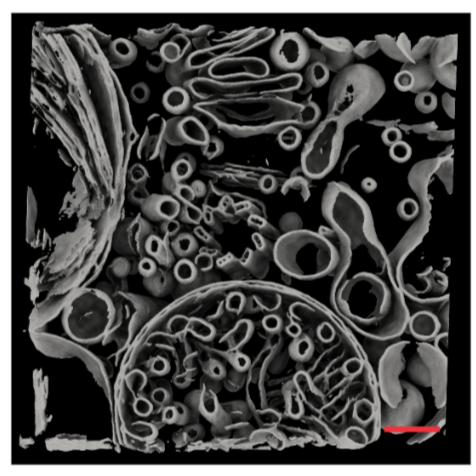
Results of the segmentation
Downstream processes
Particle picking
Sub-Tomogram Averaging
Acknowledgements
This work used the platforms of the Grenoble Instruct-ERIC center (ISBG ; UAR 3518 CNRS-CEA-UGA-EMBL) within the Grenoble Partnership for Structural Biology (PSB), supported by FRISBI (ANR-10-INBS-0005-02) and GRAL, financed within the University Grenoble Alpes graduate school (Ecoles Universitaires de Recherche) CBH-EUR-GS (ANR-17-EURE-0003).
