Oct 19, 2021
Version 2
Titan Illumina PE SARS-CoV-2 Strain Characterization Workflow for the Terra Platform V.2
This protocol is a draft, published without a DOI.
- Jill V Hagey1,
- Kevin Libuit2,
- Frank J Ambrosio2,
- Technical Outreach and Assistance for States Team1
- 1Centers for Disease Control and Prevention;
- 2Theiagen Genomics
- TOAST_publicTech. support email: [email protected]

Protocol Citation: Jill V Hagey, Kevin Libuit, Frank J Ambrosio, Technical Outreach and Assistance for States Team 2021. Titan Illumina PE SARS-CoV-2 Strain Characterization Workflow for the Terra Platform. protocols.io https://protocols.io/view/titan-illumina-pe-sars-cov-2-strain-characterizati-by77pzrnVersion created by Technical Outreach and Assistance for States Team
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: In development
We are still developing and optimizing this protocol
Created: October 19, 2021
Last Modified: January 20, 2022
Protocol Integer ID: 54239
Keywords: Illumina, Sequencing, MiSeq, iSeq, MiniSeq, NextSeq, NovaSeq, Paired-End, Next Generation Sequencing, NGS, SARS-CoV-2, Covid, Pangolin, Short-Read, Coronavirus, Genomics, Genetics, Virology, Molecular Biology, public health viral genomics titan series, input data for titan illumina pe, additional technical documentation for the titan-illumina-pe workflow, titan-illumina-pe data, consensus genome assembly, primer sequence coordinates of the pcr scheme, titan-illumina-pe workflow, theiagen genomic, generated consensus genome, bed format along with the raw illumina, consensus genome, genomic, titan illumina pe, bed file format, primer sequence coordinate, artic v3 protocol, pcr scheme, clade designation, titan illumina pe sar, quality of both raw read data, terra platform, titan-illumina-pe, titan-illumina-pe job, qiaseq primer panel, bed format, clade, pcr amplicon, common read data, input primer scheme coordinate, artic v3, raw read data, pcr, fastq files per sample
Disclaimer
The opinions expressed here do not necessarily reflect the opinions of the Centers for Disease Control and Prevention or the institutions with which the authors are affiliated. The protocol content here is under development and is for informational purposes only and does not constitute legal, medical, clinical, or safety advice, or otherwise; content added to protocols.io is not peer reviewed and may not have undergone a formal approval of any kind. Information presented in this protocol should not substitute for independent professional judgment, advice, diagnosis, or treatment. Any action you take or refrain from taking using or relying upon the information presented here is strictly at your own risk. You agree that neither the Company nor any of the authors, contributors, administrators, or anyone else associated with protocols.io, can be held responsible for your use of the information contained in or linked to this protocol or any of our Sites/Apps and Services.
Abstract
The Titan_Illumina_PE workflow is a part of the Public Health Viral Genomics Titan series for SARS-CoV-2 genomic characterization. Titan_Illumina_PE was written specifically to process Illumina paired-end (PE) read data. Input reads are assumed to be the product of sequencing tiled PCR-amplicons designed for the SARS-CoV-2 genome. The most common read data analyzed by the Titan_Illumina_PE workflow are generated with the ARTIC V3 protocol. However, alternative primer schemes such as the QIAseq Primer Panel are also suitable for this workflow. The primer sequence coordinates of the PCR scheme utilized must be provided in BED format along with the raw Illumina read data. Upon initiating a Titan_Illumina_PE job, the input primer scheme coordinates and raw paired-end Illumina read data provided for each sample will be processed to perform consensus genome assembly, infer the quality of both raw read data and the generated consensus genome, and assign lineage or clade designations as outlined in the Titan_Illumina_PE data workflow diagram below.
Additional technical documentation for the Titan_Illumina_PE workflow is available at:
Required input data for Titan Illumina PE:
Illumina paired-end read data (forward and reverse FASTQ files per sample)
Primer sequence coordinates of the PCR scheme utilized in BED file format
Video Instruction:
Theiagen Genomics: Titan Genomic Characterization
Theiagen Genomics: Titan Outputs QC
For technical assistance please contact us at: [email protected]
Attachments
Troubleshooting
Setup Terra and Google Cloud Accounts
Note
The Terra platform registration requires a Google account. If you have a Google account, you can sign in using the Terra login page:
Click on the three parallel lines in the top left-hand corner and click the 'Sign in with Google' button.
If you do not have Google email, you can set up a Google account with a non-Google email. The steps to do this are described in the following link:
Note
The Terra platform uses the Google Cloud to run workflows and store data. The following documentation will describe how to set up a Google Cloud account:
Note
To link your Terra platform account with your Google Cloud account, follow the instructions provided here:
Scroll down to Section 3 titled "Create a Terra Billing Project" and follow the instructions. It is important to note that the name you give to your Terra Billing Project must be unique across all Google Billing Projects. If the name provided is not unique, it won't immediately throw an error, but instead will not complete the process of associating the Google Billing account with the Terra Billing Project. If this occurs cancel the process and set up a new Terra Billing Project.
Import Titan Illumina PE workflow from Dockstore
Importing the Titan Workflow from Dockstore to the User Workspace
We will first walk through step by step how to import the Titan workflow and at the end there is a video showing the full process.
First create a new workspace by clicking on the three parallel lines in the top left-hand corner, followed by clicking the 'Workspaces' tab then click the "+" sign on the top right of the newly loaded page. Follow the directions to link the workspace to your billing project. The video at the end shows all steps involved. DO NOT INCLUDE SPACES IN THE WORKSPACE NAME, use underscores instead. Spaces will cause an error later in the pipeline.
Creating a new Terra workspace
The script for the Titan workflow is located in the Dockstore repository and has to be imported into the user Terra Workspace. There are two ways to get to this page and the video shows both. The first method starts by clicking on the three parallel lines in the top left-hand corner, followed by clicking the 'Library' tab and finally click the 'Workflows' button.
The library tab on the Terra Platform.
In the 'Workflows' panel, under 'Find Additional Workflows' click on the 'Dockstore' link in the grey box on the right side of the page.
The 'Code and Workflows' page in the 'Library' panel.
On the left side of the Dockstore page, search for 'theiagen/public_health_viral_genomics/Titan_Illumina_PE' in the search bar.
Search results for 'theiagen/public_health_viral_genomics/Titan_Illumina_PE' on the Dockstore page.
Click the 'theiagen/public_health_viral_genomics/Titan_Illumina_PE' link, which will take you to a new page.
The Theiagen Titan Illumina PE workflow page on the Dockstore.
On the right-hand side of the page under the 'Launch with' window, click the 'Terra' button. It should bring you back to the Terra platform within the 'Import Workflow' page. If you already have a workspace created you can simply import it into that workspace. If not, see step 2.1 to create a new workspace.
The Terra 'Import Workflow' page. The Titan Illumina PE workflow code is shown on the left.
After clicking the 'import' button, you should be automatically directed to the Titan workflow panel in the new workspace page that was just created.
Make sure you pick the latest version (or an older stable version you want to use) from the drop down version menu before continuing. The "main" and "dev" versions are under active development and aren't stable!
The Titan workflow panel within the newly created Terra workspace
Video of the entire process:
Alternatively, if you have not already made a workspace, you will need to before importing the workflow. To do this, under 'Destination Workspace' click the 'create a new workspace' button. A pop-up window titled 'Create a New Workspace' should appear. Name your new workspace and associate it with a billing account using the 'Billing project' drop-down menu. The Terra Billing Account you created in the previous step should be available. Finally, click the 'Create Workspace' button.
Creating a new workspace in the Terra 'Import Workflow' page
After clicking the 'Create Workspace' button, you should be automatically directed to the Titan workflow panel in the new workspace page that was just created.
Make sure you pick the latest version (or an older stable version you want to use) from the drop-down version menu before continuing. The "main" and "dev" versions are under active development and aren't stable!
The Titan workflow panel within the newly created Terra workspace
Upload Fastq Sequence Files
There are three options for uploading your files:
1. Upload on Terra (can only do single files).
2. Upload to Google bucket and link to Terra (can do single files or bulk files/folders).
Upload fastq sequence files one at a time to be used in the analysis.
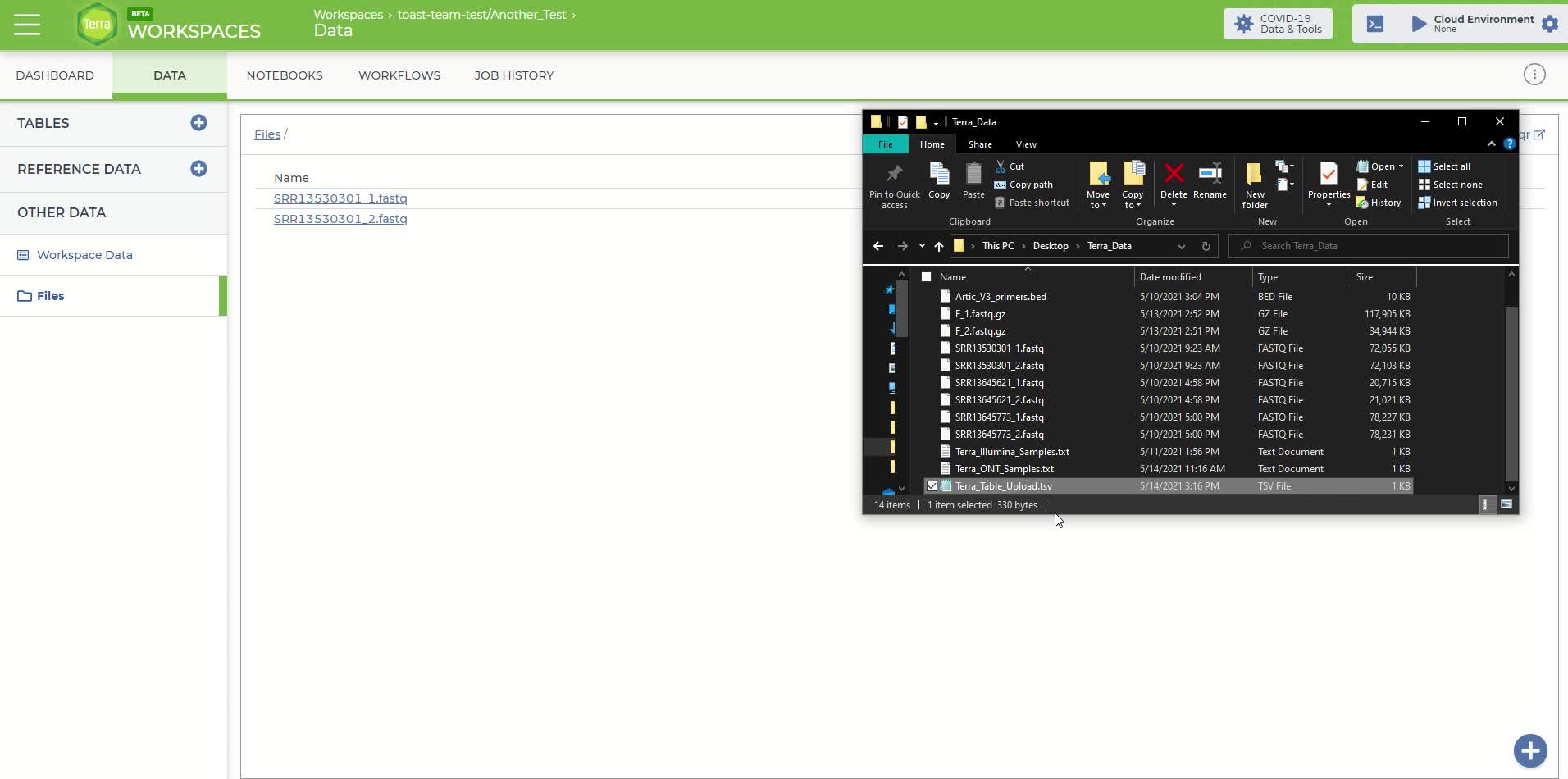
Click on the 'Data' panel in the newly created workspace and then click on the 'Files' tab
The 'Files' tab within the 'Data' panel of the newly created workspace
Under the 'Files' tab, you can either drag-and-drop your files into this space or move the mouse over the blue plus sign icon in the bottom right-hand corner and click 'upload'. Upload the sequence files you'll need for this analysis. Each fastq sequence file must be uploaded individually using this method.
Once the files are uploaded, you will need to create a table to associate your files with their corresponding link to their Google Cloud location.
Note
The Terra sample table file must follow a specific template. We've provided the template file here  Terra_Table_Upload.tsv as a downloadable tab-separated (or .tsv) file. The tab-separated table has three columns: entity:sample_id, Forward_Read, and Reverse_Read.
Terra_Table_Upload.tsv as a downloadable tab-separated (or .tsv) file. The tab-separated table has three columns: entity:sample_id, Forward_Read, and Reverse_Read.
Either by editing the text file or using spreadsheet software like Excel, fill in each column with the required information. The first column 'entity:sample_id' is the sample name that is provided by the user. The second and third columns, Forward_Read, and Reverse_Read are the file paths where the paired-end fastq files are stored within the Google Cloud.
While you had to upload your samples individually, all your samples can be organized in one data table.
To identify the Google Cloud location, right-click on each fastq file that was uploaded in the previous step and copy the link address. The file path should look like something similar to the following:
gs://fc-b1e3191a-3d9f-43fe-9743-255551ce2f38/SRR11953697_1.fastq.gz
When completed the table should look similar to the following:
Example Terra sample file
Once the table has been completed with the required information, save it in tab-separated (or tsv) format. The spreadsheet software should have an option to save as a 'tsv' file.
Note: When uploading additional sample table files to the same workspace the entity types must be unique and end in "_id" (e.g. sample1_id, sample2_id etc.)
Finally, the completed Terra sample file will need to be uploaded to the newly created Titan workspace. On the right-hand side of the workspace 'Data' panel there is a 'Tables' tab. Click the blue plus sign icon on the right edge of the 'Tables' tab. A popup window should appear titled 'Import Table Data'. Select your completed tab-separated sample file for upload and then click the 'Upload' button.
The 'Import Table Data' window for uploading the Terra sample file
If the upload is successful then the sample file should be located under the 'Tables' tab as 'sample (#)' where # is the number of samples in your file.
The workspace 'Data' panel after successfully uploading the fastq sequence files and Terra sample file
Here is a video showing the process.
You will likely have many samples to upload and you can do this by going directly to your Google bucket.
First, go to “DASHBOARD” tab in your workspace and click “Google Bucket” at the bottom right corner of the same page.
The Dashboard tab of your workspace.
This will direct you to your “Google Cloud Platform” page for data uploading. Click “UPLOAD FILES” in the middle of this page to upload single or multiple fastq files. Or you can click “UPLOAD FOLDER” to upload a folder with multiple fastq files stored inside. This will cost some google cloud credits (e.g.4.4 Gb for $0.75).
Google bucket page for uploading files and folders.
Go back to your Terra account and click “DATA” tab. The successfully uploaded fastq files will show up. If the upload is successful then the sample file should be located under the 'Tables' tab as 'sample (#)' where # is the number of samples in your file.
Recently uploaded data.
Here is a video of the process:
To upload via the Terra Data Uploader.
Select the Terra Workspace to which you would like to upload your fastq files. This will be the same workspace created in the previous step.
Click the '+ Create a new collection' link and enter a name for your new collection of fastq files. DO NOT INCLUDE SPACES IN THE COLLECTION NAME, use underscores instead. Spaces will cause an error later in the pipeline.
Note
Go to step 11 to see an example error caused by spaces present in the collection name. The process will also show you how to investigate a job failure.
Click the 'Create Collection' button and you will be routed to the data uploader page where you can drag and drop fastq files directly into your browser window to initiate upload.
Drag and drop the fastq files that you would like to upload in to the upload space.
Screen when files are uploading
Once your files have been successfully uploaded, select "NEXT>" to proceed to the metadata upload page.
To upload a metadata table and create a Terra sample table in your workspace, open excel and populate your spreadsheet with the root entity designation in cell A1 (here it is entity:sample_id), "Forward_Reads" in B1, and "Reverse_Reads" in C1 as headers; underneath these headers, proceed to populate each row.
Important note on column names: DO NOT USE SPACES! As we did before in creating our workspace and collection names use "_" instead of spaces! The first column MUST have the name "entity:sample_id", you can call the other columns whatever you like, but we recommend you use something that denotes forward and reverse reads for clarity.
Metadata table saved as a ".tsv" file.
An easy way to create this file is to run the following in the directory where your fastq files are stored:
Command
new command name
This will create the a file called Terra_Upload.txt for you. This assumes your files have the format string_1.fastq.gz and string_2.fastq.gz.
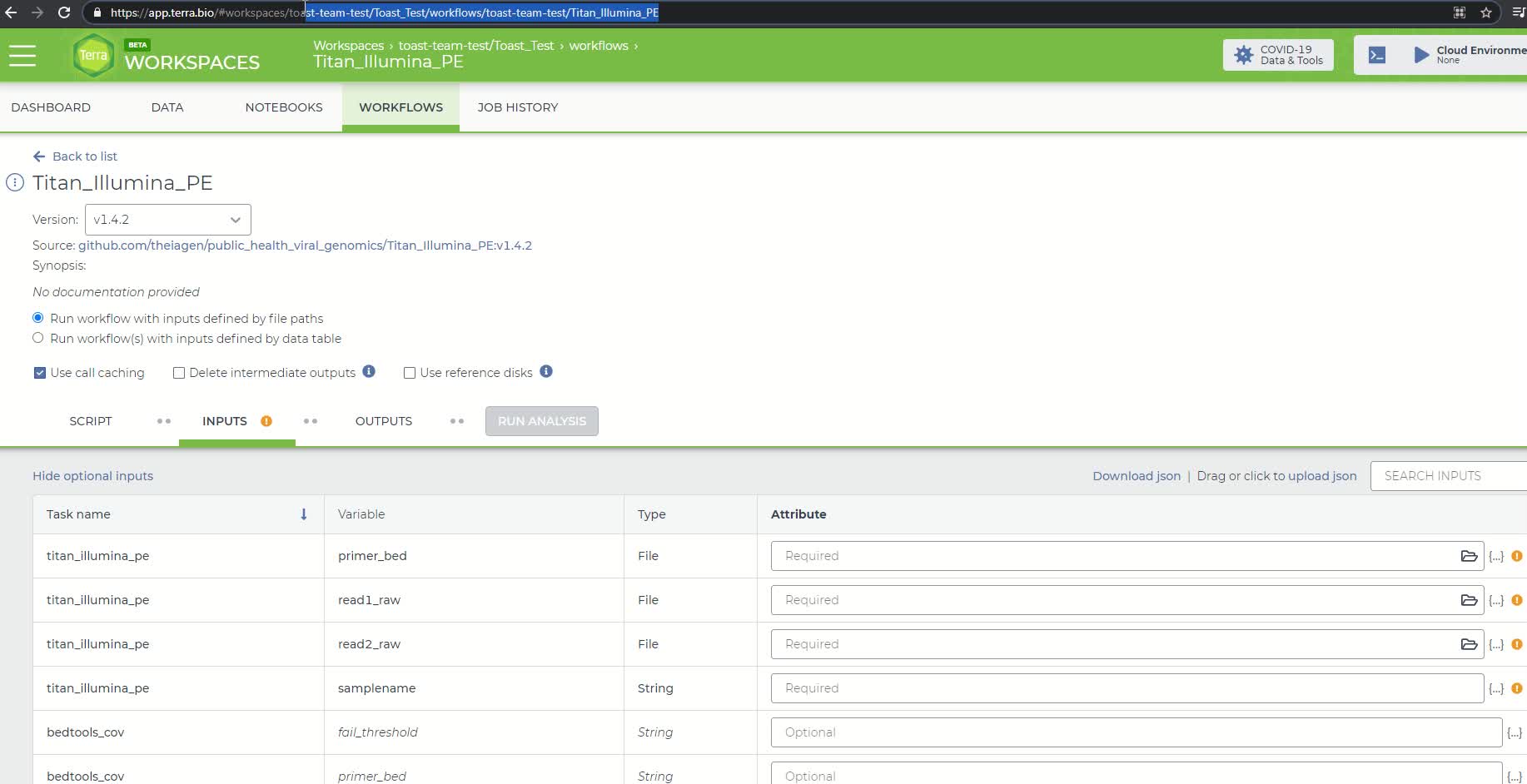
NOTE: In this example, our root entity type is "sample_id" so, in cell A1, we have written "entity:sample_id". A "root entity" is the smallest piece of data a workflow can use as input (blue square). We can check what is required from the workflow by looking at what fields are set to required (red rectangle). The root entity is always defined with "entity:" and must always end in "sample_id"
List of fields in the Titan Illumina PE workflow
Note
You can have as many columns as you want in addition to the "entity:sample_id", "Forward_Reads" and "Reverse_Reads" columns. Some examples you might want to include are:
- Ct value
- Sequencing run or plate ID
- Sampling or extraction dates
If you plan to use the Augur protocol or visualize your data with UShER or Auspice, consider including that metadata in the file you upload now. Required columns for Augur are described here:
and include:
- Collection_date
- iso_country
- iso_state
- iso_continent
- iso_county
Date columns, sequencing date for example, must use the format YYYY-MM-DD.
Whatever additional columns of metadata you populate to this Terra data table will, after a Titan run, gets populated to that same table as the final Titan output. You can then use this data to correlate assembly quality with CT values to determine cutoff thresholds for which samples to actually sequence in the future or identify issues that are sequence run specific.
If you plan to also use the Mercury workflow to prepare sequence and metadata for submission to public repositories (GISAID, GenBank and SRA) then populating this sheet now will make running the Mercury workflow easier.
Save this sheet as a tab delimited file text file.
Saving metadata file as a tab deliminated.
An example, metadata table is found below.
Note
This example file has the minimum amount of columns to be able to create a collection. Delete the sample names and populate the document with your own.
If you open this with Excel, a pop-up window will show up. Check the box "my data has headers" then click "next".
Step 1 in the pop up window
In step 2 make sure the "Tab" box is checked under "delimiters". Click next then click finish. Now you can edit!
Step 2 of the pop window
Drag-and-drop your metadata table to the Terra #Upload page and select CREATE TABLE to complete the process.
Clicking CREATE TABLE button to complete the process of making a collection.
Video of entire process:
Upload Primer Bed Files
For the Titan_lllumina_PE workflow v1.2.0, you will need to upload a bed file for the primers you used. The reasons for this and a video example can be found on YouTube. This will need to be done once and then you can use the same file for every run that uses that primer set. Visit the ARTIC Network's GitHub repository to download a bed file for the ARTIC primers.
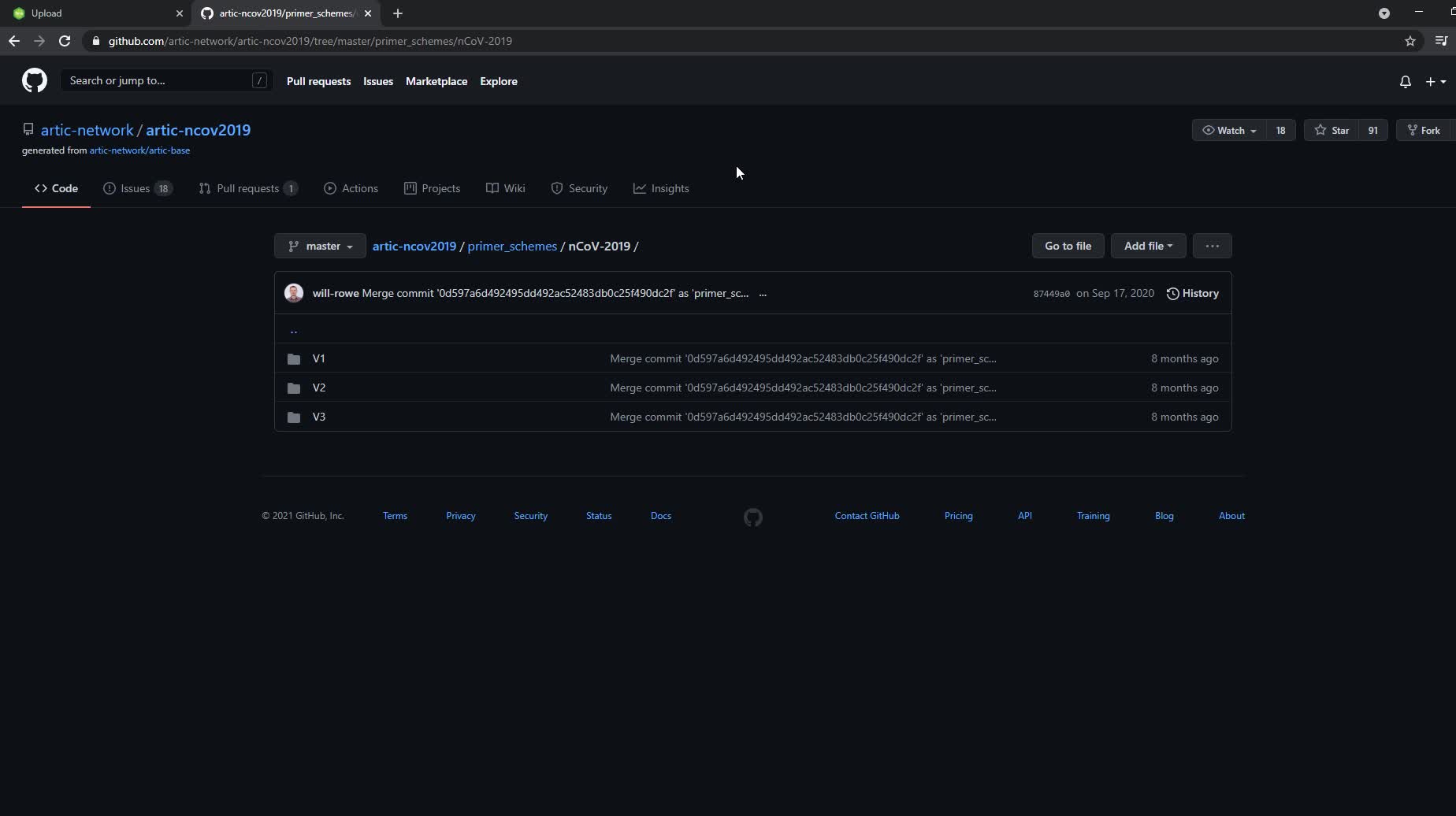
The ARTIC Network's Github Page with folders for different versions of the Artic primers.
Select the folder for the version of artic primers you are using. Then click on "nCoV-2019.primer.bed" and click the "Raw" button.
In the V3 folder, click on nCov-2019.primer.bed file.
Clicking the raw button to get the primer.bed file to save.
Copy and paste everything from this new page to a new notepad document. DO NOTE USE MICROSOFT WORD! Save as a text file with the primer version number in the filename. Then you will need to go into the folder and change the extension from ".txt" to ".bed". If necessary, click "View" at the top of the folder window (red circle) and ensure the "File name extensions" option (red rectangle) is selected.
Check the box to show file name extensions.
Then right click on metadata file and select "rename" then change ".txt" to ".bed".
Change file extension from ".txt" to ".bed"
Navigate to 'https://app.terra.bio/#upload'. Once on the upload page, click on the workspace you created before and create a new collection by clicking the blue "+" next to "select a collection" in the middle top of the page. Name the collection, here I will name it "Titan-Illumina-Files".
Create a new collection
Drag and drop the primer file you saved into this collection. The bed file will now show up as a hyperlink that actually links it from the Google bucket where it was actually uploaded to (recall Terra uses google cloud as a backend). Right click on this blue hyperlink and select "copy link address".
Copy the link address for the primer.bed file that was just uploaded
Now we will need to add the primer bed file as a data workspace element. Navigate back to your workspace and under the "Data" tab select "Workspace Data" from the menu on the right hand side. Click the "Add Variable" blue "+" in the bottom right hand corner. In the key field, name the file something like "artic-v3-nCov2019-primers" In the "value" field paste the link address to the google bucket you copied earlier. Save the file by clicking the check mark on the far right.
Add a new workspace element and save the file.
For the bed file to appear in your workspace, you must sign out and sign back in before continuing to the next step.
Here is a video of the entire process:
Running the Titan Illumina PE Workflow
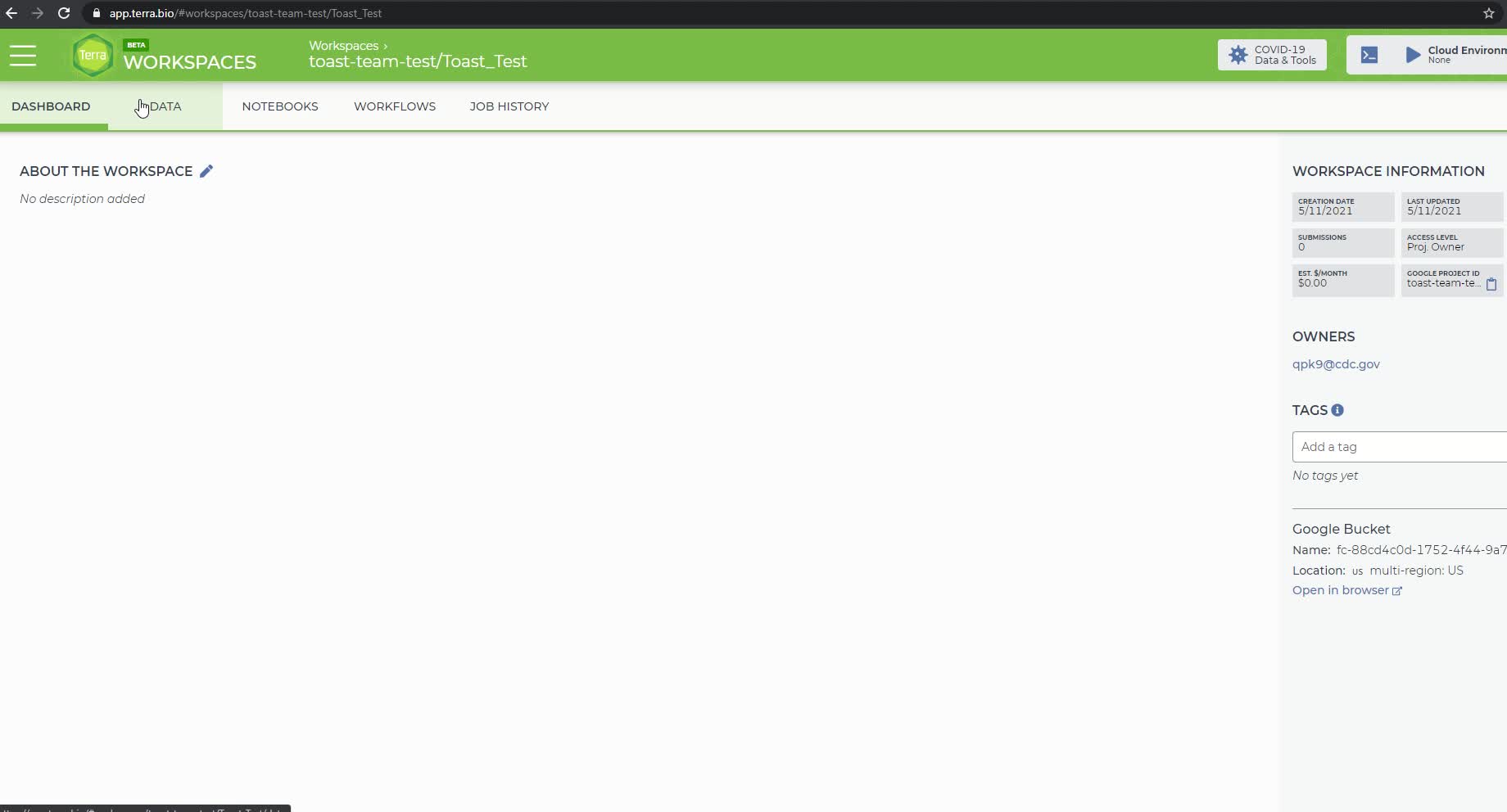
To run the Titan workflow, click on the 'Workflows' panel in the newly created workspace. It should bring you to the following page:
The workflows panel in the newly created Terra workspace
Click on the 'Titan_Illumina_PE' tile and it will take you to a new page. Double check that you are using the latest version of the workflow. Alternately, you may specify another version, but should only pick a stable version (with numbers) NOT a 'main' or 'dev' version.
The Titan Illumina PE workflow page. Check version you are using.
Click the 'Run workflow(s) with inputs defined by data table' option button and then in the 'Select root entity type' pull-down menu, select 'sample' or whichever entity type you specified in the sample table. The first four rows in the 'Inputs' tab should each have a yellow caution icon. These four rows represent variables that have to be provided by the user. This is the information that we populated the sample data table with in the previous steps.
Make sure "Use call caching" is checked and then select the root entity type for the data you wish to analyze (here it's sample).
NOTE: Call caching allows Terra to identify and skip jobs that have been run previously; this option is by default enabled to avoid unnecessary compute costs. More information on Terra call caching, including examples of when you may want to disable this feature, is available through the Terra Support Documentation.
The Titan Illumina PE workflow input panel after defining inputs by the sample data table
For the first row, click on the 'Attribute' text box and then click 'workspace.Artic_V3_nCov2019_primers' (or whatever you named the primer bed file). For the next row, the 'read1_raw' variable, click on the 'Attribute' text box and then click 'this.Forward_Read'. In the third row, the 'read2_raw' variable, click on the 'Attribute' text box and then click 'this.Reverse_Read'. Finally, in the third row, the 'samplename' variable, click on the 'Attribute' text box and then click 'this.sample_id'. Click the 'Save' button on the top right-hand side of the page. The yellow caution icons should disappear.
The Titan Illumina PE workflow input panel after setting the four required input variables
Note
VERSION UPDATE:
Beginning with v1.4.4, you have the option to specify either UShER or pangoLEARN methods for PANGO lineage assignment. The default is UShER unless "pangolearn" is entered into the "inference_engine" row.
Module 3.3: Real-time phylogenetics with UShER
Next, click the "output" tab and select "use defaults" to autofill in the names of the output files according to the sample names. If you forget to select this option, you will have trouble accessing the results! Save changes by clicking the 'Save' button.
Setting output names by clicking "use defaults"
You are now ready to run the Titan_Illumina_PE workflow! Click the blue 'select data' button and select the samples you want to run or click the check box at the very top left to select all of them to be run.
Check the box to have all samples run through the Titan_Illumina_PE workflow.
Click on the 'Run Analysis' button to the right of the 'Outputs' tab. A popup window should appear titled 'Confirm launch'. If the 'Run Analysis' button is greyed out, you need to save your recent changes by clicking the 'Save' button.
The 'Confirm launch' popup window
Click the 'Launch' button to access the 'Job History' panel where each sample will be queued for the Titan_Illumina_PE analysis.
The 'Job History' page showing each sample queued for the Titan_Illumina_PE run
The status will change from 'Queued' to 'Submitted' to 'Running.' After the workflow finishes, the status column will indicate either 'successful' or 'failed.'
Video of entire process:
This starts by loading the primer bed file, which we did in the last step, but here we show an alternative method to upload it to your workspace. After uploading, you will likely have to reload the page before it shows up as an option on the workflows page (as shown in the video).
View and Download the Titan Output Report
First, verify that all of the samples have completed the analysis run by looking at the 'Workflow Statuses' section in the top left of the 'Job History' panel. The job has completed when all the samples have a status of 'Succeeded' with a green checkmark.
The 'Job History' panel when the jobs have successfully been completed.
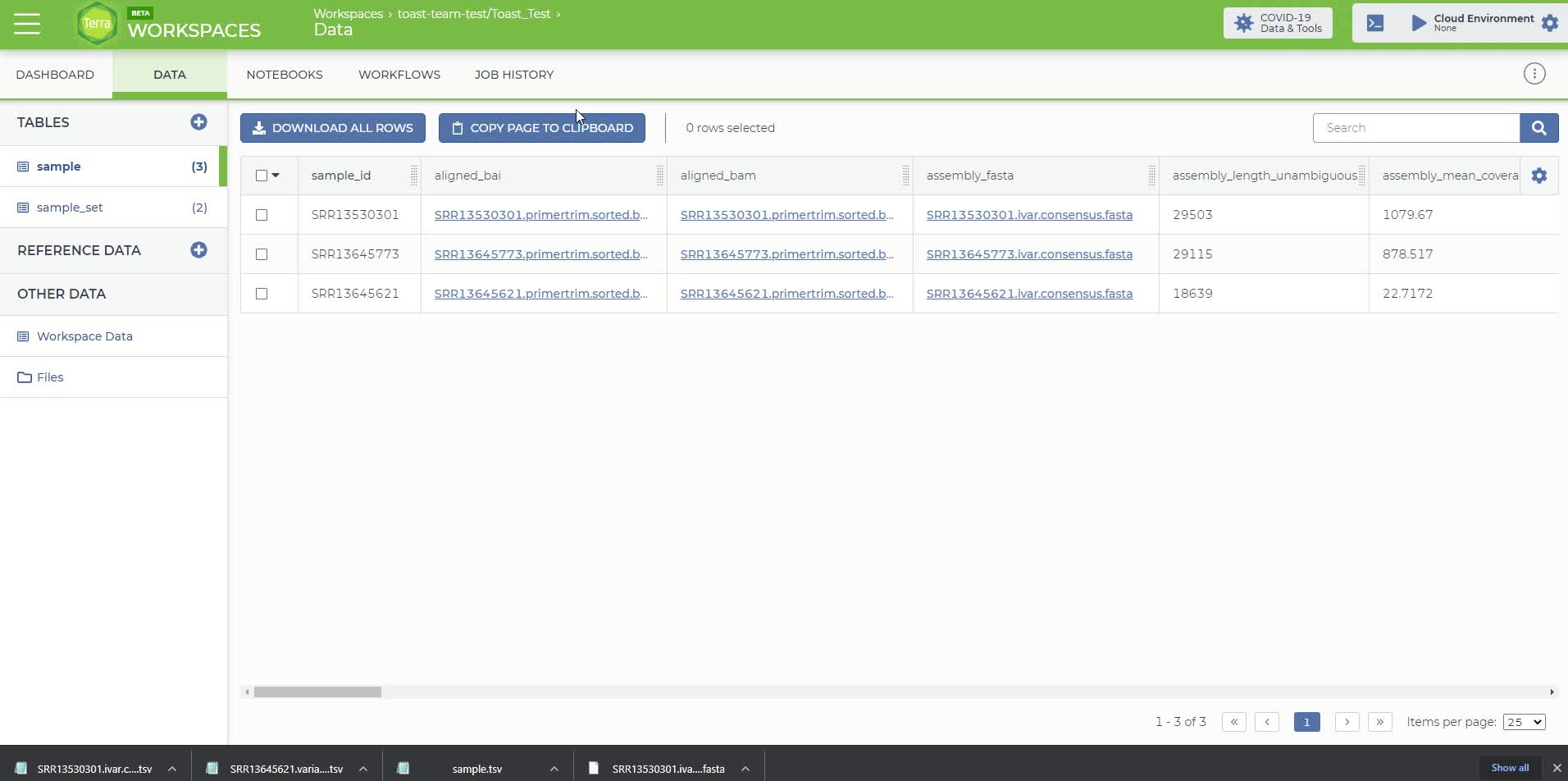
Then go to the 'Data' panel and under the 'Tables' tab click on the sample table that you created and uploaded in step 4. It will be named 'sample (#)' where # is the number of samples in your file.
The Terra sample table with the added output attributes from the Titan_Illumina_PE workflow
The Terra sample table will now include additional attributes added by the workflow, according to your specified output names (set to default in this example). You can reduce the number of visible fields by clicking the gear icon at top right. Select only the fields you want to see then click "Done".
Selecting only metrics we want to see.
To download the consensus sequence for a particular sample, click on the link in the 'assembly_fasta' attribute column and a popup window titled 'File Details' should appear. Click the 'Download For < $0.01' button, which will download the consensus sequence to your local directory. This can be uploaded to NextClade to visualize if desired (see step 10).
The 'File Details' popup window for downloading consensus sequence.
Finally, the NextClade and PANGO lineage assignments can be viewed in the Terra sample report by scrolling to the right. You can download or copy this report by using either the 'Download All Rows' or 'Copy Page To Clipboard' buttons at the top of the table.
The NextClade and PANGO lineage assignments for the 3 samples
An explanation of each output column is available here:
And also described in the video here:
Note
Additional documentation for the Titan SARS-CoV-2 workflows and the Terra platform are available.
Titan workflows: Theiagen Genomics Documentation
Terra Platform: Documentation – Terra Support
Reviewing Quality Metrics
At the end of this process, we should ideally have a complete consensus assembly sequence, which represents the entirety of the RNA sequence present in the virion in the sample. However, a full-length sequence is often not generated at the end of the protocol, but rather fragments of the genome. We can look at metrics to determine how fragmented our genome is relative to the reference sequence. There are 3 metrics in our report to help us assess this.
Assembly length unambiguous - the final count of ATCGs (i.e. unambiguous bases) within the consensus assembly.
- A higher number here will indicate a more complete consensus sequence.
The number of unambiguous bases is highlighted in orange (column 3).
Number of Ns - the converse of assembly length unambiguous - the final count of the number of Ns (i.e. ambiguous) bases. These are completely ambiguous bases due to either too little signal or too much noise for the basecaller to confidently determine what base was at this position.
- If there was an "S" (basecaller couldn't tell if it was an C or G) or "W" (basecaller couldn't tell if it was an A or T) these counts are considered "semi-ambiguous" and are not included in either the number of N or assembly length unambiguous metrics.
- A higher number of Ns will mean you have a more fragmented assembly while fewer Ns indicates a strong consensus assembly
Percent reference coverage - portion of the genome covered by consensus assembly. This uses the assembly length unambiguous metric to gauge how much of the reference sequence (Wu Han-1) is covered by the consensus assembly.
- Takes the #ATCG/length of SARS-CoV-2 reference genome (29903bp)*100
- Ideally this should to be 100%, which would mean the assembly covered 100% of the length of the reference genome unambiguously
Number of Ns and percent reference coverage columns highlighted in red (columns 1 and 2, respectively).
Here we see that the second sample has ~11k base pairs that are completely ambiguous. This has lead to only 62.33% of the reference genome being represented by this sequencing run. This indicates that the consensus sequence generated was highly fragmented. Further, depending on where this fragmentation occurred, we might not be able to trust the NextClade and PANGO lineage assignments. In contrast, the other two samples have a low number of Ns and show >97% coverage of the reference genome, suggesting these consensus sequences are fairly complete and the lineage assignments are likely to be correct if the fragmented areas are not in key regions (i.e. the spike protein). Users should refer to their lab's cut offs for quality metrics to determine if a sample requires resequencing.
If a lab is having consistent problem with contamination that is not identified to be of human origin with this workflow a larger Kraken database can be used to determine the offending organism. Contact [email protected] if you would like assistance with this or read Kraken2's documentation for how to do this on your own.
Metrics for Troubleshooting
FastQC Raw Output
These data are the number of raw reads in an input file and indicates how much sequencing occurred.
Number of reads before (raw) and after QC (clean) are highlighted in red (columns 5-6 and 7-8, respectively).
We can see that for the second sample a large number of reads were lost during the QC process, as we started with ~58k reads and only ~3k passed QC filtering parameters. We lost roughly half of the reads in the other two samples, but this might be fine if other parameters indicate that a quality consensus sequence was still recovered.
Percent of Reads Identified to be Human and SARS-CoV-2
These data indicate how much human read contamination is in the sequencing reads. Ideally, the number of reads containing human data should be as low as possible and the percent of reads assigned to SARS-CoV-2 should be high. If the sequencing run contains greater than ~15% human reads, the concentration of SARS-CoV-2 in your original sample may be too low to generate sufficient coverage depth for a good quality assembly, or there is an issue in the wet lab protocol that is leading to the low generation of SARS-CoV-2 reads.
Output from the program "Kraken" that identifies the number of human and SARS-CoV-2 reads in a sample.
The first and second sample (rows 1 and 2) are fairly free of contamination with 97.97% and 84.67% of the reads sequenced belonging to SARS-CoV-2, respectively. However, in the third sample there is a higher amount of contamination with 2.81% of reads belonging to human and only 52.73% of reads determined to be of SARS-CoV-2 origin. Thus, there is roughly ~44% of reads that are some other source of contamination that is not human. If a lab is having consistent problem with contamination that is not identified to be of human origin with this workflow, a larger Kraken database can be used to determine the offending organism. Contact [email protected] if you would like assistance with this or read Kraken2's documentation for how to do this on your own. Users should refer to their lab's cut-offs thresholds for quality metrics to determine if a sample requires resequencing.
Bam File
This file is generated during alignment and contains all information regarding the alignment to the WuHan-1 reference. This file is found in the 'aligned_bam' column and we can download this file just like we did with the consensus sequence in step 6. These files can then be visualize this with Geneious, CLC workbench or IGV.
Variants File - This is a text file that contains information regarding variants that were identified in the consensus assembly relative to the Wuhan-1 reference. This file is found in the 'ivar_tsv' column and can download this file just like we did with the consensus sequence in step 6.
Example variants file generated from iVar
If you look at the second row, this tells us that at position 1059 in the reference genome there was a "C" in the sequence that was run there was a "T" instead. You can also download and check the file in the column 'nextclade_tsv' that has specific information about exact mutations in this sample.
For Pro Users:
If for some reason you want to dig into intermediate files, you can go to the "JOB HISTORY" tab and click on the "execution directory" icon in the "Links" column. This will take us to a directory of folders that contain the output for each task.
Example google bucket with output folders generated by the Titan Illumina PE workflow
As an example we will look into into the Trimmomatic output from the sample that lost a lot of reads during the filtering process.
Submit Consensus Sequences to Public Repositories
If you plan on submitting the SARS-CoV-2 consensus sequence to either the GenBank or GISAID public repositories, please refer to the following documentation for submission criteria and minimum quality control thresholds.
GenBank Submission Criteria: About GenBank Submission (nih.gov)
GISAID Submission Criteria:  Gisaid inclusion criteria.pdf
Gisaid inclusion criteria.pdf
The Mercury workflows are designed to prepare genome assemblies and sample metadata on the Terra platform for subsequent GISAID and NCBI submission.
Mercury Overview: https://www.youtube.com/watch?v=h8YASVckOrw
After preparing consensus sequences of SARS-CoV-2 and sample metadata for submission, use the following protocols submit them to the public repositories. Completing submissions in this order allows all the sequence information to be linked together.
NextClade Visualization
After you downloading the consensus sequence for a particular sample, following directions in step 6.
Download consensus sequence
Visit to the NextClade website. Drag-and-drop the downloaded fasta file and the screen will automatically take you to the analysis page.
Drag and drop file for analysis
Hover over the different columns to get more information. Click the "Show Tree" button at the top right of the page to see where the sequence sits on the tree of other consensus sequences from SARS-Cov-2.
Clicking on 'Show Tree" button
The sequence is highlighted on the tree of SARS-CoV-2 consensus sequences.
Video of the full process:
How to debug a failed run
In this example of a failed run, we will look at an error caused by adding spaces into your collection name. If there is a failure for some other reason, you can follow similar steps to understand why your job failed.
If a run fails, you will see this indicated in the job history screen in the "status" column.
Job Failure
To understand why it failed, click the "workflow dashboard" icon in the "links" column. This will take you to a new screen and you can click the arrows next to the "message" to see what it says. Here we see there are two errors that direct us to a log file to check. To find out more, click the "execution directory" icon under the links header. This will take you to the Google Cloud location containing the run outputs.
Job failure messages
Follow the file path in the Google Cloud location to the log files that were referenced in the error messages (in red/orange text in the above photo).
Click on log file referenced in the error message.
Click on the "Authenticated URL" link that will take you to a text file.
Authenticated URL link
In the text file, we can see that there was an error that cause by there being a space between "Bad" and "Sample" in our file path that was created when we made "Bad Sample" rather than "Bad_Sample" as our collection name.
fastqc_raw.log file showing the error.
Video of the whole process.


