Nov 29, 2020
Version 1
TEA-seq V.1
This protocol is a draft, published without a DOI.
- Elliott Swanson1,
- Lucas Graybuck1
- 1Allen Institute for Immunology
- Elliott Swanson: Primary author
- Lucas Graybuck: Figures & metadata
- Human Cell Atlas Method Development Community

External link: https://www.biorxiv.org/content/10.1101/2020.09.04.283887v2
Protocol Citation: Elliott Swanson, Lucas Graybuck 2020. TEA-seq. protocols.io https://protocols.io/view/tea-seq-bpp2mmqe
Manuscript citation:
Elliott Swanson, Cara Lord, Julian Reading, Alexander T. Heubeck, Adam K. Savage, Richard Green, Xiao-jun Li, Troy R. Torgerson, Thomas F. Bumol, Lucas T. Graybuck, and Peter J. Skene. TEA-seq: a trimodal assay for integrated single cell measurement of transcription, epitopes, and chromatin accessibility. bioRxiv (2020). doi: https://doi.org/10.1101/2020.09.04.283887
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: November 13, 2020
Last Modified: November 29, 2020
Protocol Integer ID: 44506
Keywords: 10x Genomics, Trimodal, Triple, scATAC-seq, scRNA-seq, ADT, Allen Institute, Allen Institute for Immunology, AIFI, BioLegend, TotalSeq, seq tea, single cells on the 10x genomics multiome platform, 10x genomics multiome platform, use with peripheral blood mononuclear cell, transcriptomic, peripheral blood mononuclear cell, method for transcriptomic, sequencing library, seq, single cell, cell surface epitope, scrna, pbmc, cell
Disclaimer
This protocol is provided by the Allen Institute and contributors "as is" and any express or implied warranties, including, but not limited to, the implied warranties of merchantability and fitness for a particular purpose are disclaimed. In no event shall the Allen Institute or contributors be liable for any direct, indirect, incidental, special, exemplary, or consequential damages (including, but not limited to, procurement of substitute goods or services; loss of use, data, or profits; or business interruption) however caused and on any theory of liability, whether in contract, strict liability, or tort (including negligence or otherwise) arising in any way out of the use of this protocol, even if advised of the possibility of such damage.
Abstract
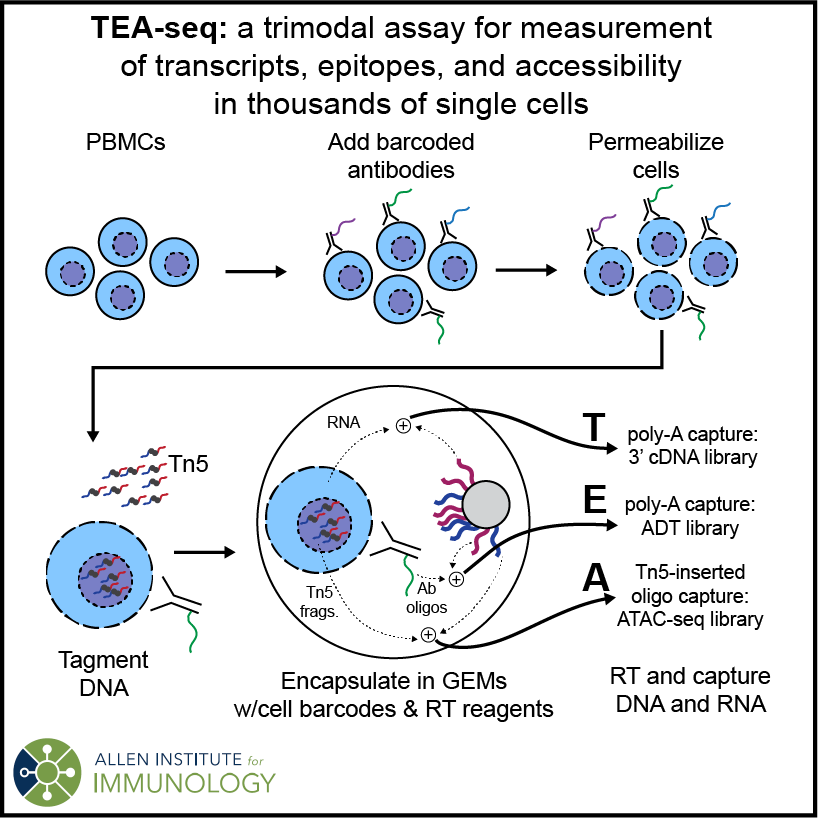
TEA-seq is a method for Transcriptomic, Epitope, and Accessibility measurement from thousands of single cells on the 10x Genomics Multiome platform. It generates scRNA-seq, scATAC-seq, and cell surface epitope (BioLegend TotalSeq) sequencing libraries linked by 10x cell barcodes for coordinated analysis. TEA-seq has been optimized for use with peripheral blood mononuclear cells (PBMCs).
Guidelines
Permeabilization testing:
Cell permeabilization in this protocol utilizes Digitonin, a naturally-derived saponin molecule. Because it's a natural product, the precise concentration and permeabilization strength may vary somewhat from source to source and batch to batch. We recommend testing your digitonin permeabilization using your cell type of interest and a live/dead cell stain (e.g. AO/PI) to find the lowest concentration that provides reliable, reproducible permeabilization (100% of cells "dead" by AO/PI stain)
Cell Handling:
Following thaw, maintain cells On ice or at 4 °C at all times.
Antibodies and titering:
Successful measurement of antibody-bound oligos depends on antibody titer and on retention of antibody binding after digitonin treatment. Because digitonin alters membrane composition, some membrane-bound proteins may be lost after permeabilization.
Antibodies for which we have noticed reduced binding after digitonin treatment:
CD20 Clone 2H7
Titrations we have used in our study:
| Target | Clone | TEA-seq ug Ab / M cells | |
| CD10 | HI10a | 0.5 | |
| CD11b | ICRF44 | 0.05 | |
| CD11c | S-HCL-3 | 0.025 | |
| CD123 | 6H6 | 0.1 | |
| CD127 (IL-7Rα) | A019D5 | 0.025 | |
| CD14 | M5E2 | 0.2 | |
| CD141 (Thrombomodulin) | M80 | 0.1 | |
| CD16 | 3G8 | 0.05 | |
| CD172a (SIRPa) | 15-414 | 0.25 | |
| CD185 (CXCR5) | J252D4 | 0.125 | |
| CD19 | HIB19 | 0.2 | |
| CD192 (CCR2) | K036C2 | 0.5 | |
| CD197 (CCR7) | G043H7 | 0.5 | |
| CD21 | Bu32 | 0.05 | |
| CD24 | ML5 | 0.5 | |
| CD25 | BC96 | 0.08 | |
| CD269 (BCMA) | 19F2 | 0.5 | |
| CD27 | LG.3A10 | 0.125 | |
| CD278 (ICOS) | C398.4A | 0.01 | |
| CD279 (PD-1) | EH12.2H7 | 0.1 | |
| CD3 | UCHT1 | 0.05 | |
| CD304 (Neuropilin-1) | 12C2 | 0.1 | |
| CD319 (CRACC) | 162.1 | 0.5 | |
| CD38 | HB-7 | 0.05 | |
| CD39 | A1 | 0.05 | |
| CD4 | RPA-T4 | 0.1 | |
| CD40 | 5C3 | 0.25 | |
| CD45RA^ | HI100 | 0.0625 | |
| CD45RO | UCHL1 | 0.1 | |
| CD56 (NCAM) | 5.1H11 | 0.15 | |
| CD66b | 6/40c | 0.25 | |
| CD71 | CY1G4 | 0.025 | |
| CD80 | 2D10 | 0.5 | |
| CD86 | IT2.2 | 0.05 | |
| CD8a | RPA-T8 | 0.2 | |
| CD95 (Fas) | DX2 | 0.1 | |
| FcεRIα | AER-37 (CRA-1) | 0.5 | |
| HLA-DR | L243 | 0.25 | |
| IgD | IA6-2 | 0.05 | |
| Mouse IgG1, κ Isotype Control | MOPC-21 | 0.5 | |
| IgM | MHM-88 | 0.05 | |
| KLRG1 (MAFA) | SA231A2 | 0.25 | |
| TCR Vα24-Jα18 (iNKT cell) | 6B11 | 0.5 | |
| TCR Vα7.2 | 3C10 | 0.05 | |
| TCR α/β | IP26 | 0.125 | |
| TCR γ/δ | B1 | 0.5 |
^ Low in our experience - may want to increase for future studies
Plate Sealing:
We recommend using an automated microplate heat-sealer for all steps following the GEM barcoding reaction. We find that we get more consistent results and higher yield by heat-sealing and vortexing or mixing by inversion, than by pipette mixing. If you do not have access to a heat-sealer Bio-Rad Microseal B adhesive seals can be used in combination with pipette mixing.
Reagent Handling:
Primers and Buffers may be thawed at room temperature. With the exception of Cleanup Buffer, store buffers and primers on ice once thawed. Reagents containing enzymes should be removed from -20 °C immediately before use and maintained on ice. Return enzymes to -20 °C storage immediately after use. Buffers and primers may be vortexed. Do not vortex enzyme containing buffers.
Materials
10x Genomics equipment and reagents:
10x Chromium Controller (10x Genomics)
Chromium Next GEM Chip J Single Cell10x GenomicsCatalog #1000230
Chromium Next GEM Single Cell Multiome ATAC/Gene Expression Reagent Bundle10x GenomicsCatalog #1000285
Dual Index Kit TT Set A10x GenomicsCatalog #1000215
Single Index Kit N Set A10x GenomicsCatalog #1000212
Other equipment (as tested - others may work):
Beckman Coulter Avanti J-15RIVD centrifuge with JS4.750 swinging bucket, B99516
C1000 Touch Thermal Cycler with 96-Deep Well Reaction Module (Bio-rad #1851197)
CFX96 Touch Real-Time PCR Detection System (Bio-rad #1855195)
BioAnalyzer (Agilent G2939A)
Vortex Mixer
Vendor-specific Reagents:
AIM V™ Medium, liquidThermo Fisher ScientificCatalog #12055091
NxGen RNAse InhibitorLucigenCatalog #30281-2
FC blocking reagent (FcX)BioLegend
SPRIselect ReagentBeckman CoulterCatalog #B23317
Dynabeads™ MyOne™ SilaneThermo FisherCatalog #37002D
Buffer EBQiagenCatalog #19086
Kapa HiFi HotStart ReadyMixKapa BiosystemsCatalog #KM2602
KAPA Library Quantification Kit for Illumina® PlatformsKapa BiosystemsCatalog #KK4835
Agilent High Sensitivity DNA KitAgilent TechnologiesCatalog #5067-4626
Generic Reagents:
Dulbecco's phosphate-buffered saline (DPBS)
Bovine serum albumin (BSA)
50% Glycerol
Digitonin
DMSO, anhydrous
MgCl2
NaCl
Tris-HCl pH 7.4, a.k.a. Tris(hydroxymethyl)aminomethane hydrochloride
Low TE Buffer (10 mM Tris-HCl, 0.1 mM EDT, pH 8.0)
Labware (as tested - others may work):
Falcon® Round-Bottom Tubes Disposable Polystyrene Corning® 5 mLCorningCatalog #352054
Eppendorf Safe-Lock Tubes 1.5 mL PCR clean colorless 500 tubesEppendorfCatalog #022363212
Falcon® 5 mL Round Bottom Polystyrene Test Tube, with Cell Strainer Snap CapCorningCatalog #352235
Eppendorf® twin.tec 96-Well PCR Plate Semi-SkirtedEppendorfCatalog #951020303
Eppendorf twin.tec® PCR 96-well plate, skirtedEppendorfCatalog #951020401
Microseal® ‘B’ Adhesive SealsBio-Rad LaboratoriesCatalog #MSB-1001
Additional Oligos:
| Name | Sequence (5'->3') | Length | |
| SI-PCR-Oligo | AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTC | 49 | |
| ADT-Rev-AMP | CCTTGGCACCCGAGAATTCC | 20 | |
| ADT-i7 | CAAGCAGAAGACGGCATACGAGATXXXXXXXXGTGACTGGAGTTCCTTGGCACCCGAGAATTCC*A | 65 |
Oligonucleotides required for ADT amplification. Note that for ADT-i7 Primers, You'll need to add 8nt index sequences in place of the stretch of 'X' nucleotides. * designates the location of a Phosphorothioate bond.
External equipment:
Illumina NovaSeq 6000
Protocol materials
Eppendorf twin.tec® PCR 96-well plate, skirtedEppendorfCatalog #951020401
NxGen RNAse InhibitorLucigenCatalog #30281-2
Dynabeads™ MyOne™ SilaneThermo FisherCatalog #37002D
Single Index Kit N Set A10x GenomicsCatalog #1000212
AIM V™ Medium, liquidThermo Fisher ScientificCatalog #12055091
Chromium Next GEM Chip J Single Cell10x GenomicsCatalog #1000230
Kapa HiFi HotStart ReadyMixKapa BiosystemsCatalog #KM2602
KAPA Library Quantification Kit for Illumina® PlatformsKapa BiosystemsCatalog #KK4835
Falcon® Round-Bottom Tubes Disposable Polystyrene Corning® 5 mLCorningCatalog #352054
Eppendorf Safe-Lock Tubes 1.5 mL PCR clean colorless 500 tubesEppendorfCatalog #022363212
Falcon® 5 mL Round Bottom Polystyrene Test Tube, with Cell Strainer Snap CapCorningCatalog #352235
Chromium Next GEM Single Cell Multiome ATAC/Gene Expression Reagent Bundle10x GenomicsCatalog #1000285
Eppendorf® twin.tec 96-Well PCR Plate Semi-SkirtedEppendorfCatalog #951020303
Microseal® ‘B’ Adhesive SealsBio-Rad LaboratoriesCatalog #MSB-1001
Dual Index Kit TT Set A10x GenomicsCatalog #1000215
Buffer EBQiagenCatalog #19086
Agilent High Sensitivity DNA KitAgilent TechnologiesCatalog #5067-4626
FC blocking reagent (FcX)BioLegend
SPRIselect ReagentBeckman CoulterCatalog #B23317
Troubleshooting
Safety warnings
Safety information
Digitonin is an acutely toxic health hazard.
Handling of digitonin-containing solutions, especially at high concentration, should be performed with personal protective equipment and in a biosafety cabinet.
Please refer to an MSDS for your Digitonin product (e.g. this version from Fisher) before beginning permeabilization experiments, and take all necessary precautions.
Before start
1. Pre-chill a swinging-bucket rotor centrifuge to 4 °C for use throughout antibody staining protocol, with holders for 5 mL polystyrene round-bottom tubes.
2. If performing bead cleanup steps (36 and onward), remove Dynabeads MyOne SILANE reagent from storage at least 00:30:00 before beginning the cleanup to allow them to equilibrate to room temperature.
Buffer preparation
30m
Stain Buffer Dulbecco's phosphate-buffered saline (DPBS) supplemented with 2% w/v bovine serum albumin.
Wash Buffer Final composition of 20 mM Tris HCl (Tris(hydroxymethyl)aminomethane hydrochloride) pH 7.4, 150 mM NaCl, 3 mM MgCl.
Perm Buffer Identical components as Wash Buffer above with the addition of digitonin to a final concentration of 0.01% w/v. Stocks of 5% w/v digitonin in DMSO stocks can be prepared ahead of time and aliquots can be stored in -20 °C for future use. We recommend limiting the number of freeze-thaw cycles to five.
Note
Digitonin concentration was optimized for PBMCs. Permeabilization of different sample types may require further optimization.
Tagmentation Buffer Identical components as Wash Buffer above with the addition of NxGen RNase Inhibitor at a final concentration of 1U / µl.
Sample preparation
1h
Remove cryopreserved cells from liquid nitrogen storage and thaw in a 37 °C water bath for 3-5 minutes until no ice is visible.
Note
This cell thaw protocol has been optimized for cryopreserved PBMCs. Alternate cell types or preservation methods may require different sample preparation procedures.
Slowly dilute cells in 10 mL of pre-warmed 37 °C AIM V™ Medium, liquidThermo Fisher ScientificCatalog #12055091 adding the first 3 mL dropwise.
Centrifuge the cells at 400 x g, 4°C, 00:05:00 and remove the supernatant.
Resuspend the cell pellet in 1 mL of ice cold Dulbecco's phosphate-buffered saline supplemented with 0.2% w/v bovine serum albumin (DPBS + 0.2% BSA) by pipette mixing. Bring the volume to 10 mL with DPBS + 0.2% BSA.
Centrifuge the cells at 400 x g, 4°C, 00:05:00 and remove the supernatant.
Resuspend the cell pellet in 1 mL of ice cold Dulbecco's phosphate-buffered saline supplemented with 0.2% w/v bovine serum albumin (DPBS + 0.2% BSA) by pipette mixing. Dilute the cell suspension in DPBS + 0.2% BSA as desired, targetting a cell concetration of 1 - 5 million cells / mL.
Count cells before proceeding to Antibody Staining.
Antibody staining
1h 30m
Centrifuge cells (recommended input of 1.5 - 5 million) at 400 x g, 4°C, 00:05:00 in a 5 mL polystyrene round bottom tube and remove the supernatant.
Resuspend the cell pellet in ice cold Stain Buffer (DPBS + 2% BSA) by gently pipette mixing.
Add 10 ul of TruStain FcX and pipette mix thoroughly. Incubate cells for 00:10:00 on ice .
10m
Add TotalSeq-A antibodies and pipette mix thoroughly. Incubate cells for 00:30:00 on ice .
30m
Dilute cells with 4 mL of ice cold Stain Buffer and gently mix.
Centrifuge cells at 400 x g, 4°C, 00:05:00 and remove the supernatant.
Resuspend the cell pellet in 4 mL of ice cold Stain Buffer by gently pipette mixing.
Centrifuge cells at 400 x g, 4°C, 00:05:00 and remove the supernatant.
5m
for a total of three washes.
10m
Resuspend the cell pellet in 1 mL of ice cold Stain Buffer and count the cells.
Permeabilization
30m
Aliquot 1 million stained cells into a 1.5 mL low bind tube and centrifuge at 400 x g, 4°C, 00:05:00 . Remove the supernatant.
5m
Resuspend the cell pellet in 100 µl of ice cold Perm Buffer by pipette mixing. Incubate cells for 00:05:00 on ice .
5m
Dilute cells with 1mL of ice cold Wash Buffer and gently mix.
Centrifuge cells at 400 x g, 4°C, 00:05:00 and remove the supernatant.
5m
Resuspend the cell pellet in 50 - 100 µl of ice cold Tagmentation Buffer depending on the desired cell input into the GEM barcoding reaction. An input of 15k cells requires a final concentration of 3k cells per µl.
Filter cell suspension using a 35 µm cell strainer to remove cell clumps. Transfer filtered solution to a new tube.
Count the cells and calculate the volume of cell suspension required for the desired cell input.
Tagmentation and GEM generation
3h
Aliquot the desired number of cells to a new 96-well skirted PCR plate and bring volume to 5 µl with Tagmentation Buffer.
Create a master mix containing 7 µl of ATAC Buffer B and 3 µl of ATAC Enzyme B. Add 10 µl of master mix to each sample and gently pipette mix ten times.
Seal the plate and incubate on a thermal cycler for 01:00:00 at 37 °C . Hold the reaction at 4 °C upon completion. Allow samples to cool to 4 °C , then proceed immediately to GEM generation (Step 28).
Perform steps 24 - 27 during the incubation time.
1h
During the tagmentation reaction remove the Single Cell Multiome Gel Beads, Template Switch Oligo, Reducing Agent B, Barcoding Reagent Mix, and Quenching Agent from storage and allow them to thaw at room temperature. First use of the Template Switch Oligo requires resuspension in 80 µl of Low TE Buffer.
Remove the Barcoding Enzyme Mix from storage 10 - 15 minutes before the end of the tagmentation reaction and maintain On ice .
Create a barcoding master mix on ice as detailed in the 10x Multiome ATAC + Gene Expression user guide.
Load a Chromium Next GEM Chip J into a secondary holder. Dispense 50% Glycerol into unused chip wells as follows:
Row 1: 70 µl
Row 2: 50 µl
Row 3: 40 µl
Remove the sample plate from the thermal cycler once it reaches 4 °C and maintain On ice . Add 60 µl of master mix to each sample well and gently pipette mix five times.
Load 70 µl of master mix plus sample into row 1 of the chip.
Vortex the Multiome Gel Beads for 00:00:30 and add 50 µl to row 2 of the chip.
30s
Add 45 µl of Partitioning Oil to row 3 of the chip.
Place a gasket over the loaded chip and load it onto the Chromium Controller. Initiate the run.
At completion of the run, slowly aspirate 100 µl of GEMs from the chip and slowly dispense into a new half-skirt plate on ice as detailed in the 10x Multiome ATAC + Gene Expression user guide (step 2.4 of the guide).
Incubate GEMs on a thermal cycler for 00:45:00 at 37 °C , followed by 00:30:00 at 25 °C , ending with a 4 °C hold.
1h 15m
Add 5 µl of Quenching Agent to each sample and mix.
Note
Optional Stopping Point: GEMs can be stored at -80 °C for for up to four weeks.
GEM cleanup
1h 30m
Remove Dynabeads MyOne SILANE reagent from storage at least 30 minutes before beginning the cleanup and allow it to equilibrate to room temperature.
Remove the Cleanup Buffer from-20 °C storage. Heat at 65 °C and thoroughly vortex until no precipitates are visible (approx. 00:20:00 ).
20m
Perform the Post GEM Incubation dynabead cleanup as detailed in the 10x Multiome ATAC + Gene Expression User Guide (step 3.1 of the guide).
Perform a 2.0x bead:sample SPRIselect cleanup. Bind beads for 00:10:00 at Room temperature . Elute DNA off the beads by resuspending in 45.5 µl of Buffer EB. Transfer 45 µl of sample to a new plate.
10m
Pre-Amplification
1h 30m
Remove Pre-Amp Primers from storage and allow them to thaw at room temperature. Remove Pre-Amp Mix from storage and thaw on ice. Prepare a 0.2 µM solution of ADT-Rev-AMP primer in Low TE Buffer (can be prepared ahead of time and frozen at -20 °C ).
On ice prepare a Pre-Amp master mix consisting of 50 µl of Pre-Amp Mix, 4 µl of Pre-Amp Primers, 1 µl of 0.2 µM ADT-Rev-AMP primer per sample.
On ice , add 55 µl of Pre-Amp master mix to 45 µl of sample and mix.
Incubate reactions on a thermal cycler using the following 7 cycle PCR protocol:
Lid Temperature: 105°C
Reaction Volume: 100 µl
| Step | Temperature | Time | |
| 1 | 72°C | 5 min | |
| 2 | 98°C | 3 min | |
| 3 | 98°C | 20 sec | |
| 4 | 63°C | 30 sec | |
| 5 | 72°C | 1 min | |
| 6 | - | Go to step 3 x6 (total of 7 cycles) | |
| 7 | 72°C | 1 min | |
| 8 | 4°C | Hold |
7 cycle Pre-Amp PCR protocol
Note
Optional Stopping Point: Following amplification, reactions can be held at 4 °C for up to 72 hours.
Perform a 2.0x bead:sample SPRIselect cleanup. Bind beads for 10 minutes at room temperature. Elute DNA off the beads by resuspending in 160.5 µl of Buffer EB. Transfer 160 µl of sample to a new plate.
Note
Optional Stopping Point: Following cleanup, libraries can be held at 4 °C for up to 72 hours or stored at -20 °C .
ATAC Library Prep
1h 30m
Remove SI-PCR Primer B and Sample Index plate N, Set A from -20 °C storage and allow them to thaw at room temperature. Remove Amp Mix from-20 °C storage and thaw on ice.
On ice , prepare a PCR master mix consisting of 50 µl Amp Mix and 7.5 µl SI-PCR Pimer B per sample.
On ice , add 57.5 µl of PCR master mix and 2.5 µl of an individual index to 40 µl of each pre-amplified sample. Mix thoroughly.
Incubate reactions on a thermal cycler using the following 9 cycle PCR protocol:
Lid Temperature: 105°C
Reaction Volume: 100 µl
| Step | Temperature | Time | |
| 1 | 98°C | 45 sec | |
| 2 | 98°C | 20 sec | |
| 3 | 67°C | 30 sec | |
| 4 | 72°C | 20 sec | |
| 5 | - | Go to step 2 x8 (total of 9 cycles) | |
| 6 | 72°C | 1 min | |
| 7 | 4°C | Hold |
9 cycle ATAC indexing PCR protocol
Note
Optional Stopping Point: Following amplification, reactions can be held at 4 °C for up to 72 hours.
Perform a dual-sided 0.6x/1.6x bead:sample SPRIselect size-selection clean-up as detailed in the 10x Multiome ATAC + Gene Expression User Guide (step 5.2 of the guide).
Note
Optional Stopping Point: Final ATAC libraries can be stored at -20 °C .
cDNA Amplification
2h
Remove cDNA Primers from storage and allow them to thaw at room temperature. Remove Amp Mix from storage and thaw on ice. Prepare a 2 µM solution of ADT-Rev-AMP primer in Low TE buffer (can be prepared ahead of time and frozen at -20 °C ).
On ice , prepare a cDNA Amplification master mix consisting of 50 µl of Amp Mix, 15 µl of cDNA Primers, and 1 µl of 2 µM ADT-Rev-AMP primer per sample.
On ice , add 66 µl of cDNA Amplification master mix to 35 µl of each pre-amplified sample. Mix thoroughly.
Incubate reactions on a thermal cycler using the following 8 cycle PCR protocol:
Lid Temperature: 105°C
Reaction Volume: 101 µl
| Step | Temperature | Time | |
| 1 | 98°C | 3 min | |
| 2 | 98°C | 15 sec | |
| 3 | 63°C | 20 sec | |
| 4 | 72°C | 1 min | |
| 5 | - | Go to step 2 x7 (total of 8 cycles) | |
| 6 | 72°C | 1 min | |
| 7 | 4°C | Hold |
8 cycle cDNA Amplification protocol
Note
Optional Stopping Point: Following amplification, reactions can be held at 4 °C for up to 72 hours.
Perform a dual-sided 0.6x/2.0x bead:sample SPRIselect size-selection clean-up. Add 60 µl of SPRIselect beads to each reaction, incubate at room temperature for 5 minutes, incubate on a magnet for 5 minutes or until clear, transfer the supernatant to a new half-skirt plate. Supernatant contains the ADT fragments and is retained for further processing.
Complete the SPRI cleanup on the large, cDNA fragments by performing two washes with 80% ethanol. Elute cDNA off the beads by resuspending in 40.5 µl of Buffer EB. Transfer 40 µl of each cDNA library to a new plate.
Add an additional 140 µl of SPRI beads to the ADT containing supernatant from the first cleanup. Incubate at room temperature for 10 minutes, incubate on a magnet and remove the supernatant. Perform two washes with 80% ethanol, elute ADTs off the beads by resuspending in 45.5 µl of Buffer EB. Transfer 45 µl of ADT solution to a new plate.
Note
Optional Stopping Point: Following cleanup, amplified cDNA can be held at 4 °C for up to 72 hours or stored at -20 °C .
Run 1 ul of each amplified cDNA library on an Agilent Bioanalyzer High Sensitivity chip at a 1:10 dilution. Set a region at 200 - 9000 bp and record the concentration in pg / µl. Calculate the total cDNA yield in ng. Consult the 10x Multiome ATAC + Gene Expression user guide if adjusting the number of PCR cycles in the Gene Expression indexing reaction is necessary.
ADT indexing PCR
2h
Remove KAPA HiFi HotStart ReadyMix from -20 °C storage and thaw on ice. Remove 10 µM SI-PCR Primer and 10 µM ADT-i7 primers from -20 °C storage and thaw at room temperature.
On ice , add 50 µl KAPA HiFi HotStart ReadyMix, 2.5 µl 10 µM SI-PCR Primer, and 2.5 µl of an individual 10 µM ADT-i7 primer to each 45 µl ADT sample. Mix thoroughly.
Incubate reactions on a thermal cycler using the following 15 cycle PCR protocol:
Lid Temperature: 105°C
Reaction Volume: 100 µl
| Step | Temperature | Time | |
| 1 | 95°C | 3 min | |
| 2 | 95°C | 20 sec | |
| 3 | 60°C | 30 sec | |
| 4 | 72°C | 20 sec | |
| 5 | - | Go to step 2 x14 (total of 15 cycles) | |
| 6 | 72°C | 5 min | |
| 7 | 4°C | Hold |
15 cycle ADT indexing PCR protocol
Note
Optional Stopping Point: Following amplification, reactions can be held at 4 °C for up to 72 hours.
Perform a 1.6x bead:sample SPRIselect cleanup. Bind beads for 5 minutes at room temperature. Elute DNA off the beads by resuspending in 30.5 µl of Buffer EB. Transfer 30 µl of final ADT library to a new plate.
Note
Optional Stopping Point: Final ADT libraries can be stored at -20 °C .
Gene Expression Library Prep
4h
Remove Fragmentation Buffer, Adapter Oligos, Ligation Buffer, and Dual Index Plate TT Set A from -20 °C storage and thaw at room temperature. Remove Fragmentation Enzyme, DNA Ligase, and Amp Mix immediately before use and maintain On ice .
On ice , prepare a fragmentation master mix conisting of 5 µl of Fragmentation Buffer and 10 µl of Fragmentation Enzyme per sample.
Transfer 10 µl of each amplified cDNA library to a new plate. On ice , add 25 µl of Buffer EB and 15 µl of fragmentation master mix to each sample. Mix thoroughly.
Incubate reactions on a pre-cooled thermal cycler holding at 4 °C . Skip to the next step to ramp to 32 °C for 00:05:00 , followed by 65 °C for 00:30:00 , ending with a 4 °C hold.
35m
Perform a dual-sided 0.6x/0.8x bead:sample SPRIselect size-selection clean-up as detailed in the 10x Multiome ATAC + Gene Expression User Guide (step 7.2 of the guide).
On ice , prepare an adapter ligation master mix consisting of 20 µl of Ligation Buffer, 10 µl of DNA Ligase, and 20 µl of Adapter Oligos.
On ice , add 50 µl of adapter ligation master mix to each sample well. Mix thoroughly.
Incubate reactions on a thermal cycler at 20 °C for 00:15:00 f, ending with a hold at 4 °C .
15m
Perform a 0.8x bead:sample SPRIselect cleanup as detailed in the 10x Multiome ATAC + Gene Expression User Guide (step 7.4 of the guide).
On ice , add 50 µl of Amp Mix and 20 µl of an individual Dual Index TT Set A to each sample. Mix thoroughly.
Incubate reactions on a thermal cycler using the following 14 cycle PCR protocol:
Lid Temperature: 105°C
Reaction Volume: 100 µl
| Step | Temperature | Time | |
| 1 | 98°C | 45 sec | |
| 2 | 98°C | 20 sec | |
| 3 | 54°C | 30 sec | |
| 4 | 72°C | 20 sec | |
| 5 | - | Go to step 2 x13 (total of 14 cycles) | |
| 6 | 72°C | 1 min | |
| 7 | 4°C | Hold |
14 cycle Gene Expression indexing PCR protocol
Note
If desired, the number of PCR cycles can be altered depending on cDNA yield. Reference the 10x Multiome ATAC + Gene Expression User Guide (step 7.5 of the guide) for vendor recommendations.
Note
Optional Stopping Point: Following amplification, reactions can be held at 4 °C for up to 72 hours.
Perform a dual-sided 0.6x/0.8x bead:sample SPRIselect size-selection clean-up as detailed in the 10x Multiome ATAC + Gene Expression User Guide (step 7.6 of the guide).
Note
Optional Stopping Point: Final Gene Expression libraries can be stored at -20 °C .
Final Library QC
2h
Run each ATAC, Gene Expression, and ADT final library on an Agilent Bioanalyzer High Sensitivity chip or equivalent electrophoresis assay.
Note
We recommend loading libraries at a 1:10 dilution for greater accuracy.
Set regions around each library trace to estimate the average base-pair size of the library. Evaluate the traces for presence of primer-dimer of other processing artifacts.
Quantify library concentration via qPCR using the KAPA Library Quantification Kit for Illumina® PlatformsKapa BiosystemsCatalog #KK4835 or equivalent kit. Specifications will vary by qPCR instrument.
Expected result
ATAC final library Bioanalyzer trace (1:10 dilution), above.
Gene Expression final library Bioanalyzer trace (1:10 dilution), above.
ADT final library Bioanalyzer trace (1:10 dilution), above.
Calculate a size-adjusted concentration (in pM) for each library using the electropherogram and qPCR data. Library concentrations determined using methods other than qPCR such as PicoGreen will be much less accurate for this assay, especially for ADT libraries.
Multiply the calculated average concentration with the following factor:
Size of DNA Standard in bp (452) / Average fragment length of library in bp
Sequencing
Sequence libraries on the desired Illumina instrument platform. ATAC libraries will require different i5 index read lengths depending on whether the platform uses the Forward or Reverse Complement indexing workflow. The Reverse Complement workflow requires ATAC libraries to be sequenced on a separate flow cell with a longer i5 read length due to the spacer in the capture sequence.
The Forward Strand indexing workflow allows for all three library types to be sequenced on the same flowcell.
Read length for Forward Strand indexing workflow joint flow cell (ATAC, Gene Expression, ADT)
Read 1: 50 bp
i7 Index: 10 bp
i5 Index: 16 bp
Read 2: 90 bp
Read length for Forward Strand indexing workflow ATAC only flow cell
Read 1: 50 bp
i7 Index: 8 bp
i5 Index: 16 bp
Read 2: 50 bp
Read length for Reverse Strand indexing workflow ATAC only flow cell
Read 1: 50 bp
i7 Index: 8 bp
i5 Index: 24 bp
Read 2: 50 bp
Read length for Gene Expression plus ADT (or Gene Expression only) flow cell
Read 1: 28 bp
i7 Index: 10 bp
i5 Index: 10 bp
Read 2: 90 bp
Read length for ADT only flow cell
Read 1: 28 bp
i7 Index: 8 bp
Read 2: 15 bp
Sequencing Depth
We recommend a minimum read count of 30k for Gene Expression, 60k for ATAC, and 12k for ADT, per cell when sequencing PBMCs. Other cell types may require additional sequencing. In our experience the required sequencing depth of the ADT library tends to scale with the size of the panel, with the exception of rare markers.
Reference the 10x Genomics Sequencing Requirements for Single Cell Multiome ATAC + Gene Expression support page for more details.
Demultiplexing
Demultiplexing of ATAC and Gene Expression data can be performed using either Cell Ranger mkfastq (10x Genomics) or bcl2fastq2 (Illumina). We prefer bcl2fastq2 as we have found that it significantly out performs mkfastq in terms of processing time and flexibility. Flow cells containing both ATAC and Gene Expression libraries must use bcl2fastq2 for demultiplexing.
Software
bcl2fastq
NAME
Illumina
DEVELOPER
Software
Cell Ranger
NAME
10x Genomics
DEVELOPER
The commands below may be used to demultiplex TEA-seq data by library type. Use of bcl2fastq2 requires a sample sheet in standard Illumina Experiment Manager format for each library type.
# demultiplex gene expression data
bcl2fastq --use-bases-mask=Y28n*,I10,I10n*,Y90n* \
--create-fastq-for-index-reads \
--minimum-trimmed-read-length=8 \
--mask-short-adapter-reads=8 \
--ignore-missing-positions \
--ignore-missing-filter \
--ignore-missing-bcls \
-r 24 -w 24 -p 80 \
-R ${FLOWCELL_DIR} \
--output-dir={OUTPUT_DIR} \
--interop-dir={INTEROP_DIR} \
--sample-sheet={GEX_SAMPLE_SHEET_PATH}
# demultiplex ATAC data
bcl2fastq --use-bases-mask=Y50n*,I8n*,Y16,Y50n* \
--create-fastq-for-index-reads \
--minimum-trimmed-read-length=8 \
--mask-short-adapter-reads=8 \
--ignore-missing-positions \
--ignore-missing-filter \
--ignore-missing-bcls \
-r 24 -w 24 -p 80 \
-R ${FLOWCELL_DIR} \
--output-dir={OUTPUT_DIR} \
--interop-dir={INTEROP_DIR} \
--sample-sheet={ATAC_SAMPLE_SHEET_PATH}
# demultiplex ADT data
bcl2fastq --use-bases-mask=Y28n*,I8n*,n*,Y90n* \
--create-fastq-for-index-reads \
--minimum-trimmed-read-length=8 \
--mask-short-adapter-reads=8 \
--ignore-missing-positions \
--ignore-missing-controls \
--ignore-missing-filter \
--ignore-missing-bcls \
-r 24 -w 24 -p 80 \
-R {FLOWCELL_DIR} \
--output-dir={OUTPUT_DIR} \
--interop-dir={INTEROP_DIR} \
--sample-sheet={ADT_SAMPLE_SHEET_PATH}
These commands have been written into a Bash script for ease of use. It is required to have bcl2fastq2 added to the environment path before use.
Data Preprocessing and analysis
Please see our Github repository for downstream data processing and analysis code:
