May 01, 2023
Version 1
- Christoph Geisenberger1,2,
- Jeroen van den Berg1,
- Vincent van Batenburg1,
- Buys De Barbanson1,
- Jeroen de Ridder1,
- Alexander van Oudenaarden1
- 1Hubrecht Institute-KNAW (Royal Netherlands Academy of Arts and Sciences), Oncode Institute, Utrecht, The Netherlands;
- 2Ludwig-Maximilians-Universität, Munich, Germany

External link: https://doi.org/10.1038/s41592-025-02847-4
Protocol Citation: Christoph Geisenberger, Jeroen van den Berg, Vincent van Batenburg, Buys De Barbanson, Jeroen de Ridder, Alexander van Oudenaarden 2023. Single-cell Epi2-Seq. protocols.io https://protocols.io/view/single-cell-epi2-seq-cqk7vuzn
Manuscript citation:
Geisenberger C, Berg Jvd, Batenburg Vv, Barbanson Bd, Lyubimova A, Verity-Legg J, Chen X, Liu Y, Song C, Ridder Jd, Oudenaarden Av (2025) Single-cell multi-omic detection of DNA methylation and histone modifications reconstructs the dynamics of epigenomic maintenance. Nature Methods 22(10). doi: 10.1038/s41592-025-02847-4
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: March 06, 2023
Last Modified: May 01, 2023
Protocol Integer ID: 78207
Keywords: Methylation, TAPS, Single-cell, Epigenomics, Chromatin, Bisulfite, dna methylation in individual cell, resolution dna methylation profiling, dna methylation, cell epi2, joint readout of histone modification, single cell, assisted pyridine borane sequencing, histone modification, dna damage, cell application, dna, targeted mnase digestion, individual cell, cell adapter, cell
Abstract
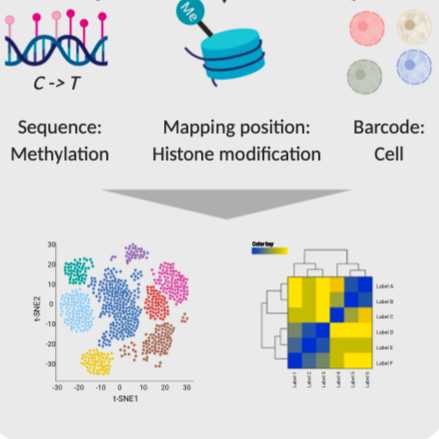
Here we describe the full protocol for single-cell Epi2-Seq which enables joint readout of histone modifications and DNA methylation in individual cells. In short, this methods combines antibody-targeted MNase digestion (ChIC, CUT&RUN) with Tet-Assisted Pyridine Borane Sequencing (TAPS), a bisulfite-free technique for base-resolution DNA methylation profiling. TAPS, unlike bisulfite, specifically converts 5mC which increases mapping rates and makes it feasible to use unmethylated single-cell adapters. Also, DNA damage is minimal which reduces input requirements and makes it well suited for single-cell applications.
Materials
General materials
- Protein lo-bind tubes (Eppendorf, 0030108094 and 0030122216)
- PBS (Thermo Fisher, AM9624)
- Nuclease-free H20 (Invitrogen, AM9932)
- Mineral oil (Sigma-Aldrich, 69794-500ML)
- Aluminum Plate sealers
- 384-well hard shell plates
- Ampure XP beads (Beckman Coulter, A63881)
- Ethanol 100% (VWR, 437435L)
Preparation of methylated spike-ins
- Unmethylated lambda phage DNA (Promega, D1521)
- NEB Buffer 2 (NEB, cat. no. M0226S)
- SAM (NEB, cat. no. M0226S)
- M.SssI (NEB, cat. no. M0226S)
- CutSmart Buffer (NEB, R0125S)
- NlaIII (NEB, R0125S)
- T4 DNA ligase buffer (NEB, M0202L)
- T4 DNA ligase (NEB, M0202L)
Chromatin Immunocleavage
- Pa-MNase (in-house production or Cell Signalin Technology, 40366)
- Cell Strainer, 70 µM (Corning, 431751)
- HEPES 1M pH 7.5 (Gibco, 15630080)
- NaCl 5 M (Thermo Fisher, AM9760G)
- Spermidine (Sigma, S2626-5G)
- Tween-20 (Sigma, P1379-100ML)
- Protease Inhibitor (Roche, 5056489001)
- EDTA (Invitrogen, 15575020)
FACS and single-cell processing
- NP-40 (Thermo Fisher, 85124)
- 70 µM cell strainers (Corning, CLS431751)
- EGTA (Thermo Fisher, 15425795)
- Klenow large fragment (NEB, M0210L)
- T4 PNK (NEB, M0201L)
- dNTPs (Promega, U1515)
- ATP (Part of Thermo Fisher Scientific, R0441 or equivalent)
- MgCl2 (part of Thermo Fisher, 4398828 or equivalent)
- PEG8000 50% (Promega, V3011)
- PNK Buffer (NEB, B0201S)
- BS 20 ng/ml (NEB, B9000S)
- AmpliTaq 360 (Thermo Fisher Scientific, 4398828)
- dATP 100mM (part of Promega, U1335)
- KCl 1 M (Thermofisher, AM9640G)
- T4 ligase (NEB, M0202L)
- MgCl21 M (ThermoFisher, AM9530G)
- Tris 1 M pH 7.5 (ThermoFisher, 15567027)
- DTT 0.1M (Invitrogen, 15846582)
Plate pooling
- VBLOK200 (Click-Bio, CBVBLOK200-1)
Tet1 Reaction Buffers
- HEPES 1M pH 7.5 (Gibco, 15630080)
- NaCl 5 M (Thermo Fisher, AM9760G)
- Ammonium Iron(II) sulfate hexahydrate (Sigma-Aldrich, 09719-50G)
- L-Ascorbic acid (Sigma-Aldrich, 95210-50G)
- α-Ketoglutaric acid disodium salt hydrate (Sigma-Aldrich, K3752-5G)
- DTT 100 mM (Part of Invitrogen, 18064022 or equivalent)
- ATP 100 mM (Part of Thermo Fisher Scientific, R0441 or equivalent)
TAPS conversion
- Proteinase K 20 mg/ml (Ambion, AM2546)
- Oligo Clean & Concentrator kit (Zymo, D4060)
- Pyridine Borane (Sigma, 179752-5G)
- Sodium Acetate 3 M, titrated to ph 4.3 (Sigma-Aldrich, S2889-250G)
In-vitro transcription
- MegaScript T7 transcription kit (Invitrogen, AMB13345)
- Potassium Acetate, KoAc (Sigma-Aldrich, 95843-100ML-F)
- Magnesium Acetate, MgOAC (Sigma-Aldrich, 63052-100ML)
- Tris Acetate ()
- RNAClean XP beads (Beckman Coulter, A63987)
Sequencing library preparation
- dNTPs (Promega, U1515)
- SuperScript™ II Reverse Transcriptase (Invitrogen, 18064022)
- RNAseOUT (Invitrogen, 10777019)
- NEBNext Ultra II Q5 Master Mix2x (NEB, M0492L)
- Qubit High Sensitivity DNA kit (Thermo Fisher, Q32851)
- Agilent High Sensitivity DNA kit (Agilent, 5067-4626)
Troubleshooting
Safety warnings
Some of the reagents in this protocol such as Pyridine Borane are hazardous and highly toxic to humans. Make sure to adhere to safety guidelines and inform your local safety officer.
PA-MNase production
(Single-cell) Chromatin immunocleave requires expression of a pA-MNase fusion protein in bacteria. We refer to Zeller et al. (Nature, 2022) and Schmid et al. (Mol. Cell, 2004) for detailed protocols (see references for more information). However, pA-MNase is also available commercially through Cell Signaling Technology (cat. no 4036).
mTet1 Protein production
Tet-assissted bisulfite sequencing (TAPS) requires expression of the catalytic domain of mouse Tet1 (mTet1CD) in an eukaryotic expression system. We refer to the detailed protocol by Liu et al. (Nature Biotechnology, 2019) for more information. mTet1CD can also be purchased from Sigma Aldrich (S9797-25UG).
Preparation of Tet1 Reaction Buffer and Fe2+ solution
Prepare 1 M solutions of Ascorbic Acid and a-Ketoglutarate:
- a-Ketoglutarate: Dissolve 0.9503 g in 5 ml nuclease-free H2O
- Ascorbic Acid: Dissolve 0.8806 g in 5 ml nuclease-free H2O
Note
Always prepare fresh Ascorbic Acid and a-KG solutions when making TET1 reaction buffer. Make sure to keep solutions on ice and protect Ascorbic Acid from direct light!
Prepare 0.5 M Fe2+ Solution
- Dissolve 1.96 g of Iron(II) sulfate hexahydrate in 10 ml nuclease-free H2O
- Store Fe2+ solution in single-use aliquots at -80°C
Assemble TET1 reaction buffer (volumes are suggestions and can be scaled up or down)
| Component | Concentration | Volume | Conc. Final | |
| H2O | 3,165.5 µl | |||
| HEPES | 1 M | 835 µl | 167 mM | |
| NaCl | 5 M | 333 µl | 333 mM | |
| a-Ketoglutarate | 1 M | 16.7 µl | 3.3 mM | |
| L-Ascorbic Acid | 1 M | 33.3 µl | 6.67 mM | |
| ATP | 100 mM | 200 µl | 4 mM | |
| DTT | 100 mM | 416.5 µl | 8.33 mM | |
| Total | 5000 µl |
Note
Store reaction buffer at -80°C in single-use aliquots and do not keep for more than 3 months!
Preparation of methylated lambda phage spike-ins
Methylation of lambda phage DNA
- Assemble the reaction below on ice
- Incubate: 2 hours @ 37°C
- Add an additional 1 µl of SAM and 0.5 µl M.SssI
- Incubate: 2 hours @ 37°C
- Perform Ampure XP SPRI cleanup (bead-to-sample ratio = 1:1)
- Elute sample in 20 µl of nuclease-free H20
- Repeat reaction once with methylated sample as input (including top-up of SAM)
- Perform Ampure XP SPRI cleanup (bead-to-sample ratio = 1:1)
- Elute sample 20 µl of nuclease-free H20
| Component | Concentration | Volume (µl) | |
| H2O | to 50.0 | ||
| NEB Buffer 2 | 10x | 5.0 | |
| SAM | 32 mM | 1.0 | |
| M.SssI | 4,000 U/ml | 0.5 | |
| Lambda phage DNA | 1 µg* |
* amount has to be adjusted based on batch-dependent concentration
NlaIII digestion
- Assemble the reaction below on ice
- Note: Adapter should be added in a ratio of 10:1, calculate molarity based on Qubit measurement and assuming full NlaIII digestion
- Incubate: 2 hours @ 37°C -> 20 min @ 65°C -> hold @ 4°C
- Perform Ampure XP SPRI cleanup (bead-to-sample ratio = 1:1)
- Elute sample 20 µl of nuclease-free H20
- Measure concentration with dye-based method (e.g. Qubit)
| Component | Concentration | Volume (µl) | |
| H2O | 24.0 | ||
| CutSmart Buffer | 10x | 5.0 | |
| NlaIII | 10,000 U/ml | 1.0 | |
| Sample | 20.0 | ||
| Total | 50 |
Adapter ligation
- Assemble the reaction below on ice
- Note: Adapter should be added in a ratio of 10:1, calculate molarity based on Qubit measurement and assuming full NlaIII digestion
- Incubate: 20 min @ 20°C -> 10 min @ 65°C -> hold @ 4°C
- Perform 2 Ampure XP SPRI cleanups (bead-to-sample ratio 0.8:1)
- Elute fully methylated, adapter-ligated controls in nuclease-free water
- Prepare aliquot with a concentration of 7 pg/µl
| Component | Concentration | Volume (µl) | |
| H2O | to 50 | ||
| T4 DNA ligase buffer | 10x | 5.0 | |
| T4 DNA ligase | 400,000 U/ml | 2.5 | |
| Sample | 20.0 | ||
| Adapter | 10:1 | variable | |
| Total | 50.0 |
Note
Adapter ligation is necessary to amplify the spike-in DNA during in-vitro transcription!
Chromatin Immuno-Cleavage (ChIC)
Recipes for wash buffers used in Chromatin Immuno-cleavage (ChIC)
| Component | Wash Buffer 1 (WB1) | Wash Buffer 2 (WB2) | Wash Buffer 3 (WB3) | |
| H2O | to 50 ml | to 50 ml | ||
| HEPES | 20 mM | 20 mM | 20 mM | |
| NaCl | 150 mM | 150 mM | 150 mM | |
| Spermidine | 0.5 µM | 0.5 µM | 0.5 µM | |
| Tween-20 | 0.05% | 0.05% | 0.05% | |
| Protease Inh. | 1 tablet | 1 tablet | ||
| EDTA | 2mM |
Fixation and permeabilization
- Harvest cells and wash twice with PBS at room temperature (centrifuge 3 min, 500 g to pellet)
- Resuspend cells in 300 µl PBS per 106 cells on ice
- Add 700 µl of ice-cold absolute ethanol per 106 cells while vortexing gently (70% ethanol final)
- Cells are fixed for two hours at -20°C
- Wash cells twice with WB1 (see above)
- Resuspend cells in 500 µl WB1
- Transfer reaction to 0.5 ml protein lo-bind Eppendorf tubes
Note
Safe stopping point: Fixed cells can be stored in WB1 supplemented with 10% DMSO at -80°C for up to 6 weeks. After thawing, wash cells twice with WB1, then continue.
Incubation with primary antibody
- Add histone-specific antibody (see table below for antibodies used in publication, others need to be titrated)
- Incubate cells overnight at 4°C with gentle agitation (e.g. on a roller)
| Antibody | Manufacturer | Cat. No. | Concentration | |
| H3K9me3 | Abcam | ab8898 | 1:100 | |
| H3K36me3 | 1:2000 | |||
| H3K27me3 | NEB | 9733S | 1:200 |
pA-MNase binding and nuclear staining
- Wash cells once with 500 µl WB2
- Add pA-MNase to a final concentration of 3 ng/µl
- Add Hoechst 34580 to a final concentration of of 5 µg/ml
- Incubate 1 hour at 4°C with gentle agitation
Washing and straining
- Wash cells twice with WB2
- Resuspend cells in 500 µl of WB3
- Filter cells through a 70 µM strainer
- Transfer to FACS tubes
FACS
Prepare sorting plates
- Add 10 µl of sterile filtered mineral oil to each well of 384-well hard-shell plates
- Plates can be prepared in advance, sealed and kept for multiple months
Cell sorting
- Cells were sorted on a BD InfluxTM cell sorter
- Depending on the machine and application, Hoechst signal can be used to select cells in G1 phase and to avoid debris
- After sorting, centrifuge plates for 1 min at 2,000 g
Note
It is critical to spin plates immediately after FACS sorting!
Single-cell processing
General notes:
- Nanoliter dispension was performed with the Innovadyne Nanodrop II platform. However, euqivalent nanoliter dispenser such as the iDOT or Mantis can be used aswell
- Adapters were copied from a source plate using the TTP Labtech Mosquito HTS liquid handler
Note
After dispensing liquids into 384-well plates, makes sure to centrifuge plates for 1 minute at 2,000 g to fuse droplets!
MNase digestion and Protease K digest
- MNase digestion is initiated by dispensing 100 nl per well of WB3 supplemented with 2 mM CaCl2 per well
- Incubate 30 mins in thermocycler set to 4°C
- to stop digestion, dispense 100 nl per well of the mix below
- Incubate: 20 min @ 4°C -> 6 hrs @ 65°C -> 20 min @ 80°C -> hold at 4°C
| Component | Concentration | Per Well (nl) | |
| Ultrapure H2O | 67 | ||
| EGTA | 0.5 M | 8 | |
| NP-40 10% | 10 % | 15 | |
| Protease K | 20 mg/ml | 10 | |
| Total | 100 |
Fragment blunting
- Dispense 150 nl per well of the mix below (350 nl total at this point)
- Incubate: 30 min @ 37°C -> 20 min @ 75°C -> hold @ 4°C
| Component | Concentration | Per well (nl) | |
| Klenow large | 5,000 U/ml | 2.5 | |
| T4 PNK | 10,000 U/ml | 2.5 | |
| dNTPs | 100 mM | 6.0 | |
| ATP | 100 mM | 3.5 | |
| MgCl | 25 mM | 10.0 | |
| PEG8000 | 50% | 7.5 | |
| PNK Buffer | 10x | 35.0 | |
| BSA | 20 mg/ml | 1.8 | |
| Ultrapure H2O | 81.3 | ||
| Total | 150 |
A-tailing
- Dispense 150 nl per well of the mix below (500 nl total at this point)
- Incubate: 15 min @ 37°C -> 10 min @ 72°C -> hold @ 4°C
| Component | Concentration | Per well (nl) | |
| AmpliTaq | 1.0 | ||
| dATP | 100 mM | 1.0 | |
| KCl | 1 M | 25.0 | |
| PEG8000 | 50% | 7.5 | |
| BSA | 20 mg/ml | 0.8 | |
| Ultrapure H2O | 114.8 | ||
| Total | 150 |
Addition of adapters and ligation
- per well, add 50 nl of adapters from source plate (5 µM) using the Mosquito HTS
- dispense 150 nl of ligation mix per well (700 nl total at this point)
- Incubate: 20 min @ 4°C -> 16 hrs @ 16°C -> 10 min @ 65°C -> hold @ 4°C
| Component | Concentration | Per well (nl) | |
| T4 ligase | 400,000 U/ml | 25.0 | |
| MgCl2 | 1 M | 3.5 | |
| Tris ph 7.5 | 1 M | 10.5 | |
| DTT | 100 mM | 52.5 | |
| ATP | 100 mM | 3.5 | |
| PEG8000 | 50% | 10.0 | |
| BSA | 20 mg/ml | 1.0 | |
| Ultrapure H2O | 44.0 | ||
| Total | 150.0 |
Plate pooling
- Remove cover, attach 384-well plates upside-down to a VBLOK200 Reservoir
- Cover with parafilm
- Centrifuge 2 min @ 500 g at room temperature to collect liquid in reservoirs
- Transfer aqueous phase to fresh 1.5 ml DNA lo-bind Eppendorf tube
- Centrifuge 1 min @ 13,000 g, transfer aqueous phase to fresh tube
- Repeat centrifugation and transfer once
- Measure volume with pipette
- Perform Ampure XP SPRI bead clean up (bead-to-sample ratio = 0.8)
- resuspend in 19 µl nuclease-free water
- transfer sample to fresh 0.5 ml DNA lo-bind Eppendorf tube
TAPS conversion
- Assemble the following reaction on ice:
| Component | Volume | |
| Sample (pooled 384-well plate) | 19 µl | |
| Methylated spike-in | 1 µl | |
| Tet1 reaction buffer | 15 µl | |
| Fe2+ solution (1.5 mM!) | 3.33 µl | |
| mTET1 enzyme | 6 to 12 µl | |
| H2O | to 50 µl |
Note
Fe2+ stock solution (0.5 M) has to be diluted 1:333 before use!
- Incubate for 80 min @ 37°C
- Add 1 µl of Proteinase K
- Vortex and centrifuge briefly to collect liquid
- Incubate 15 min @ 55°C
- Perform a 2x Ampure XP SPRI cleanup
- Option 1: to repeat Tet1 oxidation, elute in 20 µl nuclease-free water and repeat above reaction
- Option 2: to continue to Pyridine Borane incubation, elute in 33.75 µl nuclease-free H20 and transfer volume to a fresh 1.5 ml Eppendorf tube
Note
Sodium Acetate (NaAC) has to be titrated to a pH of 4.3!
Assemble Pyridine Borane reaction at room temperature:
| Component | Concentration | Volume | Final | |
| Sample | 33.75 µl | |||
| NaAc pH 4.3 | 3 M | 10 µl | 0.6 M | |
| Pyridine Borane | 8 M | 6.25 µl | 1 M | |
| Total | 50 µl |
- Incubate 16 hours @ 37°C in thermal shaker set to 850 rpm
Safety information
Warning: Pyridine Borane is highly toxic! Make sure to comply with local safety guidelines when following this protocol!
- Use Zymo Oligo Clean & Concentrator kit to clean up reactions after pyridine borane incubation
- Elute DNA with 15 µl of nuclease-free H2O heated 60°C
- To maximize DNA retrieval, repeat elution once (final volume ~ 30 µl)
- Reduce volume to 9.6 µl
- Option 1: Incubate in speed-vac, check volume regularly to prevent over-drying
- Option 2: Peform 1x Ampure XP SPRI clean-up, elute in 9.6 µl nuclease-free H2O
In-vitro transcription (IVT)
Assemble IVT reaction (all reagents are part of MegaScriptTM T7 transcription kit):
| Component | Volume | |
| Sample | 9.6 µl | |
| IVT Buffer | 2.4 µl | |
| Nucleotides (A/C/U/G) | 2.4 µl each | |
| T7 Enzyme | 2.4 µl | |
| Total: | 24 µl |
- Incubate 14 hours @ 37°C (with lid set to 70°C)
- To each reaction, add 6 µl of nuclease-free H2O and Turbo DNAse (part of MegaScriptTM T7 transcription kit)
- Incubate 15 min @ 37°C
- Add 7.88 µl of RNA fragmentation Buffer (200 mM Tris-Acetate, 500 mM KaOAc, 150 mM MgOAc)
- Incubate samples 90s at 94°C, immediately chill on ice
- Add 4.13 µl of 0.5 M EDTA to capture Mg2+
- Clean up samples with 34 µl (0.8x) of RNAClean XP beads
- Elute in 6 µl of nuclease-free H2O
Note
Run 1 µl of amplified RNA (aRNA) on a Bioanalyzer to assess quality and concentration.
Fig 1: aRNA Bioanalyzer trace from a successful experiment
Preparation of sequencing libraries
In a 0.5 ml DNA lo-bind tube, combine:
- 5 µl aRNA
- 0.5 µl of 10 mM dNTP solution
- 1 µl random hexamer RT primer 20 µM
- Heat samples to 65°C for 5 minutes
- Immediately chill samples on ice
- Assemble the reaction below
- Incubate: 10 min @ 25°C -> 60 min @ 42°C -> hold @ 4°C
| Component | Concentration | Volume (µl) | |
| Primed aRNA | 6.5 | ||
| First Strand Buffer | 5x | 2.0 | |
| DTT | 0.1 M | 1.0 | |
| SuperScript II | 200 U/µl | 0.5 | |
| Total | 10.0 |
- Assemble the reaction below
- Incubate: 30 s @ 98°C -> 10 - 13 x (10 s @ 98°C, 30 s @ 60°C, 30 s @ 72°C) -> 10 min @ 72°C -> hold @ 4°C
- Perform two AMPure XP SPRI bead cleanups (bead-to-sample-ratio 0.8)
- Elute amplified sequencing library in 15 µl of nuclease-free H2O
| Component | Concentration | Volume (µl) | |
| cDNA | 10.0 | ||
| Barcoded RPIX primer | 10 µM | 2.0 | |
| RP1 primer | 10 µM | 2.0 | |
| Ultra Q5 Master Mix | 2x | 25.0 | |
| Ultrapure H2O | 11.0 | ||
| Total | 50 |
Note
Adjust PCR cycles based on aRNA yield. In general, successful experiments should require less than 15 amplification cycles.
Sequencing library QC
- Measure concentration with a dye-based method such as Qubit
- Run 1 µl of library on Agilent High Sensitivity Bioanalzyer to assess size distribution
- Perform Illumina sequencing according to manufacturers protocol
DNA sequences
Barcoded single-cell Adapters
Example Top strand
5’-GGTGATGCCGGTAATACGACTCACTATAGGGAGTTCTACAGTCCGACGATCNNNACACACTAT
Example Bottom strand
5’-/5Phos/TAGTGTGTNNNGATCGTCGGACTGTAGAACTCCCTATAGTGAGTCGTATTACCGGCGAGCTT
Sequence features: Fork -> T7 promoter (bold) -> RA5 Illumina sequence -> 3 bp UMI (NNN) -> single-cell barcode (bold & italic) -> single-base T overhang
Library Amplification
RandomhexamerRT primer
GCCTTGGCACCCGAGAATTCCANNNNNN
Barcoded RPIX primer
5’-CAAGCAGAAGACGGCATACGAGAT-[6bp]-GTGACTGGAGTTCCTTGGCACCCGAGAATTCCA
RP1 primer
5’-AATGATACGGCGACCACCGAGATCTACACGTTCAGAGTTCTACAGTCCGA
Note
Adapter sequences are available in Supplementary Materials of the scEpi2-Seq publication!
Protocol references
Schmid M, Durussel T, Laemmli UK. ChIC and ChEC; genomic mapping of chromatin proteins. Mol Cell. 2004 Oct 8;16(1):147-57. doi: 10.1016/j.molcel.2004.09.007. PMID: 15469830.
Zeller P, Yeung J, Viñas Gaza H, de Barbanson BA, Bhardwaj V, Florescu M, van der Linden R, van Oudenaarden A. Single-cell sortChIC identifies hierarchical chromatin dynamics during hematopoiesis. Nat Genet. 2023 Feb;55(2):333-345. doi: 10.1038/s41588-022-01260-3. Epub 2022 Dec 20. PMID: 36539617; PMCID: PMC9925381.
Liu Y, Siejka-Zielińska P, Velikova G, Bi Y, Yuan F, Tomkova M, Bai C, Chen L, Schuster-Böckler B, Song CX. Bisulfite-free direct detection of 5-methylcytosine and 5-hydroxymethylcytosine at base resolution. Nat Biotechnol. 2019 Apr;37(4):424-429. doi: 10.1038/s41587-019-0041-2. Epub 2019 Feb 25. PMID: 30804537.
