Sep 24, 2025
Version 2
Shotgun Metagenomic Sequencing of Bacterial Enrichments on the Oxford Nanopore Platform Using the R10 flow cell and Ligation sequencing Kit V.2
Version 1 is forked from Nanopore Library Preparation for R10 Ligation Sequencing Kit
- Christopher Duda1,
- Padmini Ramachandran1
- 1FDA
- GenomeTrakrTech. support email: [email protected]

Protocol Citation: Christopher Duda, Padmini Ramachandran 2025. Shotgun Metagenomic Sequencing of Bacterial Enrichments on the Oxford Nanopore Platform Using the R10 flow cell and Ligation sequencing Kit. protocols.io https://dx.doi.org/10.17504/protocols.io.81wgbkjqqgpk/v2Version created by Christopher Duda
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: September 24, 2025
Last Modified: September 24, 2025
Protocol Integer ID: 228107
Keywords: metagenomics, libraries, long read, enrichments, nanopore, metagenomic sequencing of bacterial enrichment, oxford nanopore platform, amplicon libraries from dna, using nanopore, metagenomic sequencing, selective bacterial enrichment, bacterial enrichment, sequencing kit, multiplexed amplicon library, r10 flow cell, using ont v14 chemistry
Abstract
This protocol produces multiplexed amplicon libraries from DNA extracted from general and selective bacterial enrichments using Nanopore's Ligation sequencing kit, suitable for sequencing on Oxford Nanopore Technologies‱ (ONT) MinION/Gridion systems using ONT V14 chemistry (SQK-NBD114-96).
Guidelines
Overview
This protocol provides guidelines for multiplexing up to 16 bacterial enrichments on a single R10 flow cell, or up to 48 single-colony isolate DNA libraries on one flow cell. The sequencing depth generated is insufficient to close isolate genomes or to recover high-coverage metagenome-assembled genomes. However, the reads obtained are suitable for applications such as serotyping using genome indexing approaches.
Materials
| A | B | C | D | E | |
| Component | Acronym | Quantity | Color | Volume per vial | |
| Native barcode plate | NB01-96 | 3 | - | 8 µl per well | |
| DNA control sample | DCS | 3 | Yellow | 35 µl | |
| Native adapter | NA | 2 | Green | 40 µl | |
| Sequencing buffer | SB | 2 | Red | 700 µl | |
| Library beads | LIB | 2 | Pink | 600 µl | |
| Library solution | LIS | 2 | White | 600 µl | |
| Elution buffer | EB | 1 | Black | 1500 µl | |
| AMPure XP beads | AXP | 1 | Amber | 6000 µl | |
| Long fragment buffer | LFB | 1 | Orange | 7500 µl | |
| Short fragment buffer | SFB | 1 | Clear | 7500 µl | |
| EDTA | EDTA | 1 | Clear | 700 µl | |
| Flow cell flush | FCF | 1 | Blue | 15500 µl | |
| Flow cell tether | FCT | 2 | Purple | 200 µl |
Required Materials Not Included
- Nuclease-free water, molecular biology grade
- AMPure‱ XP beads (Beckman Coulter A63880) or equivalent
- 80% Ethanol (freshly prepared, molecular biology grade)
- DNA LoBind Tubes (Eppendorf‱ #022431021)
- Qubit‱ dsDNA HS Assay Kit (Thermo Fisher Scientific, Inc.‱ Q32851)
- Magnetic rack/stand (NEB #S1515, Alpaqua‱, cat. #A001322 or equivalent)
- Thermal cycler
- Vortex Mixer
- Microcentrifuge
- Agilent‱ Bioanalyzer‱ or similar fragment analyzer and associated consumables (#4150 or #4200 TapeStation System)
- DNase RNase free PCR strip tubes (USA Scientific 1402-1708)
- 1.5 ml tube magnet stand (NEB #S1506)
- NEB Blunt/TA Ligase Master Mix (NEB #M0367)
- NEBNext UltraII End repair/dA-tailing Module (NEB #E7546)
- NEBNext Quick Ligation Module (NEB #E6056)
- Bovine Serum Albumin (BSA)
Troubleshooting
Safety warnings
Please refer to Safety Data Sheets (SDS) for health and environmental hazards.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.
Before start
- Ensure you have sufficient DNA for your planned sequencing run.
- Decide on the number of samples to multiplex on a single flow cell.
- Thaw all NEB reagents and the Ligation Sequencing Kit components in advance, and confirm they are ready for use before starting the protocol.
- The library preparation here has some modifications from the original protocol in the barcoding step. Please read the step carefully before proceeding with barcoding.
Step 1: End-prep
1h
This section will take you from extracted DNA with a known concentration to a set of samples ready to be barcoded.
Before you start:
1. Start thawing on ice the reagents listed in step 3
2. You will need a significant amount of nuclease free water so make sure to have a bottle of it accessible.
Dilute DNA to 650 ng total, at 12 μL per sample
* If the concentration of DNA is too low, i.e. a negative number in column B, just use 12 μL of DNA.
The equation for this calculation is 650/DNA concentration.
The table will populate after values are entered! See attached Excel file!
For each sample, calculate a master mix of the following reagents per sample.
* Calculate for 2 extra samples to account for pipetting error. See attached Excel doc!
Add 3 μL of the above prepared master mix to each of your 650 ng/μL samples, pipetting up and down 10-20 times after each addition. Cover and spin down once all reagents are added.
Place the plate in a thermal cycler at 20 °C for 5 minutes, then 65 °C for 5 minutes.
Step 2: Barcoding and Cleanup
1h
On ice thaw:
- NEB Blunt/TA Ligase Master Mix
- EDTA blue cap tube(0.5M)
- Native Barcode (NB01-96)
At room Temperature thaw:
AMPure XP Beads (AXP). Vortex to mix.
Note: It is important to add all reagents in the order listed. Adding the Ligase Master Mix and the Native Barcode without end prepped DNA will cause the Barcode to bind to itself.
Do not vortex the sample at all during barcoding!
Barcoding happens in triplicate to maximize the number of quality read DNA fragments. To do this:
1. Get a new and separate 96 well PCR plate. (Unless you have 2 columns or less).
2. Thrice, take 3.75 μL from one sample and put it into a well on the separate plate. It should look like this:
Note: Do not use the first well as a sample for end prep and barcoding! Only use the 3 new ones from this step.

For each set of 3 end-prepped DNA samples, take from 1 well from a Native Barcode (NB01-96) plate.
Add 1.25 μL of barcode to each well, making sure each set of triplicates get the same barcode, and only one barcode goes to each set of triplicates.
Lastly, add 5 μL of Blunt/TA Ligase Master Mix to each well. Then cover and spin down the plate.
Incubate the plate for 20 minutes at room temperature.
Add 2 μL EDTA (blue cap tube) to each well to stop the reaction.
Combine all samples into one 1.5 mL LoBind Eppendorf tube. There are 13 μL of sample in each well.
For example: In an 8-sample run, there should be 24 total wells with 13 μL of sample in each well, with a total pooled volume of 312 μL
Resuspend AXP by vortexing and add 0.4X of pooled sample volume to the pooled reaction, mix by pipetting.
Incubate on a hula mixer for 10 minutes at room temperature.
Make 5 mL 80% ethanol in in nuclease free water per sample tube.
Spin down the sample and pellet on a magnet for 5 minutes until the supernatant is clear and colorless. Pipette off the supernatant and discard.
Wash the bead with 700 μL of 80% ethanol by placing the pipette tip against the side of the Eppendorf tube above the pellet and slowly dispensing the ethanol. Remove the ethanol and discard.
Repeat Step 16.
Spin down and place the tube back on the magnetic rack. Pipette off any residual ethanol, allow the pellet to dry for ~30 seconds, but do not let the pellet dry to the point of cracking.
Remove the tube from the magnet and resuspend the pellet in 35 μL of nuclease free water.
Incubate for 10 minutes at 37 °C on a hula mixer on the lowest setting, rocking for 10 seconds every 2 minutes.
Pellet the beads on a magnet until the supernatant is clear and colorless. Remove the 35 μL of supernatant and place it into a 1.5 mL LoBind Eppendorf tube.
Use a Qubit fluorometer to quantify 1 μL of barcoded sample.
Step 3: Native Adapter Ligation and Cleanup
30m
Thaw all reagents in the following table then place on ice. They are found in the NEBNext Quick Ligation Reaction Module.
| Reagent | |
| Native Adatper (NA) | |
| NEBNext Quick Ligation Buffer (5X) | |
| Quick T4 DNA Ligase | |
| Elution Buffer (EB) |
*Do not vortex the Quick T4 DNA Ligase*
Thaw all reagents, then spin them down for 5 seconds. Mix all reagents by performing 10 full volume pipette mixes.
If the Buffer has precipitate in it, vortex it until the precipitate dissolves fully.
Mix the following reagents in a 1.5 mL LoBind tube in the following order, pipetting 10-20 times between addition of reagents:
| Reagent | Volume | |
| Pooled Barcode Sample | 30 μL | |
| Native Adapter (NA) | 5 μL | |
| NEBNext Quick Ligation Buffer (5X) | 10 μL | |
| Quick T4 DNA Ligase | 5 μL | |
| Total | 50 μL |
Mix the reaction by gently pipetting up and down and briefly spinning down.
Incubate the reaction for 20 mins at room temperature
Resuspend the AMPure XP beads by vortexing.
Add 20 μL well vortexed AMPure XP Beads (AXP) to the tube and mix by flicking the tube.
Incubate for 10 minutes on a Hula mixer at room temperature (25°C).
Spin down the sample and pellet on a magnet until the eluate is clear and colorless, for at least 1 minute. Keep the tube on the magnet and pipette off the supernatant.
Wash the beads by adding 125 µL Long Fragment Buffer (LFB). Flick the beads to resuspend, spin down, and pellet the beads on a magnet until the eluate is clear and colorless, for at least 1 minute.
Remove the supernatant using a pipette and discard.
Repeat the previous step (step 31 &31.1).
Spin down and place the tube back on the magnet. Pipette off any residual supernatant. Allow to dry for 5~30 seconds. Do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend the pellet in 15 µL Elution Buffer (EB). Spin down and incubate for 10 minutes at 37°C.
Spin down the sample and pellet on a magnet until the eluate is clear and colorless, for at least 1 minute. Keep the tube on the magnet and retain 15 µl of eluate containing the DNA library into a clean 1.5 mL microtubes.
Use a Qubit fluorometer to quantify 1 μL of final library.
For short-term storage or reloading flow cells between washes, you can keep the prepared library at 4°C. While for long-term storage of more than 3 months, storing libraries at -80°C is recommended.
If you are sequencing this library within an hour, continue on to the next section where you will prime the nanopore flow cell.
Step 5: Flow cell check, Priming and loading the SpotON flow cell
Before priming the flow cell, load the flow cell onto the Gridion or minion, and complete the flow cell check. Assess the number of available pores. You need above 800 available pores for a successful run.
Thaw the flow cell, Sequencing Buffer (SQB), Library beads (LIB), BSA (not included in the kit), Flow cell flush, Flow cell Tether (FCT) at room temperature on cooling block.
Mix the flow cell priming mix with BSA in accordance with the table below:
| Reagent | Volume / flow cell | |
| Flow Cell Flush (FCF) | 1,170 μL | |
| 50 mg/mL Bovine Serum Albumin (BSA) | 5 μL | |
| Flow Cell Tether (FCT) | 30 μL | |
| Final Total Volume | 1,205 μL |
Open the MinION device lid and slide the flow cell under the clip. Press down firmly on the flow cell to ensure correct thermal and electrical contact.
Note
Make sure there are no bubbles in the flow cell.
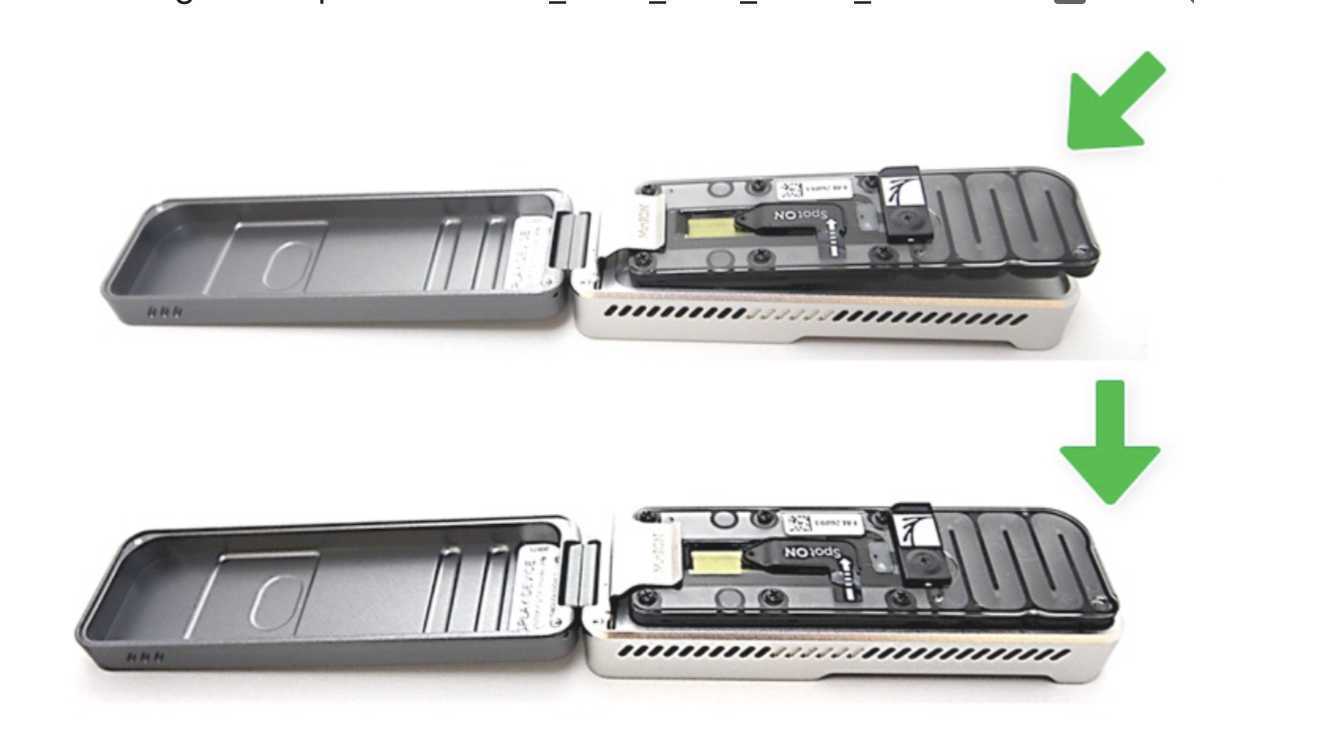

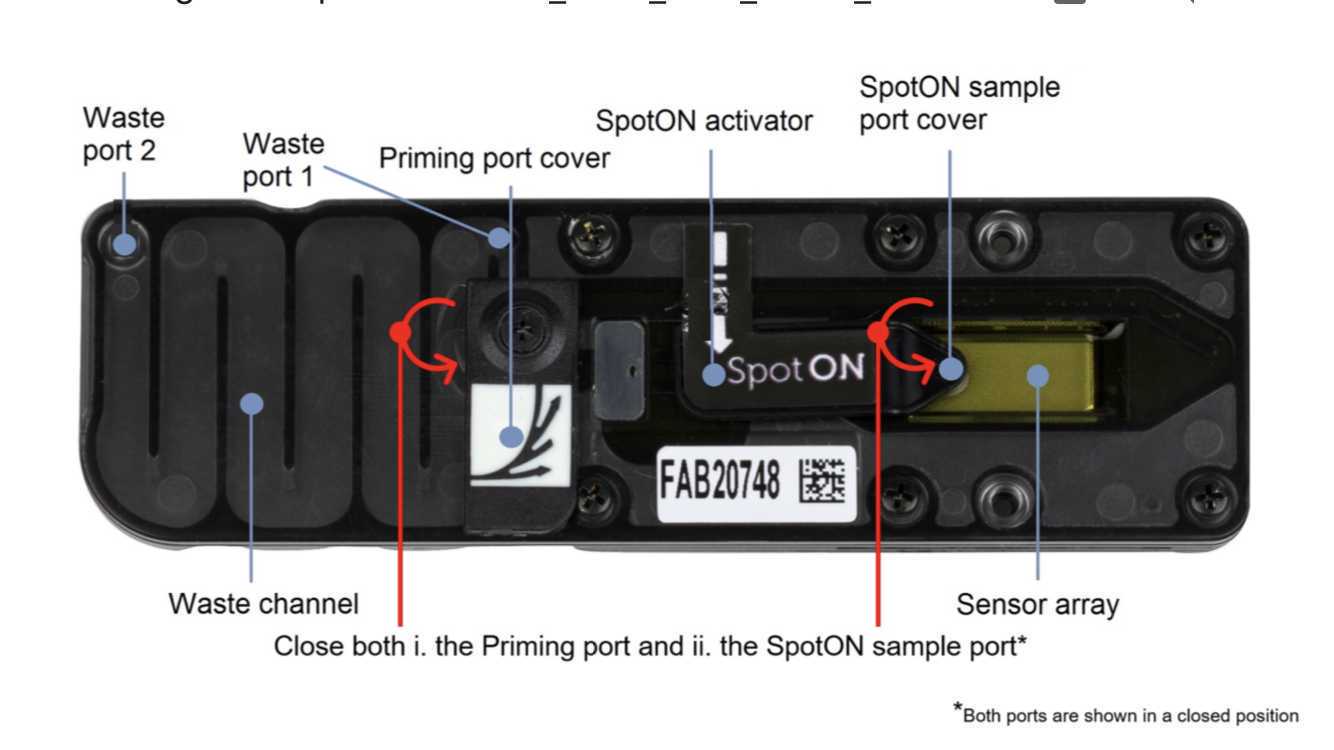
Slide the flow cell priming port cover clockwise to open the priming port.
After opening the priming port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
1. Set a P1000 pipette to 200 μl.
2. Insert the tip into the priming port.
3. Turn the wheel and aspirate fluid until the dial shows 220–230 µl, or until you can see a small volume of buffer entering the pipette tip.
Note
Visually check that there is continuous buffer from the priming port across the sensor array.
Load 800 μl of the flow cell priming mix into the flow cell via the priming port by turning the pipet wheel, avoiding the introduction of air bubbles. Wait for 5 minutes. During this time, prepare the library for loading by following the steps from step 43 and 44 below.
| Reagent | Volume | |
| Sequencing Buffer (SB) | 37.5 μl | |
| Library Beads (LIB) Mixed immediatly before use | 25.5 μl | |
| DNA Library | 12 μl | |
| Total | 75 μl |
Complete the flow cell priming:
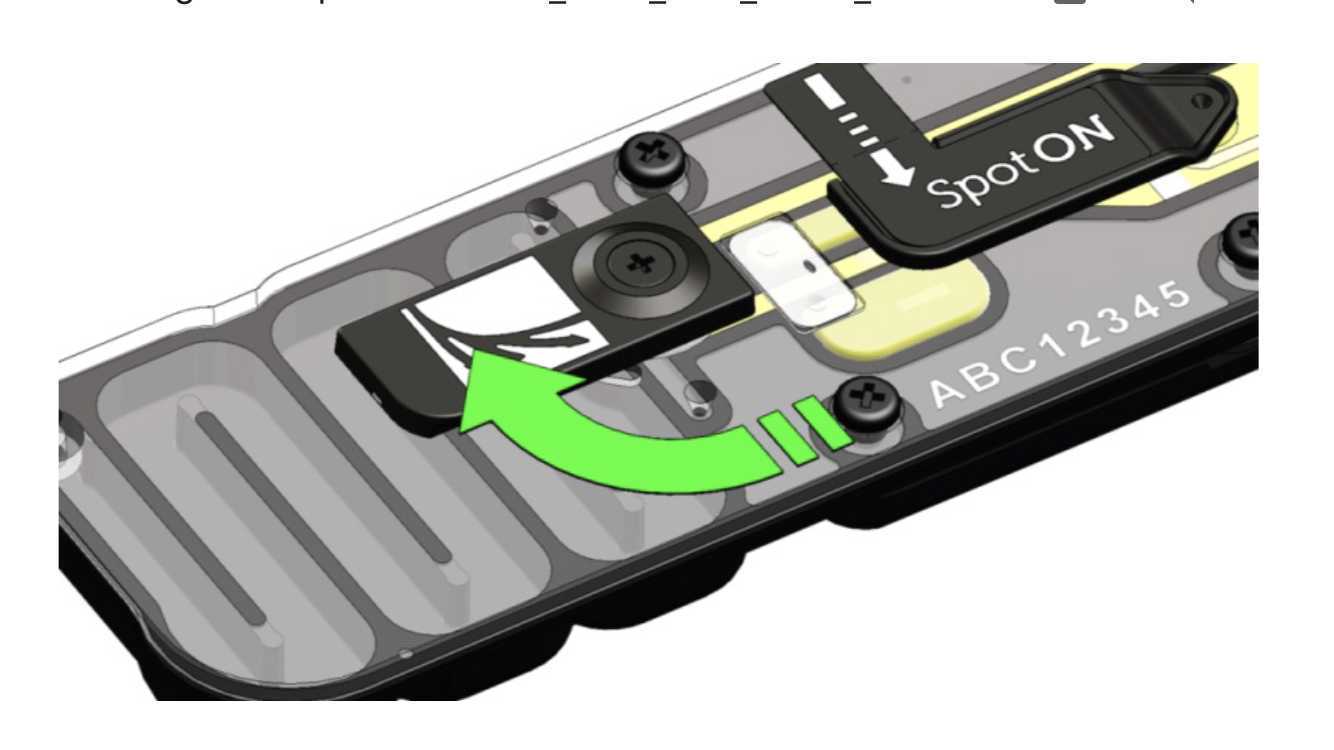
1. Gently lift the SpotON sample port cover to make the SpotON sample port accessible.
2. Load 200 μl of the priming mix into the flow cell via the priming port (not the SpotON sample port), avoiding the introduction of air bubbles.
Mix the prepared library gently by pipetting up and down in the tip using a P200 pipet just prior to loading.
Add the library to the flow cell via the SpotON sample port in a dropwise fashion. Ensure each drop flows into the port before adding the next.
Gently close the SpotON sample port cover, making sure the bung enters the SpotON port, close the priming port, cover the sensor array with the sensory cover, and close the MinION device lid.
Setting up parameters of sequencing
Name the experiment, sample id and library preparation kit (Native barcoding kit 114-96) and choose 48 hrs of sequencing while collecting fastqs every hour. Hit start sequencing.
