Oct 05, 2025
Sexing - General
- Tim Frasier1,
- Brenna Frasier2
- 1Saint Mary's University;
- 2Nova Scotia Museum

Protocol Citation: Tim Frasier, Brenna Frasier 2025. Sexing - General . protocols.io https://dx.doi.org/10.17504/protocols.io.q26g7n8o9lwz/v1
Manuscript citation:
Gilson A, Syvanen M, Levine K, Banks J (1998) Deer gender determination by polymerase chain reaction: Validation study and application to tissues, bloodstains, and hair forensic samples from California. California Fish and Game 84: 159-169.
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: July 28, 2025
Last Modified: October 05, 2025
Protocol Integer ID: 223455
Keywords: sexing, PCR, polymerase chain reaction, electrophoresis, ZFX, ZFY, SRY, primers, X chromosome, Y chromosome, mammal, cetacean, sex determination, sexing method, other for the sry gene, sry gene, results for male, sexing, results for female, pcr product, primer pair, agarose gel, zfx, range of mammalian species, mammalian species, female, male, gel, pcr, gene
Abstract
This sexing method is adapted from Gilson et al. (1998) and works well across a range of mammalian species. It involves the use of two primer pairs, one covering the ZFX/ZFY genes the other for the SRY gene. The PCR products are ~400bp long for the X chromosome and ~200bp for the Y chromosome fragment. When viewed on an agarose gel, results for females will show one band and results for males will be two bands.
Guidelines
If opening a new reagent or starting PCR for the first time, aliquots of each reagent should be made to decrease the number of freeze thaw cycles and to avoid contaminating whole stock bottles. Mix reagents well before aliquoting. Label each aliquot with a number and keep track of which aliquots have been previously used by putting a star on the lid and recording the aliquot number in your lab book.
Dilutions of primer stock should also be tracked to reduce freeze thaw cycles. If you see primer dilutions getting low, make more. Be sure to check the stock concentration to ensure you are correctly diluting to a concentration of 10 µm using TE.
A dNTP mix is created by combining each of the 4 bases to a concentration of 2mM each. This mix is what is used in the PCR.
Materials
Most of the supplies needed for this protocol are general/generic lab supplies. However, a few key materials for which it is important to know the details are below.
| Materials | Supplier | Catalogue # | |
| BSA (non acetylated) | Invitrogen | AM2616 | |
| dNTPs kit | Invitrogen | 10297018 | |
| GoTaq Flexi DNA Polymerase kit | Promega | M8295 | |
| Low Mass Ladder | Invitrogen | 10068013 | |
| HPLC water | Fisher Scientific | 270733-4L |
Supplies list
Primer sequences are listed below:
For ZFX/ZFY region (Originally from Aason & Medrano 1990)
P1-5EZ: 5' - ATA ATC ACA TGG AGA GCC ACA AGC T - 3'
P2-3EZ: 5' - GCA CTT CTT TGG TAT CTG AGA AAG T - 3'
For SRY region (Originally from Fain & LeMay 1995)
Y53-3C: 5' - CCC ATG AAC GCA TTC ATT GTG TGG - 3'
Y53-3D: 5' - ATT TTA GCC TTC CGA GGT CGA TA -3'
Troubleshooting
Safety warnings
Ethidium bromide is a hazardous chemical and should be used with caution. Read the SDS before using. Proper PPE, including a lab coat, gloves, long pants, and closed toe shoes should be worn while working with this chemical. Always be aware of equipment that has been contaminated and ensure it does not leave the ethidium bromide contaminated bench. Any ethidium bromide contaminated waste should be disposed of properly.
Before start
Note that this protocol has been optimized for use with biopsy samples. Samples from dead cetaceans may not work well with this method depending on quality and DNA degradation. See the protocol "Sexing - short fragment (cetacean only)" when working with samples from dead cetaceans.
Preparing sample concentrations for PCR
Whether to make a dilution or run straight DNA stock for PCR is decided based on Qubit readings. Generally, 10 ng of DNA is used (2µl of a 5 ng/µl dilution), however, if samples are already low concentration then the stock itself may be used. Use the flow chart below to determine what concentration of DNA you should use.
Flow chart for deciding how to prepare samples for sexing PCR.
Along with your samples of interest, you will also need to include known male and female controls from your project for verification of sex and functionality.
Sexing PCR
Pull out all aliquots of PCR reagents needed from the freezer:
- Reagent water (HPLC water)
- 5X Buffer
- dNTPs
- BSA
- MgCl2
- Primers - Y53-3C, Y53-3D, P1-5EZ, P2-3EZ
- DNA (samples and 2 controls – 1 male 1 female)
Allow reagents to thaw and then mix well.
Calculate the volume of reagents to add using the C1V1=C2V2 formula based on stock concentration, desired concentration, and number of samples. See the table below for an example. Record all calculations in your lab book. See Appendix for concentration justifications.
| Number of Samples | Volumes | |
| Controls: 2 | Total reactions = 30 | |
| PCR Negative: 1 | Each reaction volume = 10µl | |
| Samples: 24 | Total cocktail volume (“V2”) = 300µl | |
| Extras: 3 |
| Reagent | Initial Concentration ("C1") | Desired Concentration ("C2") | Volume to Add ("V1") | |
| Buffer | 5X | 1X | 60 µl | |
| dNTPs | 2 mM (each) | 0.2 mM (each) | 30 µl | |
| BSA | 3 µg/µl | 0.4 µg/µl | 40 µl | |
| MgCl2 | 25 mM | 1.5 mM | 18 µl | |
| Y53-3C | 10 µm | 0.3 µm | 9 µl | |
| Y53-3D | 10 µm | 0.3 µm | 9 µl | |
| P1-5EZ | 10 µm | 0.3 µm | 9 µl | |
| P2-3EZ | 10 µm | 0.3 µm | 9 µl | |
| Taq | 5 U/µl | 0.05 U/ | 3 µl | |
| DNA | 5 ng/μl | 2 µl/rxn | 60 µl | |
| ddH2O | - | To volume (300 μl) | 5 μl |
Example table of calculating volumes of reagents needed for the number of samples.
Example of C1V1=C2V2 calculation for 5X buffer:
To make the PCR cocktail, mix together the water, buffer, dNTPs, BSA, MgCl2, primers, and Taq. Add the Taq last and only right before adding to the samples.
Add 8 µL of cocktail to the 2 µL of DNA for each sample.
If using PCR tubes and not a plate, distribute the tubes across the PCR machine to distribute the weight and ensure even temperature.
Set up the PCR machine to run the following PROPDNR cycle conditions.
| Temperature | Time | Cycles | |
| 94°C | 5 minutes | 1 | |
| 94°C | 30 seconds | 30 | |
| 55°C | 1 minute | ||
| 72°C | 1 minute | ||
| 72°C | 10 minutes | 1 |
PCR cycle conditions for sexing PCR
Record the name of the PCR machine used in your lab book.
Making and Running Gel
Refer to the agarose gel protocol for how to make the 1.5% agarose gel.
Add 3 µL of Orange G dye to the 10 µL of PCR product. The ratio of dye:sample should always be >1:5 in order to make the solution dense enough to sink into the well.
Load 10 µL of PCR product and orange G mix into gel wells and run.
Visualize the gel under a UV box and take a picture of the gel for your records.
Assessing Results
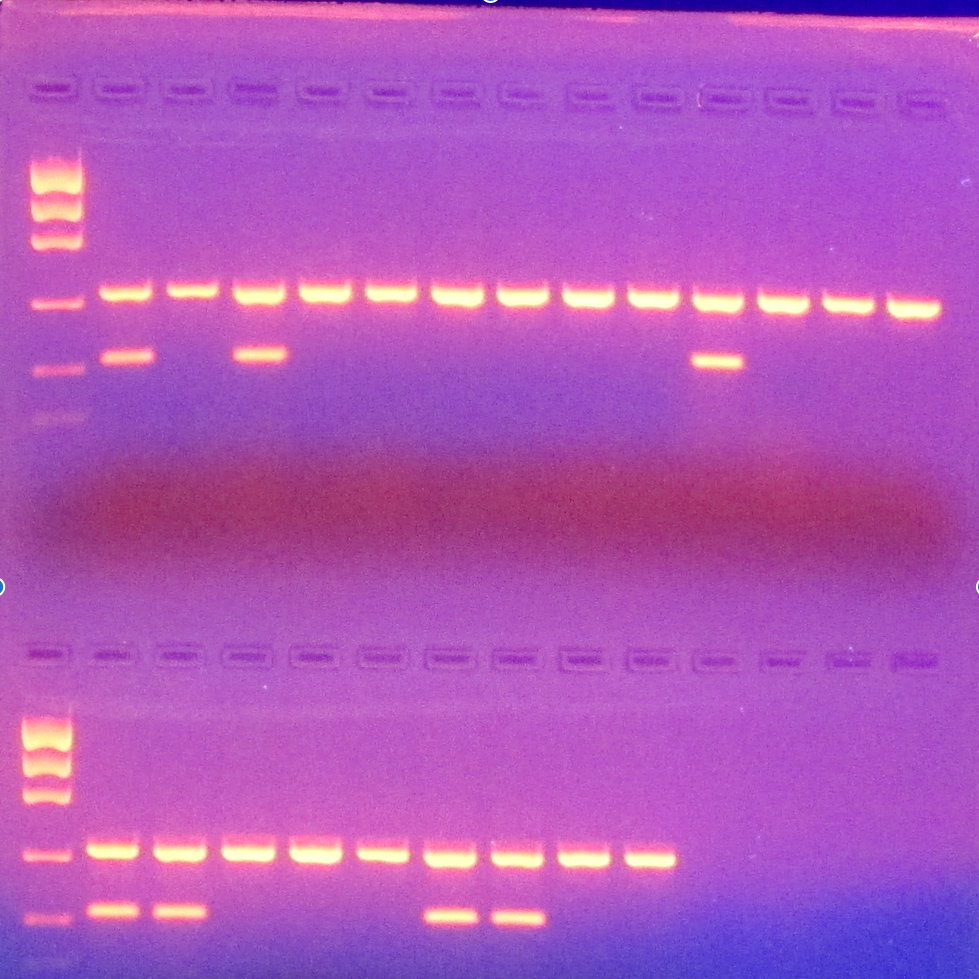
Print the gel image and paste/tape it into your lab book. Write the appropriate sample ID over each well as well as the sexing result. If there are two bands (one at ~400 bp and another ~200 bp) that is a male. If there is only one band ~400 bp then it is female. Check your positive and negative controls to make sure your reaction worked and there was no contamination.
Example of sexing gel image. The samples with one band are a female and the samples with two bands are male.
Save sexing gel to appropriate directory on the lab computer (associated with a particular project)
Enter sexes into the database, including the sexing gel information.
Appendix - Justification for PCR concentrations
As an example we will use a 10µL reaction to show the justification for each reagent concentration.
DNA - The average weight of the chromosomes in a single human cell (and therefore representing two copies of each region) is ~6.46 picograms (pg) (Piovesan et al. 2019). We use 10 ng of DNA in our sexing reactions, which therefore represents:
(0.00646 ng / 1 cell) = (10 ng / x cells)
x ~ 1,548 cells or ~3,000 copies.
In theory, the number of copies will double with each cycle of PCR, starting in cycle 3, resulting in a total of about ~1x1012 copies at the end of 30 cycles.
Primers - Because primers get used up during PCR (i.e., they become part of the newly synthesized fragments of DNA), they need to be added in excess compared to the amount of desired PCR products. Therefore, there needs to be more than 1x1012 molecules. Using Avogadro's number of 6.022x1023 molecules/mole, this means that we need to add:
(6.022x1023 molecules / 1 mole) = (1x1012 molecules / x moles)
x = 1.66x10-12 moles of each primer.
If our reaction volume is 10 µl, then this means that we need our primers at a concentration of at least:
1.66x10-12 moles / 0.00001 L = 0.000000166 molar (M), or 0.166 µM
We double this amount to ensure primers aren't limiting, therefore we add 0.3µM.
dNTPs - Similar to primers, dNTPS get used up in the reaction, and we therefore need to plan accordingly. Let’s assume the region is ~1,000 bp long. Each amplicon then requires 1,000 molecules of dNTPs. Therefore, we need 1,000 X more dNTP molecules than DNA (or primer) molecules.
1x1012 molecules X 1000 = 1x1015 molecules
Using Avogadro's number:
(6.022x1023 molecules / 1 mole) = (1x1015 molecules / x moles)
x = is 1.66x10-9 moles. In a 10 µl reaction 1.66x10-9 moles is:
1.66x10-9 moles / 0.00001 L = 0.000166 molar (M), or 0.166 mM
We use a bit more than this to ensure dNTPs aren’t limiting, therefore we add 0.2 mM each. Note that the actual concentration of all dNTPs combined is 4 times this (0.8 mM).
MgCl2 - Each dNTP will bind one magnesium ion. Based on this alone, we need to add at least 0.8 mM of MgCl2. Taq also requires MgCl2 as a co-factor. Lastly, magnesium is needed to counteract the EDTA in TE0.1, the solution in which our DNA is stored. Therefore, we add double what is needed just for the dNTPs (0.8 mM) to ensure that it is not a limiting factor, so 1.5 mM is used.
Taq Polymerase - 1 unit (U) of Taq will incorporate 10 nmol of dNTPs within 30 min at 72°C. One unit of Taq is ~4x1012 molecules (Sambrook & Russell 2001). From above, we have a total of 1.66x10-9 moles of dNTPS, which is 1.66 nanomoles (nm). We therefore only need about 0.2 units of Taq polymerase in our reaction. However, we use more than double this amount (0.5 U) to ensure it is not a limiting factor. In a 10 µl reaction this translates to 0.05 U/µl for a total of 0.5 U.
Protocol references
Aasen E, Medrano JF (1990) Amplification of the ZFY and ZFX genes for sex identification in humans, cattle, sheep and goats. Biotechniques 8: 1279-1281.
Fain SR, LeMay JP (1995) Gender identification of humans and mammalian wildlife
species from PCR amplified sex linked genes. Proceedings of the American Academy of
Forensic Science 1: 34.
Gilson A, Syvanen M, Levine K, Banks J (1998) Deer gender determination by polymerase chain reaction: Validation study and application to tissue, bloodstains, and hair forensic samples from California. California Fish and Game, 84: 159-169.
Piovesan A, Pelleri MC, Antonaros F, Strippoli P, Caracausi M, Vitale L (2019) On the length, weight and GC content of the human genome. BMC Research Notes, 12: 106.
Sambrook, J., & Russell, D.W. (2001) Molecular Cloning: A Laboratory Manual. 3rd Edition, Vol. 1, Cold Spring Harbor Laboratory Press, New York.
