Jan 07, 2026
Segmentation of motor pools in cleared spinal cord
- John-Paul Fuller-Jackson1,
- Peregrine B Osborne1,
- Janet R Keast1
- 1University of Melbourne
- SPARCTech. support email: [email protected]

Protocol Citation: John-Paul Fuller-Jackson, Peregrine B Osborne, Janet R Keast 2026. Segmentation of motor pools in cleared spinal cord . protocols.io https://dx.doi.org/10.17504/protocols.io.yxmvmenkbg3p/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: May 02, 2024
Last Modified: January 07, 2026
Protocol Integer ID: 99160
Keywords: spinal cord, imaris, motor neurons, motor pools, somatic motor neurons, motor columns, autonomic, rat, light sheet microscopy, 3d segmentation of motoneuron pool, spinal cord cytoarchitecture, intact spinal cord of the rat, segmentation of motor pool, motoneuron pool, cleared intact spinal cord, cholinergic neuron identification, cleared spinal cord, 3d segmentation, segmentation, microscopy, reference materials for motor pool identification, motor pool identification, motor pool, light sheet microscopy
Funders Acknowledgements:
NIH SPARC
Grant ID: 3OT2OD023872
Abstract
This protocol is for the 3D segmentation of motoneuron pools in large-volume images (light sheet microscopy) of the cleared intact spinal cord of the rat. Segmentation is based on cholinergic neuron identification and spinal cord cytoarchitecture, relying on reference materials for motor pool identification.
Citation
LINK
Citation
LINK
Materials
Software
Imaris File Converter
NAME
Bitplane
DEVELOPER
Software
Imaris Stitcher
NAME
Bitplane
DEVELOPER
Troubleshooting
Data preprocessing
Convert image stack (.OME-TIFF) to Imaris file type (.IMS) using Imaris File Converter.
Software
Imaris File Converter
NAME
Bitplane
DEVELOPER
For mosaic/tile scan acquisitions, stitch the individual .IMS files with Imaris Sticher.
Software
Imaris Stitcher
NAME
Bitplane
DEVELOPER
Spinal cord visualization and segment localization
Open the final, stitched .IMS file in Imaris.
Using the Orthogonal Slicer in Imaris, optimize the display properties of each relevant channel, focusing the slicer on the motor neurons of interest while doing so. This will typically involve changing the minimum and maximum display and increasing the gamma correction to 1.2 to 1.6.
Identify spinal cord segment boundaries with the following fiduciary markers:
- ventral and/or dorsal roots
- segment-specific cytoarchitecture
Mark these segment boundaries with transverse orientated Orthogonal Slicers.
Motor pool 3D segmentation
In Imaris, create a Surface by clicking on the blue Surface icon.
Click on the large button with the "X" symbol - "Skip automatic creation, edit manually"
In the Draw tab of the Surfaces, go to Contour sub-tab and select the Orientation of orthogonal slice on which contours will be drawn. The best resolution for light sheet images is always in XY, and was chosen for this example, which is the horizontal/longitudinal orientation relative to the spinal cord.
Move the orthogonal slice to the desired position in the spinal cord i.e. beginning of the motor pool to be segmented.
Click Draw to enable the creation of the contour on the orthogonal slice.
Create a polygonal contour outline on the orthogonal slice, clicking once to lay down each vertex. To close the contour, simply click on the starting point.
Repeat the drawing of contours multiple times at different levels, going from the start of the motor pool to the end. The frequency of contours required depends on how detailed the final segmentation needs to be. For exact outlines of neurons, a high number of contours is recommended. In our example, we traced loosely around the motor pool, and hence a low number of contours was required.
When all contours are drawn for the motor pool, click on Create Surface in the Contour sub-tab of the Draw tab to create the 3D surface.
Repeat Steps 6 to 13 for each motor pool or spinal cord nuclei to be segmented.
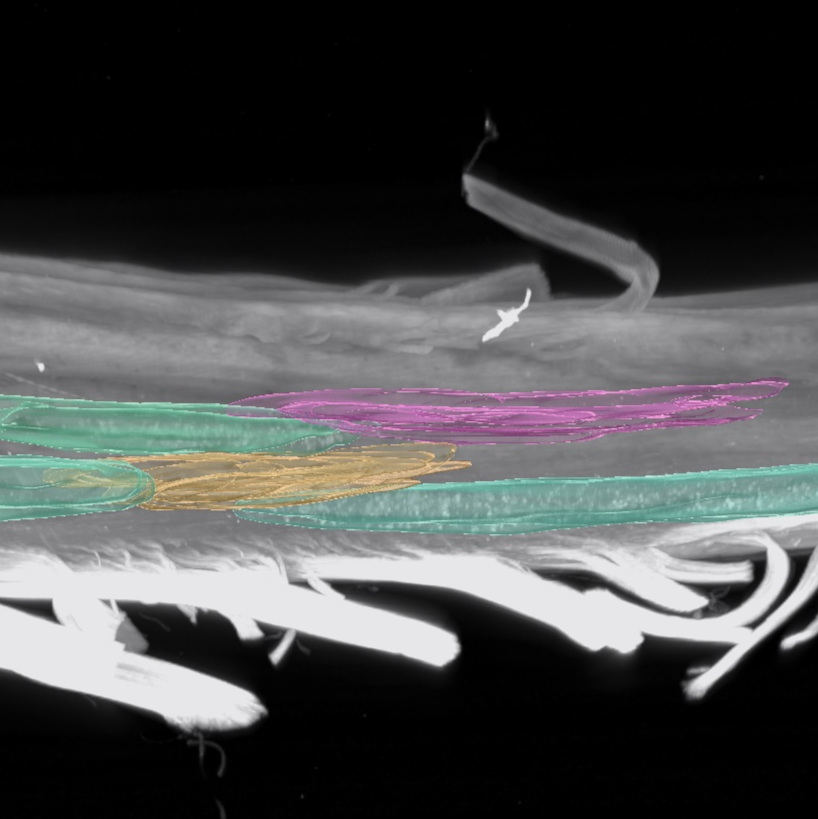
Example collection of segmented motor pools using Imaris Surfaces.
Protocol references
Citation
LINK
Citation
LINK
Citations
Charles Watson, George Paxinos, Gulgun Kayalioglu, Claire Heise. Chapter 15 - Atlas of the Rat Spinal Cord
https://doi.org/10.1016/B978-0-12-374247-6.50019-5Henrik Daa Schrøder. Organization of the motoneurons innervating the pelvic muscles of the male rat
https://doi.org/10.1002/cne.901920313