Jul 11, 2025
- Peter Rugg-Gunn1,
- Yang Wang1
- 1Babraham Institute
- Human Developmental Biology Initiative (HDBI)

Protocol Citation: Peter Rugg-Gunn, Yang Wang 2025. scMTR-seq. protocols.io https://dx.doi.org/10.17504/protocols.io.yxmvmb7dog3p/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: July 11, 2025
Last Modified: July 11, 2025
Protocol Integer ID: 222312
Keywords: single cell sequencing, single cell multiomics, scMTR-seq, histone modifications, transcription factors, CUT&Tag, CUT&RUN, ChIP-seq, transcriptomes in single cell, transcriptome in the same single cell, specific for each target histone modification, multiple histone modification, sequencing technology, target histone modification, other chromatin, scmtr, protocol for scmtr, applied scmtr, transcriptome, situ reverse transcription, histone modification, including pluripotent stem cell differentiation, pluripotent stem cell differentiation, single cell, same single cell, high cell recovery, nuclear mrna, mrna, indexed proteina, transcription
Funders Acknowledgements:
Wellcome
Grant ID: 215116/Z/18/Z
Wellcome
Grant ID: 225839/Z/22/Z
BBSRC
Grant ID: BBS/E/B/000C0522
BBSRC
Grant ID: BBS/E/B/000C0523
MRC
Grant ID: MR/T011769/1
MRC
Grant ID: MR/V02969X/1
Abstract
Here, we describe a protocol for scMTR-seq (single cell multi-targets and mRNA sequencing), which is a multi-omics sequencing technology that can simultaneously profile multiple histone modifications or other chromatin-bound proteins and transcriptome in the same single cells. scMTR-seq has high sensitivity, high cell recovery and is highly scalable in terms of starting material requirement. The protocol has the following steps: i) pre-assemble antibodies specific for each target histone modification or protein with indexed proteinA-Tn5-adapters; ii) perform in situ Tn5-mediated tagmentation with indexed complexes; iii) capture nuclear mRNA with a barcoded poly-T primer followed by in situ reverse transcription; iv) label single cells using split-pool combinatorial barcoding. We have applied scMTR-seq to profile histone modifications and transcriptomes in single cells from a broad range of heterogeneous samples including pluripotent stem cell differentiation and mouse pre-implantation embryos.
Attachments
Protocol materials
Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Roche Complete Protease Inhibitor EDTA-Free tablets Merck MilliporeSigma (Sigma-Aldrich)Catalog #5056489001
DPBS (no Ca, no Mg)ThermofisherCatalog #14190144
Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
Trypan Blue Solution, 0.4%Thermo FisherCatalog #15250061
16% Formaldehyde (w/v) Methanol-freeThermo Fisher ScientificCatalog #28906
Concanavalin-coated magnetic beads Bangs LaboratoriesCatalog #BP531
EDTA (0.5 M), pH 8.0, RNase-freeThermo FisherCatalog #AM9260G
Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
1M MgCl2Invitrogen - Thermo FisherCatalog #AM9530G
Anti-Histone H3 (acetyl K27) antibody - ChIP GradeAbcamCatalog #ab4729
Anti-Histone H3 (mono methyl K4) antibody - ChIP GradeAbcamCatalog #ab8895
Anti-Histone H3 (tri methyl K36) antibody - ChIP GradeAbcamCatalog #ab9050
Tri-Methyl-Histone H3 (Lys27) (C36B11) Rabbit mAbCell Signaling TechnologyCatalog #9733
Anti-Histone H3 (tri methyl K4) antibody - ChIP GradeAbcamCatalog #ab8580
Anti-Histone H3 (tri methyl K9) antibody - ChIP GradeAbcamCatalog #ab8898
Normal Rabbit IgG Control R&D SystemsCatalog #AB-105-C
PEG 6000 (Poly(ethylene glycol))Bio Basic Inc.Catalog #PB0432.SIZE.500g
dNTP Set 100 mM SolutionsThermo Fisher ScientificCatalog #R0182
Protector RNase InhibitorMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335399001
SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696
Maxima H Minus Reverse TranscriptaseThermo Fisher ScientificCatalog ##EP0741
T4 DNA Ligase - 100,000 unitsNew England BiolabsCatalog #M0202L
Protector RNase InhibitorMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335402001
10% Triton X-100Merck MilliporeSigma (Sigma-Aldrich)
SDS, 10% Solution, RNase-freeThermo FisherCatalog #AM9822
1M Tris-HCl (pH 8.0)Thermo Fisher ScientificCatalog #15568025
Proteinase KThermo Fisher ScientificCatalog #EO0491
Dynabeads MyOne Streptavidin C1Invitrogen - Thermo FisherCatalog #65001
PMSFMerck MilliporeSigma (Sigma-Aldrich)Catalog #P7626
Ficoll PM‐400Merck MilliporeSigma (Sigma-Aldrich)Catalog #F5415-50ML
HotStart ReadyMix (KAPA HiFi PCR kit)Kapa BiosystemsCatalog #KK2601
Ampure XP beads Beckman CoulterCatalog #A63881
NEBNext® High-Fidelity 2X PCR Master MixNew England BiolabsCatalog #M0541
GlycerolMerck MilliporeSigma (Sigma-Aldrich)Catalog #G5516
1 M Hydroxyethyl piperazineethanesulfonic acid pH 7.9 (HEPES (K ))Merck MilliporeSigma (Sigma-Aldrich)Catalog #H3375
1 M Potassium Chloride (KCl)Merck MilliporeSigma (Sigma-Aldrich)Catalog #P3911
2 M Spermidine Merck MilliporeSigma (Sigma-Aldrich)Catalog #S2501
PBS Thermo Fisher ScientificCatalog #28374
Glycine
1 M Hydroxyethyl piperazineethanesulfonic acid pH 7.5 (HEPES (Na ))Merck MilliporeSigma (Sigma-Aldrich)Catalog #H3375
5M NaClAmbionCatalog #AM9760G
1 M Calcium Chloride (CaCl2)Fisher ScientificCatalog #BP510
1 M Manganese Chloride (MnCl2)Merck MilliporeSigma (Sigma-Aldrich)Catalog #203734
Trizma® hydrochloride solutionMerck MilliporeSigma (Sigma-Aldrich)Catalog #T2319
NP-40 10% Merck MilliporeSigma (Sigma-Aldrich)Catalog #11332473001
10% Tween 20Bio-Rad LaboratoriesCatalog #1662404
Tris HCl Buffer 1M Solution, Sterile pH 7.5Bio Basic Inc.Catalog #SD8124.SIZE.450ml
Pierce™ Dimethylformamide (DMF), Sequencing gradeThermo Fisher ScientificCatalog #20672
5M NaCl solutionThermo Fisher ScientificCatalog #AM9759
DTTMerck MilliporeSigma (Sigma-Aldrich)Catalog #D0632
Troubleshooting
Solution Preparation
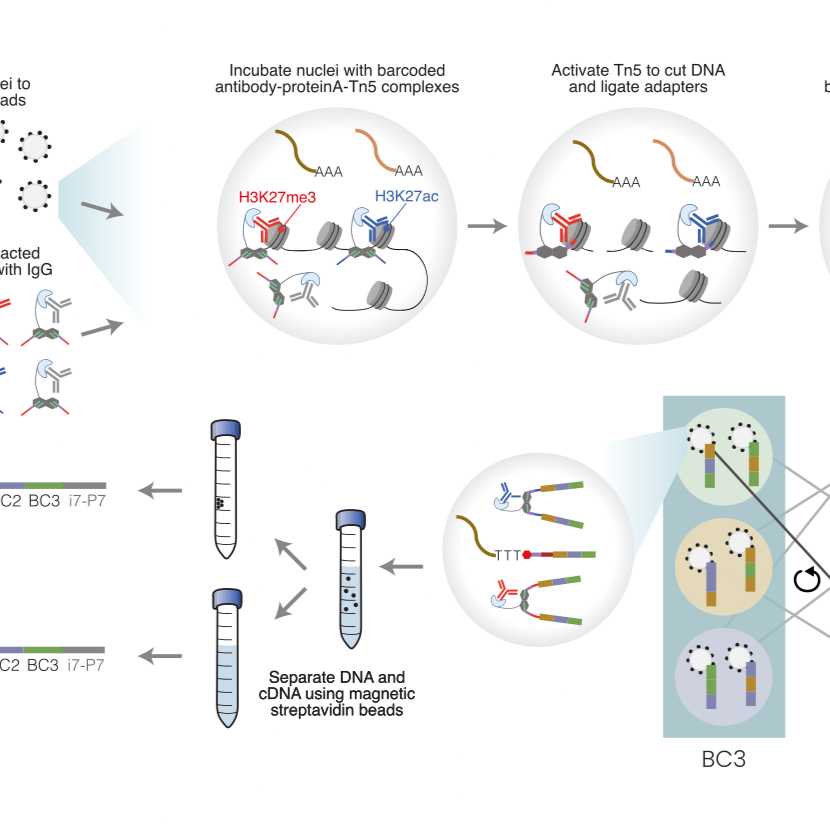
Overview of scMTR-seq
Solution Preparation
Protease Inhibitor Cocktail (PIC) (100x):
Dissolve 1 tablet of
Roche Complete Protease Inhibitor EDTA-Free tablets Merck MilliporeSigma (Sigma-Aldrich)Catalog #5056489001
in 500 µL Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932 , aliquot and store at -20 °C .
Avoid repeated freeze-thaw cycles
RNase Inhibitor (RI) mix (80x)
Mix:
- SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696
- Protector RNase InhibitorMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335399001
at a 1:1 ratio, and store at -20 °C
PBS with inhibitors (PBSI)
- 1x PIC
- 0.4x RI mix
- 0.04% BSA Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037 diluted in DPBS (no Ca, no Mg)ThermofisherCatalog #14190144
eg: 2 µL PIC + 1 µL RI + 1.06 µL BSA +196 µL DPBS
2x Nuclei Extraction (NE) buffer
| A | B | C | D | |
| Reagents | Stocking | Working 1x | 2x /μL | |
| HEPES(K+) pH 7.9 | 1M | 20 mM | 1000 | |
| KCl | 1M | 10 mM | 500 | |
| Spermidine | 2M | 0.5 mM | 12.5 | |
| Triton-X100 | 10% (vol/vol) | 0.1% | 500 | |
| Glycerol | 100% | 20% | 10000 | |
| PIC* | 100x | 2x | 500 | |
| RI* | 80x | 2x | 625 | |
| Nuclease-free water | 11862.5 | |||
| total | 25000 |
* Only add 2x PIC and 2x RI mix just before use.
Prior to these additions, the buffer can be stored at 4 °C for up to 3 months.
NB To achieve 40% glycerol in the 2x NE buffer, we typically add 12.5g of 100% glycerol.
- 1 M Hydroxyethyl piperazineethanesulfonic acid pH 7.9 (HEPES (K ))Merck MilliporeSigma (Sigma-Aldrich)Catalog #H3375
- 1 M Potassium Chloride (KCl)Merck MilliporeSigma (Sigma-Aldrich)Catalog #P3911
- 2 M Spermidine Merck MilliporeSigma (Sigma-Aldrich)Catalog #S2501
- 10% Triton X-100Merck MilliporeSigma (Sigma-Aldrich)
- GlycerolMerck MilliporeSigma (Sigma-Aldrich)Catalog #G5516
- PIC (100x)
- RI (80x)
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
5x Fixation quench buffer
| A | B | C | D | |
| Reagents | Stocking | Working 1x | /μL | |
| Glycine | 2.5M | 125 mM | 50 | |
| Tris-HCl pH 8.0 | 1M | 50 mM | 50 | |
| Triton-X100 | 10% | 0.1% (in buffer) | 2 | |
| BSA* | 7.5% | 0.1% | 13.4 | |
| PBS | 84.6 | |||
| total | 200 |
* Only add BSA just before use. Store 4 °C .
- Glycine
- 1M Tris-HCl (pH 8.0)Thermo Fisher ScientificCatalog #15568025
- 10% Triton X-100Merck MilliporeSigma (Sigma-Aldrich)
- Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- PBS Thermo Fisher ScientificCatalog #28374
Wash Buffer
| A | B | C | D | |
| Reagents | Stocking | Working 1x | /μL | |
| HEPES(Na+) pH 7.5 | 1 M | 20 mM | 1000 | |
| NaCl | 5 M | 150 mM | 1500 | |
| Spermidine | 2 M | 0.5 mM | 12.5 | |
| BSA* | 7.5% | 1% | 6666 | |
| PIC* | 100x | 1x | 500 | |
| RI* | 80x | 1x | 625 | |
| Nuclease-free water | 39696 | |||
| total | 50000 |
* Only add 1x PIC, 1x RI mix, and 1% BSA just before use.
Prior to these additions, buffer can be stored at 4 °C for up to 3 months.
- 1 M Hydroxyethyl piperazineethanesulfonic acid pH 7.5 (HEPES (Na ))Merck MilliporeSigma (Sigma-Aldrich)Catalog #H3375
- 5M NaClAmbionCatalog #AM9760G
- 2 M Spermidine Merck MilliporeSigma (Sigma-Aldrich)Catalog #S2501
- Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- PIC (100x)
- RI mix (80x)
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Binding Buffer
| A | B | C | D | |
| Reagents | Stocking | Working 1x | /μL | |
| HEPES (Na+) pH 7.5 | 1 M | 20 mM | 200 | |
| KCl | 1 M | 10 mM | 100 | |
| CaCl2 | 1 M | 1 mM | 10 | |
| MnCl2 | 1 M | 1 mM | 10 | |
| Nuclease-free water | 9680 | |||
| total | 10000 |
Store the buffer at 4 °C for up to 6 months
- 1 M Hydroxyethyl piperazineethanesulfonic acid pH 7.5 (HEPES (Na ))Merck MilliporeSigma (Sigma-Aldrich)Catalog #H3375
- 1 M Potassium Chloride (KCl)Merck MilliporeSigma (Sigma-Aldrich)Catalog #P3911
- 1 M Calcium Chloride (CaCl2)Fisher ScientificCatalog #BP510
- 1 M Manganese Chloride (MnCl2)Merck MilliporeSigma (Sigma-Aldrich)Catalog #203734
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Nuclei Isolation Buffer (NIB)
| A | B | C | D | E | |
| Reagents | Stocking | Working 1x | /μL | ||
| Trizma buffer pH 7.5 | 1 M | 10 mM | 500 | ||
| NaCl | 5 M | 10 mM | 100 | ||
| MgCl2 | 1 M | 3 mM | 150 | ||
| NP-40 | 10% | 0.1% | 500 | ||
| Nuclease-free water | 48750 | ||||
| total | 50000 |
Add PIC and RI mix just before use
- Trizma® hydrochloride solutionMerck MilliporeSigma (Sigma-Aldrich)Catalog #T2319
- Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
- 1M MgCl2Invitrogen - Thermo FisherCatalog #AM9530G
- NP-40 10% Merck MilliporeSigma (Sigma-Aldrich)Catalog #11332473001
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
STE buffer
| A | B | C | D | |
| Reagents | Stocking | Working 1x | /μL | |
| Tris pH 8.0 | 1 M | 10 mM | 500 | |
| NaCl | 5 M | 50 mM | 500 | |
| EDTA | 0.5 M | 1 mM | 100 | |
| Nuclease-free water | 48900 | |||
| total | 50000 |
- 1M Tris-HCl (pH 8.0)Thermo Fisher ScientificCatalog #15568025
- Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
- EDTA (0.5 M), pH 8.0, RNase-freeThermo FisherCatalog #AM9260G
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
1x B&W-T buffer
| A | B | C | D | |
| Reagents | Stocking | In buffer | /μL | |
| Tris pH 8.0 | 1M | 5 mM | 50 | |
| NaCl | 5M | 1 M | 2000 | |
| EDTA | 0.5 M | 0.5 mM | 10 | |
| Tween 20 | 10% | 0.05% | 50 | |
| Nuclease-free water | 7890 | |||
| total | 10000 |
- 1M Tris-HCl (pH 8.0)Thermo Fisher ScientificCatalog #15568025
- Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
- EDTA (0.5 M), pH 8.0, RNase-freeThermo FisherCatalog #AM9260G
- 10% Tween 20Bio-Rad LaboratoriesCatalog #1662404
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
2x Binding & Washing buffer
| A | B | C | D | |
| Reagents | Stocking | In buffer | /μL | |
| Tris pH 8.0 | 1M | 10 mM | 10 | |
| NaCl | 5M | 2 M | 400 | |
| EDTA | 0.5 M | 1 mM | 2 | |
| Nuclease-free water | 588 | |||
| total | 1000 |
Add Superase RI before use (1μL RI to 45μL 2x Binding & Washing buffer)
- 1M Tris-HCl (pH 8.0)Thermo Fisher ScientificCatalog #15568025
- Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
- EDTA (0.5 M), pH 8.0, RNase-freeThermo FisherCatalog #AM9260G
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
4x Tag Buffer for cDNA tagmentation
| A | B | C | D | |
| Reagents | Stocking | In Buffer | /μL | |
| Tris-HCl pH 7.5 | 1M | 40mM | 40 | |
| MgCl2 | 1M | 20mM | 20 | |
| Dimethylformamide (DMF) | 100% | 20% | 200 | |
| Nuclease-free water | 740 | |||
| total | 1000 |
- Tris HCl Buffer 1M Solution, Sterile pH 7.5Bio Basic Inc.Catalog #SD8124.SIZE.450ml
- 1M MgCl2Invitrogen - Thermo FisherCatalog #AM9530G
- Pierce™ Dimethylformamide (DMF), Sequencing gradeThermo Fisher ScientificCatalog #20672
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Tn5 dilution buffer
| A | B | C | D | |
| Reagents | Stocking | In Buffer | /uL | |
| Tris-HCl pH7.5 | 1M | 10mM | 10 | |
| NaCL | 5M | 100mM | 20 | |
| Glycerol | 100% | 50% (0.625g) | 500 | |
| DTT | 1mM | 0.1M | 10 | |
| Nuclease-free water | 460 | |||
| total | 1000 |
- Tris HCl Buffer 1M Solution, Sterile pH 7.5Bio Basic Inc.Catalog #SD8124.SIZE.450ml
- 5M NaCl solutionThermo Fisher ScientificCatalog #AM9759
- GlycerolMerck MilliporeSigma (Sigma-Aldrich)Catalog #G5516
- DTTMerck MilliporeSigma (Sigma-Aldrich)Catalog #D0632
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Prepare oligonucleotides for split pool barcoding
This section prepares the oligonucleotides that will add the sequence used to identify individual cells.
Example Annealed Barcodes & Linkers
Equipment:
- PCR machine (thermocycler)
- RNAse-free 96-well plates (Twin-Tec PRC plate, 96 Lobind, Semi-skirted)
Equipment
Eppendorf Twin tec PCR plates, 96 wells
NAME
96 well PCR plate
TYPE
Eppendorf
BRAND
EP0030128770
SKU
LINK
Primers
Linker Primers
| A | B | |
| Name | Sequence | |
| Linker_1 | CGAATGCTCTGGCCTCTCAAGCACGTGGAT | |
| Linker_2 | GGTCTGAGTTCGCACACGCTCCTCGAATCC | |
| Linker_3 | AGTCGTACGCCGATGCGAAACATCGGCCAC |
Barcode Primers
Split pool barcoding for cell identification, example primer sequence and index position:
BC1_#01
GTGCGAACTCAGACC CTGTAGCC ATCCACGTGCTTGAG
| Index |
BC1
| A | B | |
| Name | Sequence | |
| BC1_#01 | /5Phos/GTGCGAACTCAGACCAACGTGATATCCACGTGCTTGAG | |
| BC1_#02 | /5Phos/GTGCGAACTCAGACCAAACATCGATCCACGTGCTTGAG | |
| BC1_#03 | /5Phos/GTGCGAACTCAGACCACCACTGTATCCACGTGCTTGAG | |
| BC1_#04 | /5Phos/GTGCGAACTCAGACCCATCAAGTATCCACGTGCTTGAG | |
| BC1_#05 | /5Phos/GTGCGAACTCAGACCCTGTAGCCATCCACGTGCTTGAG | |
| BC1_#06 | /5Phos/GTGCGAACTCAGACCAACCGAGAATCCACGTGCTTGAG | |
| BC1_#07 | /5Phos/GTGCGAACTCAGACCAAGGTACAATCCACGTGCTTGAG | |
| BC1_#08 | /5Phos/GTGCGAACTCAGACCACAGCAGAATCCACGTGCTTGAG | |
| BC1_#09 | /5Phos/GTGCGAACTCAGACCACCTCCAAATCCACGTGCTTGAG | |
| BC1_#10 | /5Phos/GTGCGAACTCAGACCACGCTCGAATCCACGTGCTTGAG | |
| BC1_#11 | /5Phos/GTGCGAACTCAGACCACTATGCAATCCACGTGCTTGAG | |
| BC1_#12 | /5Phos/GTGCGAACTCAGACCAGAGTCAAATCCACGTGCTTGAG | |
| BC1_#13 | /5Phos/GTGCGAACTCAGACCAGATCGCAATCCACGTGCTTGAG | |
| BC1_#14 | /5Phos/GTGCGAACTCAGACCCAATGGAAATCCACGTGCTTGAG | |
| BC1_#15 | /5Phos/GTGCGAACTCAGACCCAGCGTTAATCCACGTGCTTGAG | |
| BC1_#16 | /5Phos/GTGCGAACTCAGACCCATACCAAATCCACGTGCTTGAG | |
| BC1_#17 | /5Phos/GTGCGAACTCAGACCCCAGTTCAATCCACGTGCTTGAG | |
| BC1_#18 | /5Phos/GTGCGAACTCAGACCCGAACTTAATCCACGTGCTTGAG | |
| BC1_#19 | /5Phos/GTGCGAACTCAGACCCGACTGGAATCCACGTGCTTGAG | |
| BC1_#20 | /5Phos/GTGCGAACTCAGACCCTCAATGAATCCACGTGCTTGAG | |
| BC1_#21 | /5Phos/GTGCGAACTCAGACCCTGAGCCAATCCACGTGCTTGAG | |
| BC1_#22 | /5Phos/GTGCGAACTCAGACCGAATCTGAATCCACGTGCTTGAG | |
| BC1_#23 | /5Phos/GTGCGAACTCAGACCGAGCTGAAATCCACGTGCTTGAG | |
| BC1_#24 | /5Phos/GTGCGAACTCAGACCGATAGACAATCCACGTGCTTGAG | |
| BC1_#25 | /5Phos/GTGCGAACTCAGACCGCTAACGAATCCACGTGCTTGAG | |
| BC1_#26 | /5Phos/GTGCGAACTCAGACCGCTCGGTAATCCACGTGCTTGAG | |
| BC1_#27 | /5Phos/GTGCGAACTCAGACCGGAGAACAATCCACGTGCTTGAG | |
| BC1_#28 | /5Phos/GTGCGAACTCAGACCGGTGCGAAATCCACGTGCTTGAG | |
| BC1_#29 | /5Phos/GTGCGAACTCAGACCGTACGCAAATCCACGTGCTTGAG | |
| BC1_#30 | /5Phos/GTGCGAACTCAGACCGTCGTAGAATCCACGTGCTTGAG | |
| BC1_#31 | /5Phos/GTGCGAACTCAGACCGTGTTCTAATCCACGTGCTTGAG | |
| BC1_#32 | /5Phos/GTGCGAACTCAGACCTCTTCACAATCCACGTGCTTGAG | |
| BC1_#33 | /5Phos/GTGCGAACTCAGACCTGGAACAAATCCACGTGCTTGAG | |
| BC1_#34 | /5Phos/GTGCGAACTCAGACCTGGCTTCAATCCACGTGCTTGAG | |
| BC1_#35 | /5Phos/GTGCGAACTCAGACCTGGTGGTAATCCACGTGCTTGAG | |
| BC1_#36 | /5Phos/GTGCGAACTCAGACCTTCACGCAATCCACGTGCTTGAG | |
| BC1_#37 | /5Phos/GTGCGAACTCAGACCAACTCACCATCCACGTGCTTGAG | |
| BC1_#38 | /5Phos/GTGCGAACTCAGACCAAGAGATCATCCACGTGCTTGAG | |
| BC1_#39 | /5Phos/GTGCGAACTCAGACCAAGGACACATCCACGTGCTTGAG | |
| BC1_#40 | /5Phos/GTGCGAACTCAGACCAATCCGTCATCCACGTGCTTGAG | |
| BC1_#41 | /5Phos/GTGCGAACTCAGACCACACGACCATCCACGTGCTTGAG | |
| BC1_#42 | /5Phos/GTGCGAACTCAGACCAGCCATGCATCCACGTGCTTGAG | |
| BC1_#43 | /5Phos/GTGCGAACTCAGACCATCATTCCATCCACGTGCTTGAG | |
| BC1_#44 | /5Phos/GTGCGAACTCAGACCCACCTTACATCCACGTGCTTGAG | |
| BC1_#45 | /5Phos/GTGCGAACTCAGACCCCTAATCCATCCACGTGCTTGAG | |
| BC1_#46 | /5Phos/GTGCGAACTCAGACCCCTCTATCATCCACGTGCTTGAG | |
| BC1_#47 | /5Phos/GTGCGAACTCAGACCGAACAGGCATCCACGTGCTTGAG | |
| BC1_#48 | /5Phos/GTGCGAACTCAGACCGATGAATCATCCACGTGCTTGAG |
BC2
| A | B | |
| Name | Sequence | |
| BC2_#01 | /5Phos/CATCGGCGTACGACTAACGTGATGGATTCGAGGAGCGT | |
| BC2_#02 | /5Phos/CATCGGCGTACGACTAAACATCGGGATTCGAGGAGCGT | |
| BC2_#03 | /5Phos/CATCGGCGTACGACTACCACTGTGGATTCGAGGAGCGT | |
| BC2_#04 | /5Phos/CATCGGCGTACGACTCATCAAGTGGATTCGAGGAGCGT | |
| BC2_#05 | /5Phos/CATCGGCGTACGACTCTGTAGCCGGATTCGAGGAGCGT | |
| BC2_#06 | /5Phos/CATCGGCGTACGACTAACCGAGAGGATTCGAGGAGCGT | |
| BC2_#07 | /5Phos/CATCGGCGTACGACTAAGGTACAGGATTCGAGGAGCGT | |
| BC2_#08 | /5Phos/CATCGGCGTACGACTACAGCAGAGGATTCGAGGAGCGT | |
| BC2_#09 | /5Phos/CATCGGCGTACGACTACCTCCAAGGATTCGAGGAGCGT | |
| BC2_#10 | /5Phos/CATCGGCGTACGACTACGCTCGAGGATTCGAGGAGCGT | |
| BC2_#11 | /5Phos/CATCGGCGTACGACTACTATGCAGGATTCGAGGAGCGT | |
| BC2_#12 | /5Phos/CATCGGCGTACGACTAGAGTCAAGGATTCGAGGAGCGT | |
| BC2_#13 | /5Phos/CATCGGCGTACGACTAGATCGCAGGATTCGAGGAGCGT | |
| BC2_#14 | /5Phos/CATCGGCGTACGACTCAATGGAAGGATTCGAGGAGCGT | |
| BC2_#15 | /5Phos/CATCGGCGTACGACTCAGCGTTAGGATTCGAGGAGCGT | |
| BC2_#16 | /5Phos/CATCGGCGTACGACTCATACCAAGGATTCGAGGAGCGT | |
| BC2_#17 | /5Phos/CATCGGCGTACGACTCCAGTTCAGGATTCGAGGAGCGT | |
| BC2_#18 | /5Phos/CATCGGCGTACGACTCGAACTTAGGATTCGAGGAGCGT | |
| BC2_#19 | /5Phos/CATCGGCGTACGACTCGACTGGAGGATTCGAGGAGCGT | |
| BC2_#20 | /5Phos/CATCGGCGTACGACTCTCAATGAGGATTCGAGGAGCGT | |
| BC2_#21 | /5Phos/CATCGGCGTACGACTCTGAGCCAGGATTCGAGGAGCGT | |
| BC2_#22 | /5Phos/CATCGGCGTACGACTGAATCTGAGGATTCGAGGAGCGT | |
| BC2_#23 | /5Phos/CATCGGCGTACGACTGAGCTGAAGGATTCGAGGAGCGT | |
| BC2_#24 | /5Phos/CATCGGCGTACGACTGATAGACAGGATTCGAGGAGCGT | |
| BC2_#25 | /5Phos/CATCGGCGTACGACTGCTAACGAGGATTCGAGGAGCGT | |
| BC2_#26 | /5Phos/CATCGGCGTACGACTGCTCGGTAGGATTCGAGGAGCGT | |
| BC2_#27 | /5Phos/CATCGGCGTACGACTGGAGAACAGGATTCGAGGAGCGT | |
| BC2_#28 | /5Phos/CATCGGCGTACGACTGGTGCGAAGGATTCGAGGAGCGT | |
| BC2_#29 | /5Phos/CATCGGCGTACGACTGTACGCAAGGATTCGAGGAGCGT | |
| BC2_#30 | /5Phos/CATCGGCGTACGACTGTCGTAGAGGATTCGAGGAGCGT | |
| BC2_#31 | /5Phos/CATCGGCGTACGACTGTGTTCTAGGATTCGAGGAGCGT | |
| BC2_#32 | /5Phos/CATCGGCGTACGACTTCTTCACAGGATTCGAGGAGCGT | |
| BC2_#33 | /5Phos/CATCGGCGTACGACTTGGAACAAGGATTCGAGGAGCGT | |
| BC2_#34 | /5Phos/CATCGGCGTACGACTTGGCTTCAGGATTCGAGGAGCGT | |
| BC2_#35 | /5Phos/CATCGGCGTACGACTTGGTGGTAGGATTCGAGGAGCGT | |
| BC2_#36 | /5Phos/CATCGGCGTACGACTTTCACGCAGGATTCGAGGAGCGT | |
| BC2_#37 | /5Phos/CATCGGCGTACGACTAACTCACCGGATTCGAGGAGCGT | |
| BC2_#38 | /5Phos/CATCGGCGTACGACTAAGAGATCGGATTCGAGGAGCGT | |
| BC2_#39 | /5Phos/CATCGGCGTACGACTAAGGACACGGATTCGAGGAGCGT | |
| BC2_#40 | /5Phos/CATCGGCGTACGACTAATCCGTCGGATTCGAGGAGCGT | |
| BC2_#41 | /5Phos/CATCGGCGTACGACTACACGACCGGATTCGAGGAGCGT | |
| BC2_#42 | /5Phos/CATCGGCGTACGACTAGCCATGCGGATTCGAGGAGCGT | |
| BC2_#43 | /5Phos/CATCGGCGTACGACTATCATTCCGGATTCGAGGAGCGT | |
| BC2_#44 | /5Phos/CATCGGCGTACGACTCACCTTACGGATTCGAGGAGCGT | |
| BC2_#45 | /5Phos/CATCGGCGTACGACTCCTAATCCGGATTCGAGGAGCGT | |
| BC2_#46 | /5Phos/CATCGGCGTACGACTCCTCTATCGGATTCGAGGAGCGT | |
| BC2_#47 | /5Phos/CATCGGCGTACGACTGAACAGGCGGATTCGAGGAGCGT | |
| BC2_#48 | /5Phos/CATCGGCGTACGACTGATGAATCGGATTCGAGGAGCGT |
BC3
| A | B | |
| Name | Sequence | |
| BC3_#01 | CAGACGTGTGCTCTTCCGATCTAACGTGATGTGGCCGATGTTTCG | |
| BC3_#02 | CAGACGTGTGCTCTTCCGATCTAAACATCGGTGGCCGATGTTTCG | |
| BC3_#03 | CAGACGTGTGCTCTTCCGATCTACCACTGTGTGGCCGATGTTTCG | |
| BC3_#04 | CAGACGTGTGCTCTTCCGATCTCATCAAGTGTGGCCGATGTTTCG | |
| BC3_#05 | CAGACGTGTGCTCTTCCGATCTCTGTAGCCGTGGCCGATGTTTCG | |
| BC3_#06 | CAGACGTGTGCTCTTCCGATCTAACCGAGAGTGGCCGATGTTTCG | |
| BC3_#07 | CAGACGTGTGCTCTTCCGATCTAAGGTACAGTGGCCGATGTTTCG | |
| BC3_#08 | CAGACGTGTGCTCTTCCGATCTACAGCAGAGTGGCCGATGTTTCG | |
| BC3_#09 | CAGACGTGTGCTCTTCCGATCTACCTCCAAGTGGCCGATGTTTCG | |
| BC3_#10 | CAGACGTGTGCTCTTCCGATCTACGCTCGAGTGGCCGATGTTTCG | |
| BC3_#11 | CAGACGTGTGCTCTTCCGATCTACTATGCAGTGGCCGATGTTTCG | |
| BC3_#12 | CAGACGTGTGCTCTTCCGATCTAGAGTCAAGTGGCCGATGTTTCG | |
| BC3_#13 | CAGACGTGTGCTCTTCCGATCTAGATCGCAGTGGCCGATGTTTCG | |
| BC3_#14 | CAGACGTGTGCTCTTCCGATCTCAATGGAAGTGGCCGATGTTTCG | |
| BC3_#15 | CAGACGTGTGCTCTTCCGATCTCAGCGTTAGTGGCCGATGTTTCG | |
| BC3_#16 | CAGACGTGTGCTCTTCCGATCTCATACCAAGTGGCCGATGTTTCG | |
| BC3_#17 | CAGACGTGTGCTCTTCCGATCTCCAGTTCAGTGGCCGATGTTTCG | |
| BC3_#18 | CAGACGTGTGCTCTTCCGATCTCGAACTTAGTGGCCGATGTTTCG | |
| BC3_#19 | CAGACGTGTGCTCTTCCGATCTCGACTGGAGTGGCCGATGTTTCG | |
| BC3_#20 | CAGACGTGTGCTCTTCCGATCTCTCAATGAGTGGCCGATGTTTCG | |
| BC3_#21 | CAGACGTGTGCTCTTCCGATCTCTGAGCCAGTGGCCGATGTTTCG | |
| BC3_#22 | CAGACGTGTGCTCTTCCGATCTGAATCTGAGTGGCCGATGTTTCG | |
| BC3_#23 | CAGACGTGTGCTCTTCCGATCTGAGCTGAAGTGGCCGATGTTTCG | |
| BC3_#24 | CAGACGTGTGCTCTTCCGATCTGATAGACAGTGGCCGATGTTTCG | |
| BC3_#25 | CAGACGTGTGCTCTTCCGATCTGCTAACGAGTGGCCGATGTTTCG | |
| BC3_#26 | CAGACGTGTGCTCTTCCGATCTGCTCGGTAGTGGCCGATGTTTCG | |
| BC3_#27 | CAGACGTGTGCTCTTCCGATCTGGAGAACAGTGGCCGATGTTTCG | |
| BC3_#28 | CAGACGTGTGCTCTTCCGATCTGGTGCGAAGTGGCCGATGTTTCG | |
| BC3_#29 | CAGACGTGTGCTCTTCCGATCTGTACGCAAGTGGCCGATGTTTCG | |
| BC3_#30 | CAGACGTGTGCTCTTCCGATCTGTCGTAGAGTGGCCGATGTTTCG | |
| BC3_#31 | CAGACGTGTGCTCTTCCGATCTGTGTTCTAGTGGCCGATGTTTCG | |
| BC3_#32 | CAGACGTGTGCTCTTCCGATCTTCTTCACAGTGGCCGATGTTTCG | |
| BC3_#33 | CAGACGTGTGCTCTTCCGATCTTGGAACAAGTGGCCGATGTTTCG | |
| BC3_#34 | CAGACGTGTGCTCTTCCGATCTTGGCTTCAGTGGCCGATGTTTCG | |
| BC3_#35 | CAGACGTGTGCTCTTCCGATCTTGGTGGTAGTGGCCGATGTTTCG | |
| BC3_#36 | CAGACGTGTGCTCTTCCGATCTTTCACGCAGTGGCCGATGTTTCG | |
| BC3_#37 | CAGACGTGTGCTCTTCCGATCTAACTCACCGTGGCCGATGTTTCG | |
| BC3_#38 | CAGACGTGTGCTCTTCCGATCTAAGAGATCGTGGCCGATGTTTCG | |
| BC3_#39 | CAGACGTGTGCTCTTCCGATCTAAGGACACGTGGCCGATGTTTCG | |
| BC3_#40 | CAGACGTGTGCTCTTCCGATCTAATCCGTCGTGGCCGATGTTTCG | |
| BC3_#41 | CAGACGTGTGCTCTTCCGATCTACACGACCGTGGCCGATGTTTCG | |
| BC3_#42 | CAGACGTGTGCTCTTCCGATCTAGCCATGCGTGGCCGATGTTTCG | |
| BC3_#43 | CAGACGTGTGCTCTTCCGATCTATCATTCCGTGGCCGATGTTTCG | |
| BC3_#44 | CAGACGTGTGCTCTTCCGATCTCACCTTACGTGGCCGATGTTTCG | |
| BC3_#45 | CAGACGTGTGCTCTTCCGATCTCCTAATCCGTGGCCGATGTTTCG | |
| BC3_#46 | CAGACGTGTGCTCTTCCGATCTCCTCTATCGTGGCCGATGTTTCG | |
| BC3_#47 | CAGACGTGTGCTCTTCCGATCTGAACAGGCGTGGCCGATGTTTCG | |
| BC3_#48 | CAGACGTGTGCTCTTCCGATCTGATGAATCGTGGCCGATGTTTCG |
The following sub-steps describe annealing linker primers with barcode primers in RNAse-free 96-well plates (Twin-Tec PRC plate, 96 Lobind, Semi-skirted) to a total volume of 10 µL per well. At the end of the resulting plate will contain oligos with the following concentrations:
BC1-Linker_1: in round 1 plates, containing:
- 9 µL Linker_1 and
- 10 µL BC1 barcodes
BC2-Linker_2: in round 2 plates, containing:
- 11 µL Linker_2 and
- 12 µL BC2 barcodes
BC3-Linker_3: in round 3 plates, containing:
- 13 µL Linker_3 and
- 14 µL BC3 barcodes
To prepare stock of 9 plates of 48 barcodes:
For each linker-barcode, prepare 10 µL /well x 9 x 1.1 = 100 µL ;
- BC1-Linker1:
- Linker_1 (1 millimolar (mM) stock) 0.9 µL
- BC1 (100 micromolar (µM) stock) 10 µL
- STE buffer 89.1 µL ( STE )
- BC2-Linker2:
- Linker_2 (1 millimolar (mM) stock) 1.1 µL
- BC2 (100 micromolar (µM) stock) 12 µL
- STE buffer 86.9 µL
- BC3-Linker3:
- Linker_3 (1 millimolar (mM) stock) 1.3 µL
- BC3 (100 micromolar (µM) stock) 14 µL
- STE buffer 84.7 µL
Prepare linker stocks (x 48 x 1.1 = x 53; in each round, linker is the same one)
- Linker_1 47.7 µL + STE buffer 4722.3 µL
- Linker_2 58.3 µL + STE buffer 4605.7 µL
- Linker_3 68.9 µL + STE buffer 4489.1 µL
Aliquot diluted linker into 48 wells: Linker_1 90 µL ; Linker_2 88 µL ; Linker_3 86 µL .
- Add barcodes of BC1 (1-48) 10 µL to Linker_1
- Add barcodes of BC2 (1-48) 12 µL to Linker_2
- Add barcodes of BC3 (1-48) 14 µL to Linker_3
Place sealed plates with annealing linker and barcodes in PCR machine with the programme:
95 °C 00:02:00 , cooling down to 20 °C at -1 °C /min
Aliquot mixture 10 µL to each well into 96-well plates
Seal plates and store at -20 °C .
Solution Preparation
Equipment:
- PCR machine (thermocycler)
Primers
Barcoding for antibody Identification, with example primer sequence and index position:
| A | B | |
| Name | Sequence | |
| MEB_dU_#01 | /5Phos/AGGCCAGAGCATTCGAATGGNNNNNNN/ideoxyU/AGATGTGTATAAGAGACAG | |
| MEB_dU_#02 | /5Phos/AGGCCAGAGCATTCGTGAGANNNNNNN/ideoxyU/AGATGTGTATAAGAGACAG | |
| MEB_dU_#03 | /5Phos/AGGCCAGAGCATTCGCCATANNNNNNN/ideoxyU/AGATGTGTATAAGAGACAG | |
| MEB_dU_#04 | /5Phos/AGGCCAGAGCATTCGGGTGTNNNNNNN/ideoxyU/AGATGTGTATAAGAGACAG | |
| ME_R | /5Phos/C*T*G*T*C*T*C*T*T*A*T*A*C*A*/3ddC/ | |
| ME_A | TCGTCGGCAGCGTCAGATGTGTATAAGAGACAG |
*N indicates a phosphorothioate bond (primer sequences are in the IDT DNA format).
Dilute oligonucleotides to 200 micromolar (µM) in STE buffer ( STE )
pA-Tn5-MeBdU-MeR
For each of the four MEB_dU oligonucleotides, in separate tubes mix:
- 2 µL 200 micromolar (µM) phosphorylated Mosaic end adapter B dU (MEB_dU_#N)
- 2 µL 200 micromolar (µM) blocked Mosaic end-reverse (ME_R) oligonucleotides
to generate 100 micromolar (µM) pre-annealed MEBdU-MER
pA-Tn5-MeA-MeR
Mix:
- 2 µL 200 micromolar (µM) Mosaic end adapter A (ME_A)
- 2 µL 200 micromolar (µM) blocked Mosaic end-reverse (ME-R) oligonucleotides
to generate 100 micromolar (µM) pre-annealed MEA-MER
Place the tubes in a thermocycler and run 95 °C 00:02:00 , slowly cool to 20 °C with ramp rate of -1 °C /min
2m
Mix 4 µL 100 micromolar (µM) pre-annealed MEBdU-MER or MEA-MER with 40 µL of 5.5 micromolar (µM) pA-Tn5 fusion protein and incubate on a rotating platform for 01:00:00 at room temperature. Store at -20 °C for up to 1 year.
Note
We use home-made pA-Tn5 fusion protein generated from plasmid #124601 from Addgene https://www.addgene.org/124601/ following this protocol:
Citation
LINK
Commercially available alternatives are:
- pA-Tn5 Transposase (10 µg)Active MotifCatalog #53161
- CUTANA™ Uncharged pAG-Tn5 for CUT&TagEpiCypherCatalog #15-1025
1h
Single-cell dissociation
Equipment:
- Centrifuge with swing-bucket rotor
- 1.5 mL protein lobind tube
Dissociate samples into single-cell suspension, and transfer cells to 1.5 mL protein lobind tube
Wash cells twice with PBSI ( PBSI ) by centrifuging cells in a swing-bucket rotor at 300 x g, 00:03:00 , and remove the supernatant.
Nuclei Preparation
To prepare nuclei:
Measure the the volume of the the single cell suspension and add PBSI ( PBSI ) to cells to a final volume of 80 µL
Add 80 µL 2x NE buffer ( 2x NE Nuffer ) (with 2x PIC and 2x RI)
briefly vortex to mix and put cells on ice for 00:10:00 (adjust incubation time if needed; Trypan Blue Solution, 0.4%Thermo FisherCatalog #15250061 , trypan blue positive > 95%).
Light Fixation and Quench
To help maintain intact nuclei for split-pool barcoding, we use a light fixation with 0.2% formaldehyde.
Equipment:
- Centrifuge with swing bucket rotor
- (optional) Mr. Frosty container
Fixation wash buffer
| A | B | C | |
| Reagent | Stocking | /µL | |
| Base Wash Buffer | 867 | ||
| BSA | 7.5% | 133 | |
| PIC | 100x | 10 | |
| RI | 80x | 12.5 |
Quickly spin (100 x g, 00:00:05 ) to collect solution into the bottom, add 2 µL 16% Formaldehyde (w/v) Methanol-freeThermo Fisher ScientificCatalog #28906 to final 0.2%, immediately tap to mix and start stopwatch. Incubate for 00:05:00 exactly .
Quickly spin (100 x g, 00:00:05 ) to collect solution into the bottom, add 40 µL 5x Fixation Quench Buffer ( 5x Fixation Quench Buffer ) to quench nuclei, invert to mix, place on ice for 00:05:00 - 00:10:00 .
Centrifuge nuclei in a swing-bucket rotor (pre-cooled) at 600 x g, 4°C, 00:03:00 , carefully take out tubes and remove 180 µL from the top.
Leave ~20 µL in the tube without disturbing nuclei at the bottom.
Add 100 µL Fixation wash buffer to re-suspend cells, invert to mix.
Note
Optional Stopping Point
Quickly spin 100 x g, 00:00:05 , add 15 µL 100% DMSO to each sample (10-15% in final), invert to mix. Quickly spin 100 x g, 00:00:05 , place samples in a Mr. Frosty container and store at -80 °C overnight to freeze samples. The next day, samples are ready for shipment on dry ice or can be stored at -80 °C freezer for a longer term.
Binding nuclei to beads
15m
To immobilise nuclei, we bind the lightly-fixed nuclei with Concanavalin A (ConA)-coated magnetic beads.
Equipment:
- Magnet stand
Beads Preparation
Transfer (5 x N* x 1.1) μL
Concanavalin-coated magnetic beads Bangs LaboratoriesCatalog #BP531 bead slurry into a 1.5 mL tube , wash twice with 1 mL binding buffer, mix by pipetting, and place the tube on a magnetic stand to allow the mixture to clear, and drain liquid completely.
*where N is the number of samples
Note
Using more bead slurry could alleviate the formation of clumps. We typically use around 1 µL per 1,000 cells.
Re-suspend beads in (5 x N x 1.1) μL binding buffer, and hold on ice until nuclei are ready.
Preparing
Thawing Wash Buffer:
| A | B | C | D | |
| Reagents | Stocking | Working | /μL | |
| Base Wash Buffer | 867 | |||
| BSA | 7.5% | 1% | 133 | |
| PIC | 100x | 1x | 10 | |
| RI | 80x | 0.4x | 5 |
- 867 µL wash buffer ( wash buffer )
- 133 µL Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- 5 µL RI mix
- 10 µL PIC
Thawing Wash buffer + EDTA:
| A | B | C | D | |
| Reagents | Stocking | Working | /μL | |
| Thawing Wash Buffer | 500 | |||
| EDTA | 0.5 M | 2 µM | 2 |
- 500 µL Thawing Wash Buffer
- 2 µL EDTA (0.5 M), pH 8.0, RNase-freeThermo FisherCatalog #AM9260G
Thaw nuclei on ice, add 500 µL Thawing Wash Buffer
Slowly add prepared ConA beads from step 12.2 into cell suspension while vortexing gently 1100 rpm Vortex Gently , place tubes on an end-over-end rotator for 00:05:00 - 00:10:00 Room temperature
15m
Place the tubes on a magnetic stand to allow the mixtures to clear. Carefully discard the liquid (taking care due to surface tension).
Wash once with 100 µL Thawing Wash Buffer + EDTA while tubes are on the magnetic stand.
DNA tagmentation
The steps below describe how to simultaneously tagment and profile multiple indexed antibodies using scMTR-seq. We use indexed MEBdU to multiplex each primary antibody of choice with different pA-Tn5-MEBdU-MER. To minimise cross-contamination between antibodies, we then block any unreacted pA-Tn5-MEBdu_MER by incubation with normal IgG. We have successfully profiled five different histone modifications simaltaenously with only minor off-target signals detected. Including an additional IgG profile helps to identify and further reduce off-target signals using computational methods.
As an alternative approach, we have also performed tagmentation in multiple sequential steps, rather than simultaneously (see step 13.12). Sequential profiling increases hands-on time and may lead to greater loss of nuclei, although the advantages of this alternative approach include reduced off-target signals and the opportunity to profile more targets in different rounds.
Solution Preparation
Equipment:
- end-over-end rotator
- cold room
- PCR machine (thermocycler)
Preparing
High RI wash 300 buffer
| A | B | C | |
| Reagents | Stocking | /μL | |
| Base Wash Buffer | 867 | ||
| NaCl | 5M | 30 | |
| BSA | 7.5% | 133 | |
| PIC | 100x | 10 | |
| RI | 80x | 12.5 | |
- 867 µL wash buffer ( wash buffer )
- 30 µL 5 Molarity (M) Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
- 133 µL Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- 12.5 µL RI mix
- 10 µL PIC
(make 100 µL per sample)
Low RI Wash 300 buffer
| A | B | C | |
| Reagents | Stocking | /μL | |
| Base Wash Buffer | 867 | ||
| NaCl | 5 M | 30 | |
| BSA | 7.5% | 133 | |
| PIC | 100x | 10 | |
| RI | 80x | 5 |
- 867 µL wash buffer ( wash buffer )
- 30 µL 5 Molarity (M) Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
- 133 µL Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- 10 µL PIC
- 5 µL RI mix
(make 100 µL per sample)
Tag buffer
| A | B | C | |
| Reagent | Stocking | /μL | |
| Base Wash Buffer | 867 | ||
| NaCl | 5 M | 30 | |
| BSA | 7.5% | 13.3 | |
| PIC | 100x | 10 | |
| RI | 80x | 25 | |
| Nuclease-free water | 119.7 | ||
| MgCl2 | 1M | 10 |
- 867 µL wash buffer ( wash buffer )
- 30 µL 5 Molarity (M) Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
- 13.3 µL Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- 25 µL RI mix
- 10 µL PIC
- 119.7 µL Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
- 10 µL 1 Molarity (M) 1M MgCl2Invitrogen - Thermo FisherCatalog #AM9530G
(make 100 µL per sample)
Pre-assemble selected primary antibody (Ab) and indexed pA-Tn5-MeBdU-MeR (0.5-1h). We list below several antibodies that we have used successfully in scMTR-seq. Alternative antibodies can be tested first using bulk CUT&Tag to check for signal specificity.
Example Antibodies:
- Anti-Histone H3 (tri methyl K4) antibody - ChIP GradeAbcamCatalog #ab8580
- Anti-Histone H3 (tri methyl K9) antibody - ChIP GradeAbcamCatalog #ab8898
- Anti-Histone H3 (acetyl K27) antibody - ChIP GradeAbcamCatalog #ab4729
- Anti-Histone H3 (mono methyl K4) antibody - ChIP GradeAbcamCatalog #ab8895
- Tri-Methyl-Histone H3 (Lys27) (C36B11) Rabbit mAbCell Signaling TechnologyCatalog #9733
- Anti-Histone H3 (tri methyl K36) antibody - ChIP GradeAbcamCatalog #ab9050
Mix:
- 0.5 µL 1 µg/µL antibody
- 0.5 µL pA-Tn5-MeBdU-MeR
- 4.25 µL High RI wash 300 buffer
place tubes on an end-over-end rotator
10 rpm, Room temperature, 01:00:00 , end-over-end rotator
1h
Block any remaining unreacted proteinA-Tn5:
Add 1 µL 1 µg/µL Normal Rabbit IgG Control R&D SystemsCatalog #AB-105-C and incubate Room temperature for 00:30:00
30m
Mix pre-assembled Ab-pA-Tn5-MeBdU-MeR (6 µL x n of indexed antibodies) with ConA-bound nuclei, add High RI wash 300 to 99.6 µL , and then add 0.4 µL 0.5 Molarity (M) EDTA.
Gently mix, briefly spin and incubate in cold room (4 °C ) Overnight at 25-30 rpm on rotator or 1-2 hrs at Room temperature
2h
Wash three times with Low RI Wash 300 buffer, using the magnetic stand, and invert to mix between washes
Re-suspend with 50 µL Tag buffer, gently mix and place in PCR machine, incubate 37 °C for 01:00:00
1h
Directly place tube on pre-cooled rack, and add 1.67 µL 0.5 Molarity (M) EDTA each sample to stop tagmentation, gently tap to mix
Add 8 µL Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037 to each sample to avoid clumps forming. Mix by pipetting or gently tapping, place on ice for 00:05:00
5m
(Optional)
For sequential tagmentation, wash twice with Low RI wash 300 buffer after adding stop buffer, and repeat from to step 13.11.
In situ Reverse Transcription
17m 12s
In situ reverse transcribe polyA mRNA with biotinylated dT RT primers in the nuclei.
Equipment:
- PCR machine
Primers
| A | B | |
| Name | Sequence | |
| RT_#01 | /5Phos/AGGCCAGAGCATTCGAGCAANNNNNNNN /iBiodT/TTTTTTTTTTTTTTVN | |
| RT_#02 | /5Phos/AGGCCAGAGCATTCGCCTAANNNNNNNN /iBiodT/TTTTTTTTTTTTTTVN | |
| RT_#03 | /5Phos/AGGCCAGAGCATTCGGTCGANNNNNNNN /iBiodT/TTTTTTTTTTTTTTVN | |
| RT_#04 | /5Phos/AGGCCAGAGCATTCGTGTGANNNNNNNN /iBiodT/TTTTTTTTTTTTTTVN |
Preparing
NIB-RI buffer
- 867 µL NIB buffer ( Nuclei Isolation Buffer )
- 133 µL Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- 5 µL RI mix
- 10 µL PIC
(make 350 µL per sample)
RT mix:
| A | B | C | D | |
| Reagent | Stocking | Working 1x | 1x volume /μL | |
| RT buffer | 5x | 1x | 10 | |
| PEG 6000 | 50 % | 15 % | 15 | |
| RT_#N primer | 100 μM | 10 μM | 5 | |
| dNTPs | 25 mM/each | 0.5 mM | 1 | |
| RNAse Protector | 40 U/μL | 0.5 U/μL | 0.625 | |
| Superase RI | 20 U/μL | 0.5 U/μL | 1.25 | |
| Maxima H Minus RT | 200 U/μL | 20 U/μl | 5 | |
| Nuclease-free water | 12.125 | |||
| total | 50 |
- PEG 6000 (Poly(ethylene glycol))Bio Basic Inc.Catalog #PB0432.SIZE.500g
- RT_#N
- dNTP Set 100 mM SolutionsThermo Fisher ScientificCatalog #R0182
- Protector RNase InhibitorMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335399001
- SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696
- Maxima H Minus Reverse TranscriptaseThermo Fisher ScientificCatalog ##EP0741
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Wash in situ-tagged nuclei twice with 100 µL NIB-RI buffer.
Add 10 µL of RT mix per 10,000 nuclei to each tube, gently mix and briefly centrifuge
- Run following programme to synthesise the first strand of cDNA: 50 °C 00:10:00
- 3 cycles of annealing:
8 °C 00:00:12
15 °C 00:00:45
20 °C 00:00:45
30 °C 00:00:30
42 °C 00:02:00
50 °C 00:03:00
End cycles
- 50 °C 5 min
- 4 °C Hold
17m 12s
Add 50 µL NIB-RI to dilute sample, place tubes on PCR magnet and remove liquid
Wash once with 100 µL NIB-RI and remove liquid
Split-pool barcoding
3h 14m
Three rounds of split-pool barcoding were used for barcoding single cells. As there is 48 different barcodes for each round in our assay, this will generate over 110,000 barcodes combinations (# of BC1 x # of BC2 x # of BC3) after three rounds of the split-pool barcoding procedure. Both cDNA and tagged gDNA molecules will get the same combination of barcodes within the same single nuclei.
Note
To ensure there are enough barcode combinations for barcoding each single cell/nuclei with a unique cell barcode, we adjust the conditions to keep the collision rate below 5%. The collision rate is calculated by dividing the number of cells or nuclei by the number of barcode combinations.
The number of cells or nuclei: as the recovery rate from the starting point is normally ~20-40%, we use the number of starting cells multiplied by the recovery rate to estimate the number of cells/nuclei.
The number of barcode combinations: in addition to the split-pool steps, we could use different RT primers and MEB adapters for different samples, and different nuclei could be aliquoted into different sub-libraries and amplified with different i5-i7 primer sets, so all these steps will contribute to the total number of barcodes combinations. Therefore, the number of barcode combinations = the number of RT primers or MEB adapters x number of BC1 x number of BC2 x number of BC3 x number of sub-libraries.
In practice, we normally adjust the number of RT primers and MEB adapters used and the number of sub-libraries to keep the collision rate below 5%.
Example calculations are included in the attached spreadsheet.
Equipment:
- Plate centrifuge
- Multichannel pipettor
- Thermomixer
Primers
| A | B | |
| Name | Sequence | |
| Blocker_1 | ATCCACGTGCTTGAGAGGCCAGAGCATTCG | |
| Blocker_2 | GGATTCGAGGAGCGTGTGCGAACTCAGACC | |
| Blocker_3 | GTGGCCGATGTTTCGCATCGGCGTACGACT |
Preparing
Hybridisation mix: (2 mL per sample)
| A | B | C | D | |
| Reagents | Stocking | Working 1x | /μl | |
| T4 ligation buffer | 10x | 1x | 200 | |
| RNase Inhibitor | 40 U/μL | 0.32 U/μl | 16 | |
| SUPERase RI | 20 U/μL | 0.05 U/μl | 5 | |
| PIC | 100x | 1x | 20 | |
| Triton X-100 | 10% (vol/vol) | 0.1% | 20 | |
| BSA | 7.5% | 0.1% | 27 | |
| NIB | 0.25x | 500 | ||
| Nuclease-free water | 1212 | |||
| total | 2000 |
- T4 DNA Ligase - 100,000 unitsNew England BiolabsCatalog #M0202L
- Protector RNase InhibitorMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335402001
- SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696
- PIC
- 10% Triton X-100Merck MilliporeSigma (Sigma-Aldrich)
- Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- NIB
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
block1 mix:
| A | B | C | D | |
| Blocker_1 | 1 mM | 22 μM | 12.1 μL | |
| T4 Ligation buffer | 10x | 2x | 110 μL | |
| Nuclease-free water | 427.9 μL | |||
| 550 μL |
- T4 DNA Ligase - 100,000 unitsNew England BiolabsCatalog #M0202L
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
block2 mix:
| A | B | C | D | |
| Blocker_2 | 1mM | 26.4μM | 14.52 μL | |
| T4 Ligation buffer | 10x | 2x | 110 μL | |
| Nuclease-free water | 425.48 μL | |||
| 550 μL |
- T4 DNA Ligase - 100,000 unitsNew England BiolabsCatalog #M0202L
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
block3 mix:
| A | B | C | D | |
| Blocker_3 | 1mM | 23 μM | 12.65μL | |
| Triton X100 | 10% | 0.1% | 5.5μL | |
| Nuclease-free water | 531.85μL | |||
| 550μL |
- 10% Triton X-100Merck MilliporeSigma (Sigma-Aldrich)
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Ligation mix: (0.5 mL per sample)
| A | B | C | D | |
| Reagents | Stocking | Working 1x | /μL | |
| T4 ligation buffer | 10x | 1x | 50 | |
| RNase Inhibitor | 40 U/μL | 0.32 U/μl | 4 | |
| Superase RI | 20 U/μL | 0.05 U/μL | 1.25 | |
| PIC | 100x | 1x | 5 | |
| T4 DNA ligase | 2000 U/μL | 20 U/μl | 5 | |
| Triton X-100 | 10% (vol/vol) | 0.1% | 5 | |
| BSA | 7.5% | 0.1% | 6.7 | |
| NIB | 0.2x | 100 | ||
| Nuclease-free water | 323.05 | |||
| total | 500 |
- T4 ligation buffer
- Protector RNase InhibitorMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335402001
- SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696
- PIC
- T4 DNA Ligase - 100,000 unitsNew England BiolabsCatalog #M0202L
- 10% Triton X-100Merck MilliporeSigma (Sigma-Aldrich)
- Bovine Albumin Fraction V (7.5% solution)Thermo FisherCatalog #15260037
- NIB
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Split-pool barcoding
Prepare 1% BSA in DPBS for rinsing plasticware, which helps to reduce sample loss.
Take out pre-prepared annealed linker-barcodes 96 well plates , spin at 700 x g, Room temperature, 00:02:00 , thaw and bring to Room temperature
2m
Re-suspend ConA-bead bound nuclei with 2 mL hybridisation mix for 48 barcodes in each round
Round 1
- Add 40 µL of nuclei per well to linker1-BC1 plate , 300 rpm, Room temperature, 00:30:00 , incubate on thermomixer (or 23 °C )
- Use multichannel pipettor to add 10 µL block1, pipetting up and down to mix,300 rpm, Room temperature, 00:30:00 , incubate on thermomixer (or 23 °C )
- Centrifuge 300 x g, 00:02:00
- Use multichannel pipette (rinsed with 1% BSA in DPBS) to collect cells into reservoir (rinsed with 1% BSA in DPBS)
1h 2m
Round 2
- Add 55 µL of nuclei per well to linker2-BC2 plate, 300 rpm, Room temperature, 00:30:00 , incubate on thermomixer
- Use multichannel pipette to add 10 µL block2, pipetting up and down to mix,300 rpm, Room temperature, 00:30:00 , incubate on thermomixer (or 23 °C )
- Centrifuge 300 x g, 00:02:00
- Use multichannel pipette (rinsed tips with 1% BSA in DPBS) to collect nuclei into reservoir (also pre-rinsed with 1% BSA in DPBS)
1h 2m
Round 3
- Add 70 µL of nuclei per well to linker3-BC3 plate, 300 rpm, Room temperature, 00:30:00 , incubate on thermomixer (or 23 °C )
- Use multichannel pipette to add 10 µL block3, pipetting up and down to mix
- Centrifuge 300 x g, 00:02:00
- Use multichannel pipette (rinsed tips with 1% BSA in DPBS) to collect nuclei into reservoir (also pre-rinsed with 1% BSA in DPBS)
32m
Transfer nuclei suspension to 5 mL tube, centrifuge 600 x g, 4°C, 00:03:00 , remove and discard the supernatant
3m
Transfer the pellet into a new 1.5 mL protein lobind tube , wash twice with 0.5 mL NIB (no RI, PIC), invert to mix in between
Ligation:
Re-suspend nuclei with 0.5 mL Ligation buffer, 300 rpm, 25°C, 00:30:00 , incubate on thermomixer
30m
Briefly spin, and wash with 0.5 mL NIB (no RI, PIC),
re-suspend in0.5 mL NIB (no RI, PIC),
filter through 30 µL strainer,
rinse strainer with 0.5 mL NIB
(Optional) Put strainer on top of a new tube, centrifuge 600 x g, 4°C, 00:03:00
3m
Collect all nuclei into one tube, manually count nuclei by labelling nuclei with trypan blue (Trypan Blue Solution, 0.4%Thermo FisherCatalog #15250061 )
Based on estimated collision rate, aliquot desired number of nuclei into each sub-library.
Reverse crosslinking and lysis
1h
Reversing the light fixation performed in
Equipment:
- PCR machine
Preparing
2x reverse cross-linking buffer:
| A | B | C | D | |
| Reagents | Stocking | In buffer | /μL | |
| Tris pH 8.0 | 1M | 100 mM | 100 | |
| NaCl | 5M | 100 mM | 20 | |
| SDS | 10% | 0.04% | 4 | |
| Nuclease-free water | 876 | |||
| total | 1000 |
- 1M Tris-HCl (pH 8.0)Thermo Fisher ScientificCatalog #15568025
- Sodium Chloride (5M)Invitrogen - Thermo FisherCatalog #AM9760G
- SDS, 10% Solution, RNase-freeThermo FisherCatalog #AM9822
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Reverse crosslinking and lysis
Add NIB ( Nuclei Isolation Buffer ) to each sample to bring the volume to 20 µL in total
20 µL of 2× reverse cross-linking buffer, 2 µL of 20 mg/mL proteinase K (Proteinase KThermo Fisher ScientificCatalog #EO0491 1 µg/µL in final), and 1.6 µL of RI mix (SUPERase RI:Protector 1:1) were mixed with each sample and incubated at 55 °C for 01:00:00 in PCR machine
1h
OPTIONAL STOP POINT
After adding lysis buffer, sub-libraries can be stored at -20 °C .
Add 2.5 µL of 100 millimolar (mM) PMSFMerck MilliporeSigma (Sigma-Aldrich)Catalog #P7626 to the reverse crosslinked sample to inactivate proteinase K and incubate at Room temperature for 00:10:00 .
10m
Attach to magnet and transfer supernatant into a new tube to remove ConA beads.
Myone C1 Streptavidin beads to separate tagged cDNA and gDNA
1h
Equipment:
- end-to-end-rotator
Wash Dynabeads MyOne Streptavidin C1Invitrogen - Thermo FisherCatalog #65001 (10 µL for each sample, total 10 x N* μL) with 800 µL 1x B&W-T buffer three times, and re-suspend in 45 µL 2x Binding & Washing buffer + 1 µL SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696 .
*where N is the number of samples
Add 46 µL re-suspended beads into each lysed sample (45.6 µL ), mixed and rotated on an end-to-end rotator at 10 rpm, Room temperature, 01:00:00 , end-to-end rotator
1h
Put tubes on a magnetic stand to separate supernatant (transposed DNA fragments) and beads (cDNA).
RNA library preparation
3h 12m
Equipment:
- Qubit fluorometer
- PCR machine (thermocycler)
- end-to-end-rotator
Primers
| A | B | |
| Name | Sequence | |
| TSO | AAGCAGTGGTATCAACGCAGAGTGAATrGrG+G | |
| PA_F | CAGACGTGTGCTCTTCCGATCT | |
| PA_R | AAGCAGTGGTATCAACGCAGAGT | |
| P7_#1 | CAAGCAGAAGACGGCATACGAGATGATCTGGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7_#2 | CAAGCAGAAGACGGCATACGAGATTCAAGTGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7_#3 | CAAGCAGAAGACGGCATACGAGATCTGATCGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7_#4 | CAAGCAGAAGACGGCATACGAGATAAGCTAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7_#5 | CAAGCAGAAGACGGCATACGAGATGTAGCCGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7_#6 | CAAGCAGAAGACGGCATACGAGATTACAAGGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7_#7 | CAAGCAGAAGACGGCATACGAGATTTGACTGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7_#8 | CAAGCAGAAGACGGCATACGAGATGGAACTGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P5_#1 | AATGATACGGCGACCACCGAGATCTACACTAGATCGCTCGTCGGCAGCGTCAGATGTGTAT | |
| P5_#2 | AATGATACGGCGACCACCGAGATCTACACCTCTCTATTCGTCGGCAGCGTCAGATGTGTAT | |
| P5_#3 | AATGATACGGCGACCACCGAGATCTACACTATCCTCTTCGTCGGCAGCGTCAGATGTGTAT | |
| P5_#4 | AATGATACGGCGACCACCGAGATCTACACAGAGTAGATCGTCGGCAGCGTCAGATGTGTAT | |
| P5_#5 | AATGATACGGCGACCACCGAGATCTACACGTAAGGAGTCGTCGGCAGCGTCAGATGTGTAT | |
| P5_#6 | AATGATACGGCGACCACCGAGATCTACACACTGCATATCGTCGGCAGCGTCAGATGTGTAT | |
| P5_#7 | AATGATACGGCGACCACCGAGATCTACACAAGGAGTATCGTCGGCAGCGTCAGATGTGTAT | |
| P5_#8 | AATGATACGGCGACCACCGAGATCTACACCTAAGCCTTCGTCGGCAGCGTCAGATGTGTAT |
Preparing
1x B&W-T-RI buffer:
- 1 mL B&W-T buffer ( B&W-T buffer )
- 5 µL SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696 .
(300 µL per sub-library)
STE-RI buffer:
- 1 mL STE buffer ( STE buffer )
- 5 µL SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696 .
(100 µL per sub-library)
Template switch mix:
| A | B | C | D | |
| Reagent | Stocking | Working 1x | 1x /μL | |
| RT buffer | 5x | 1x | 10 | |
| Ficoll PM-400 | 20 % | 4 % | 10 | |
| PEG 6000 | 50 % | 15 % | 15 | |
| TSO primer | 100 μM | 2.5 μM | 1.25 | |
| dNTPs | 25 mM/each | 1 mM | 2 | |
| RNAse Protector | 40 U/μL | 0.5 U/μL | 0.625 | |
| Superase RI | 20 U/μL | 0.25 U/μL | 0.625 | |
| Maxima H Minus RT | 200 U/μL | 10 U/μl | 2.5 | |
| Nuclease-free water | 8 | |||
| total | 50 |
- RT buffer (Maxima H minus RT)
- Ficoll PM‐400Merck MilliporeSigma (Sigma-Aldrich)Catalog #F5415-50ML
- PEG 6000 (Poly(ethylene glycol))Bio Basic Inc.Catalog #PB0432.SIZE.500g
- TSO (Template Switching Optimised) primer
- dNTP Set 100 mM SolutionsThermo Fisher ScientificCatalog #R0182
- Protector RNase InhibitorMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335402001
- SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696
- Maxima H Minus Reverse TranscriptaseThermo Fisher ScientificCatalog ##EP0741
This is an optimised Moloney murine leukemia virus Reverse Transcriptase (M-MuLV RT) which has terminal transferase activity. This means it can perform non-template addition of nucleotides to 3'-ends with a preference for dCTP. The resulting poly C overhang allows the TSO primer, which ends in 3 rGs, to anneal. This RT is also capable of template switching, meaning that it can read through the gap in the backbone between the end of the RNA template and the annealed TSO and continue elongating with the TSO as the template.
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
RNA library preparation
Wash beads three times using 100 µL 1x B&W-T buffer with SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696 0.1 µL mix by inverting between washes
Wash beads once using 100 µL STE buffer with SUPERaseIN RNase InhibitorThermo Fisher ScientificCatalog #AM2696 0.1 µL while tube on magnet, without re-suspending beads
Re-suspend beads with 50 µL Template switch mix by pipetting
Rotate beads 10 rpm, Room temperature, 00:30:00 , end-to-end-rotator
Re-suspend by pipetting
Incubate for 01:30:00 at 42 °C in PCR machine
2h
Pre-Amplification
After template switching, add 100 µL of STE to each tube to dilute the sample.
Remove the supernatant by placing the sample on a magnetic stand.
Wash beads with 200 µL of STE without disturbing the bead pellet
Beads were then re-suspended in 55 µL of PCR mix:
| A | B | C | D | |
| Reagent | Stocking | Working 1x | 1x /μL | |
| Kapa HiFi PCR mix | 2x | 1x | 27.5 | |
| PA_F | 10 μM | 400 nM | 2.2 | |
| PA_R | 10 μM | 400 nM | 2.2 | |
| Nuclease-free water | 23.1 | |||
| total | 55 |
- HotStart ReadyMix (KAPA HiFi PCR kit)Kapa BiosystemsCatalog #KK2601
- PA_F & PA_R primers
- Nuclease-free water, not DEPC-treated Life TechnologiesCatalog #AM9932
Run following programme to pre-amplify cDNA:
- 95 °C 00:03:00
- 12 cycles of:
- 98 °C 00:00:30
- 65 °C 00:00:45
- 72 °C 00:03:00
- 72 °C 00:05:00
- 4 °C Hold
12m 15s
Purify by 0.8x Ampure XP beads Beckman CoulterCatalog #A63881 and elute in 32 µL 0.1x EB (Elution Buffer)
Measure concentration with Qubit
Note
Libraries made from ~1000-5000 cells and amplified for 12-15 cycles typically yield around 500 ng after purification
Tagging MEA and PCR amplification
Setup following reaction; incubate in PCR machine at 37 °C for 00:30:00
| A | B | |
| 1:20 pA-Tn5-MeA-MeR | 2 μl | |
| 4x Tag Buffer | 5 μl | |
| Sample | Volume to give 50ng | |
| Nuclease-free water | To 20 μl |
30m
Add 2 µL 0.2% SDS mix and incubate at Room temperature for 00:10:00 to release tagged fragments; neutralise SDS with 1 µL 4% Triton X100
- SDS, 10% Solution, RNase-freeThermo FisherCatalog #AM9822
- 10% Triton X-100Merck MilliporeSigma (Sigma-Aldrich)
10m
Amplify libraries with NEBnext mix as following
| A | B | |
| Sample | 23 μl | |
| 2x NEBnext mix | 25 μl | |
| P5_#N 25μM | 1 μl | |
| P7_#N 25uM | 1 μl |
Run the following programme to amplify libraries:
- 72 °C 00:10:00
- 98 °C 00:03:00
- 11 cycles of:
- 98 °C 00:00:10
- 65 °C 00:00:30
- 72 °C 00:01:00
- 72 °C 00:05:00
- 4 °C hold
19m 45s
Purify the amplified library using 0.7x Ampure XP beads Beckman CoulterCatalog #A63881 three times and elute with 32 µL of EB.
DNA library amplification
15m 40s
Equipment:
- PCR machine (thermocycler)
- Qubit fluorometer
- bioanalyzer
Primers
In addition to the P7_#N and P5_#N primers listed above in the RNA library prep section you will need:
| A | B | |
| Name | Sequence | |
| MEA_LNA | TCGTCGGCAGCGTC AGATGTGTA+TA+AG+AG+AC+AG/3InvdT/ |
Purify the DNA component using 1.2x Ampure XP beads Beckman CoulterCatalog #A63881 , elute with 21 µL 0.1x EB buffer
Add MEA by adapter switching
Modified from Mulqueen et al. 2021 "[symmetrical strand single-cell combinatorial indexing (s3)]
uses single-adapter transposition to incorporate the forward primer sequence, the Tn5 mosaic end sequence and a reaction-specific DNA barcode. As with standard tagmentation workflows, extension through the bottom strand is then performed to provide adaptor sequences on both ends of each molecule; however, the s3 transposome complexes contain a uracil base immediately following the mosaic end sequence. Use of a uracil-intolerant polymerase therefore prevents extension beyond the mosaic end into the DNA barcode and forward adaptor sequence. A second template oligo is then introduced that contains a 3’-blocked locked nucleic acid (LNA) mosaic end reverse complement sequence with a reverse adaptor sequence 5’ overhang. This oligo favorably anneals to the copied mosaic end sequence, due to the higher melting temperature of LNA, and acts as a template for the library molecule to extend through and copy the reverse adaptor. This results in all library fragments having both a forward and reverse adaptor sequence. The LNA-templated extension is carried out over multiple rounds of thermocycling to ensure maximum efficiency of reverse adaptor incorporation."
Citation
LINK
Gap filling reaction
20 µL sample (use all of the sample) + 8 µL NEBNext® High-Fidelity 2X PCR Master MixNew England BiolabsCatalog #M0541
run: 72 °C 00:10:00
10m
Adapter switching reaction
Add 3 µL 1 micromolar (µM) MEA_LNA
- run: 98 °C 00:00:30
- 10 cycles of :
- 98 °C 00:00:10
- 59 °C 00:00:20
- 72 °C 00:00:10
1m 10s
Amplification reaction
Add:
- 50 µL Q5U 2x readymix DNA polymerase
- 15 µL H2O
- 2 µL P7_#N (25 micromolar (µM) )
- 2 µL P5_#N (25 micromolar (µM) )
run: 98 °C 00:00:30
- 10 cycles of:
- 98 °C 00:00:10
- 55 °C 00:00:20
- 72 °C 00:00:30
- 72 °C 00:03:00
- 4 °C hold
4m 30s
Take out 50 µL into a new tube and store at -20 °C as a backup, and add extra cycles (4-10 cycles, depending on the sample) to the remaining libraries to generate enough material for sequencing.
Purify libraries with 0.7x Ampure XP beads Beckman CoulterCatalog #A63881 twice, elute with 32 µL EB, check concentration with Qubit, and run samples on Bioanalyzer.
Sequencing
Use a 150 bp paired-end sequencing kit with the following customised parameters:
- Read 2 contains 128 bp for cell barcodes and UMI sequences, plus 72 bp for genomic sequence.
- i7 index is 6bp
- i5 index is 8bp
Normally, we add 10% PhiX spike-in for Illumina sequencing machines, and 2% PhiX spike-in for Element AVITI machine.
Data Processing & Analysis
Data processing can be performed with our analysis pipeline.
See the README file in the pipeline repository for additional details.
Software
nf_mtr_seq
NAME
SOURCE LINK
Protocol references
3XFlag-pATn5: https://dx.doi.org/10.17504/protocols.io.8yrhxv6
CUT&Tag: https://doi.org/10.1038/s41467-019-09982-5
Split-seq: https://www.science.org/doi/10.1126/science.aam8999
Split-seq protocol: https://www.seeliglab.org/uploads/5/9/9/7/59974251/split-seq_protocol_v3.0
Paired-seq: https://doi.org/10.1038/s41594-019-0323-x
Paired-Tag: https://doi.org/10.1038/s41592-021-01060-3
Adapter switch S3: https://doi.org/10.1038/s41587-021-00962-z
Citations
Step 26
Mulqueen RM, Pokholok D, O'Connell BL, Thornton CA, Zhang F, O'Roak BJ, Link J, Yardımcı GG, Sears RC, Steemers FJ, Adey AC. High-content single-cell combinatorial indexing.
https://doi.org/10.1038/s41587-021-00962-z