Feb 22, 2026
SARS-CoV-2 incursion scenario in the city Fantastica v3
This protocol is a draft, published without a DOI.
- Benjamin Schwessinger1
- 1Australian National University

Protocol Citation: Benjamin Schwessinger 2026. SARS-CoV-2 incursion scenario in the city Fantastica v3. protocols.io https://protocols.io/view/sars-cov-2-incursion-scenario-in-the-city-fantasti-hsbnb6amf
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: February 22, 2026
Last Modified: February 22, 2026
Protocol Integer ID: 243790
Keywords: genomic surveillance of sar, following about sar, spread into of sar, spread of sar, genomic surveillance, likely infection scenario for the family infection cluster, family infection cluster, classic epidemiological data with genomic data, sar, available sar, australia on sar, official sar, known outbreak location, exclusive genomic surveillance, genome per infection cycle, pathogen, biosecurity sector, outbreak reference id area, cov2, outbreak reference id area of residence age date, weakness of exclusive genomic surveillance, genomic, combining classic epidemiological data, human biosecurity, used sar, outbreak reference id, genomic data, potential exposure vaccination status oversea, tutorials around human biosecurity, likely infection scenario, quarantine, fantasticasarscov2sequence, matching genomic dataset, genome, viral mutation rate, studying incursion scenario, cov, hospitalcluster1, likely source of infection, vaccination coverage, incursion scenario, infection cycle, city fantastica v2, quarantin
Abstract
This protocol is part of the mini-project on genomic enhance epidemiology of the MAE program delivered by NCEPH at ANU.
It focuses on the 'dry-lab' by investigating a hypothetical incursion scenario in the so-called city Fantastica. You will combine genomic surveillance of SARS-CoV-2 with case interview data to trace the spread into of SARS-CoV-2 in the community and into high risk settings. We will provide you with an (annotated) tree and fantasized case interviews. You will put these two together to trace the spread and suggest potential improvements in containment strategies with a focus on high risk settings.
This is a creative version of similar scenarios investigated during the official SARS-CoV-2 incursion. The main objective of mini-project is to solidify concepts you just learned. We will combine fictional case interview information with a matching genomic dataset of SARS-CoV-2 genomes to investigate the incursion. Hopefully this will show you the power combining these two data types brings when compared to having only one or the other. In the larger perspective of the course, this hopefully illustrates to you that one needs to consider a multitude of perspectives and data types when operating in the biosecurity sector.
I had a lot of fun coming up with this incursion scenario and I hope you will enjoy working on it with your detective hat on. Of course this complete scenario is absolutely fictional. All the used SARS-CoV-2 sequences are publicly available on GISAID as described in this publication (Hall et. al. 2023).
The incursion scenario:
Imagine a city called Fantastica in the middle of the SARS-CoV-2 pandemic mid-2021 in a country where vaccination coverage and COVID-19 case numbers are very low. Fantastica is located on a continental scale island nation and the international borders to this nation are highly regulated to prevent new COVID-19 cases from entering. The main public health measures employed to contain the spread of SARS-CoV-2 are social distancing, mask wearing, mass testing, contact tracing, isolating and quarantining of confirmed cases and lock-downs.
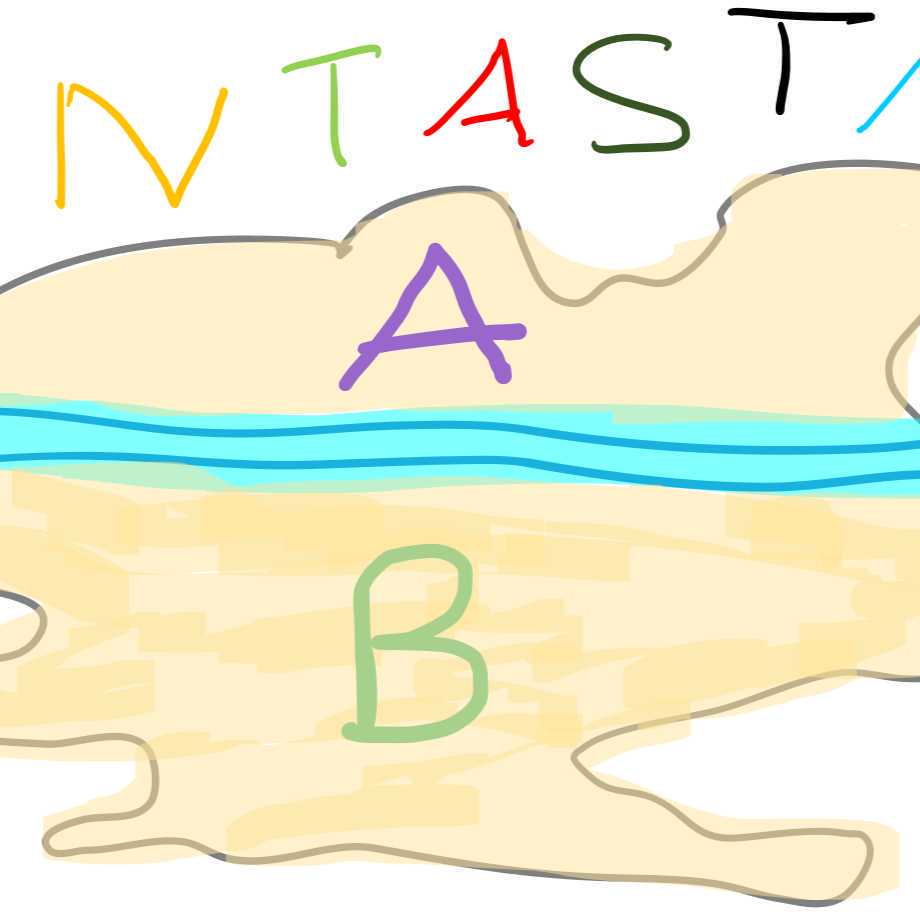
Fantastica has two main areas of residence with A being the affluent North and B being the less well off South (Figure 1). These two areas are separated by a river. The main hospital is located right at the river.

City map Fantastica
In mid-2021 the city experiences its first COVID-19 case for a long time (Outbreak reference ID: Fantastica034), which was successfully contained in hotel quarantine for overseas travelers. The following months Fantastica experiences a larger COVID-19 outbreak that it aims to contain with lockdowns including restricting movements from 12 September 2021 till 20 November 2021. The public health unit achieves to sequence all SARS-CoV-2 genomes of all identified COVID-19 cases in this time frame.
In our simplified scenario we assume the following about SARS-CoV-2:
- Infectious period: 48 hrs before and after onset of symptoms.
- Asymptomatic cases can also cause forward transmission.
- Viral mutation rate: on average 0.5 mutations in each genome per infection cycle.
You are now provided with the following material to start your investigation and address the specific questions below. All the information is idealized and fictionalized.
Provided main material can be found on Canvas:
1. An excel file (1_ContactTracingCaseInterviews) containing case interview information (not exhaustive and simplified) including the following columns:
- Outbreak Reference ID
- Area of Residence
- Age
- Date of symptom onset
- Date of specimen collection
- Symptoms
- Household contact
- Contact with known COVID-19 case
- Case associated with known outbreak
- Locations of potential exposure
- Vaccination Status
- Overseas travel
2. and 3. are rooted phylogenetic trees of all sequenced cases' SARS-CoV-2 genomes (2_MarkedUpTree and 3_NonMarkedUpTree)
4. to 7. are various subtrees that might be useful to address some of the questions.
8. and 9. are the fasta files of SARS-CoV-2 genomes of all identified COVID-19 cases in Fantastica in the indicated study period (plus Fantastica034) (8_FantasticaSARSCoV2Sequences) and the original SARS-CoV-2 genome from Wuhan (9_MN908947.3)
What you need for the mini-project:
- A detective hat.
- Your computer.
- Pen and paper including different colored pens.
Specific questions to be addressed in the mini-project:
- Describe the overall LargeClusterA1. What drove the transmission in this cluster? Was it contained successfully with public health measures such as testing, tracing, lockdowns and quarantine? Has the index case been clearly identified? Is the index case the likely first case in this cluster? Do you think most cases in this cluster have been identified? Explain your reasoning.
- Describe the overall LargeClusterB1. What drove the transmission in this cluster? Was it contained successfully with public health measures such as testing, tracing, lockdowns and quarantine? Has the index case been clearly identified? Is the index case the likely first case in this cluster? Do you think most cases in this cluster have been identified? Include later appearing mini-clusters in your analysis:
MC1: Fantastica063, Fantastica062, Fantastica058, Fantastica064 and Fantastica059.
MC2: Fantastica067, Fantastica068, Fantastica069, Fantastica072, Fantastica074, Fantastica070, Fantastica071, Fantastica073, Fantastica075
In your analysis speculate how, these genetically linked subclusters could potentially physically linked (or not) to the main cluster?
- What is a likely infection scenario for the family infection cluster containing Fantastica014, 016, 017?
- How can you explain that case Fantastica019 is so distinct from all other cases?
- Describe the case Fantastica033. What cluster does this case belong to? When could this case have caught COVID-19? Who could be the potential source cases? Where could this case got infected? Explain your reasoning.
- Describe the "HospitalCluster1 (non-COVID Ward)"? Was it a single incursion? What was the likely transmission chain? How could such an incursion scenario be better managed? Explain your reasoning.
- Describe the "ElderlyHomeClusterB"? Was it a single incursion? What was the likely transmission chain? How could such an incursion scenario be better managed? Explain your reasoning.
- How would you have interpreted case Fantastica076 without contact tracing data? What does this case reveal for the strength and weakness of exclusive genomic surveillance without epi data?
- There is one case that lied on the contact tracing form. Identify this case, its most likely source of infection, and who they passed it on to.
For all these questions we are looking for the most parsimonious answers. The simplest and most plausible answers.
Troubleshooting
Safety warnings
This protocol does not require any hazardous substances or infectious agents. However, maintain a proper posture while working on your computer.
Before start
You must study the protocol carefully before you start. If anything is unclear post questions directly here on protocols.io.
Overlay case interview information on top of the genetic data
On each table you have a print out of the tree that looks like the following

So you have a skeleton (aka tree) of the genetic relationship of all samples and hence COVID-19 cases.
Now you have to overlay the case interview data to answer the specific questions in the description section of the protocol above. For this you can use the printed trees to draw on (with different coloured pens) or annotate it on your computer. Make sure to make good use of the sort and filter functions in Excel when going over the case interview data to ease your analysis.

^^Screenshot of case interview data.
We will walk through some of these questions from the description section in class and will be guided by your questions.
Now annotate the tree with the case information something like the following maybe a bit prettier

Now try to answer some of the questions which are part of the instructions.
We will walk through some of these questions from the description section in class and will be guided by your questions.
