Jan 06, 2025
RLB-MALBAC: A non-transposase method for amplifying and barcoding DNA using Multiple Annealing and Looping Based Amplification Chemistry (MALBAC) for Oxford Nanopore sequencing.
- Vijay J Gadkar1,2,
- David M. Goldfarb1,2,
- Jonathan Gubbay1,2,
- Peter Tilley1,2,
- Sukh Dhaliwal1,2
- 1Department of Pathology & Laboratory Medicine, Division of Microbiology, Virology & Infection Control, BC Children’s and Women’s Hospital + Sunny Health Center, Vancouver;
- 2Department of Pathology & Laboratory Medicine, Faculty of Medicine, University of British Columbia, Vancouver
- Vijay J Gadkar: Corresponding author;
- High molecular weight DNA extraction from all kingdoms
- BCCH

External link: https://doi.org/10.1186/s44330-025-00024-9
Protocol Citation: Vijay J Gadkar, David M. Goldfarb, Jonathan Gubbay, Peter Tilley, Sukh Dhaliwal 2025. RLB-MALBAC: A non-transposase method for amplifying and barcoding DNA using Multiple Annealing and Looping Based Amplification Chemistry (MALBAC) for Oxford Nanopore sequencing.. protocols.io https://dx.doi.org/10.17504/protocols.io.dm6gpzbr1lzp/v1
Manuscript citation:
Gadkar, V.J., Goldfarb, D.M., Dhaliwal, S. et al. RLB-MALBAC: a rapid, non-transposase method for synthesizing unbiased sequencing libraries from low amount of genomic DNA for Nanopore sequencing. BMC Methods 2, 4 (2025). https://doi.org/10.1186/s44330-025-00024-9
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: April 30, 2024
Last Modified: January 06, 2025
Protocol Integer ID: 99137
Keywords: Nanopore, Transposase, SQK-RPB114.24, Non-tranposase, Vent polymerase, whole genome amplification, quasilinear whole genome amplification, barcoding dna, based amplification chemistry, amplification chemistry, libraries for oxford nanopore, amplification step, amplification protocol, based amplification cycle, amplification cycle, sequencing, amplifying, low amount of input dna, oxford nanopore, input dna, genomic dna of escherichia coli bacteriophage lambda dna, malbac procedure, genomic dna, escherichia coli bacteriophage lambda dna, multiple displacement amplification, protocol of the rlb, viral genome, transcriptomic, malbac protocol
Abstract
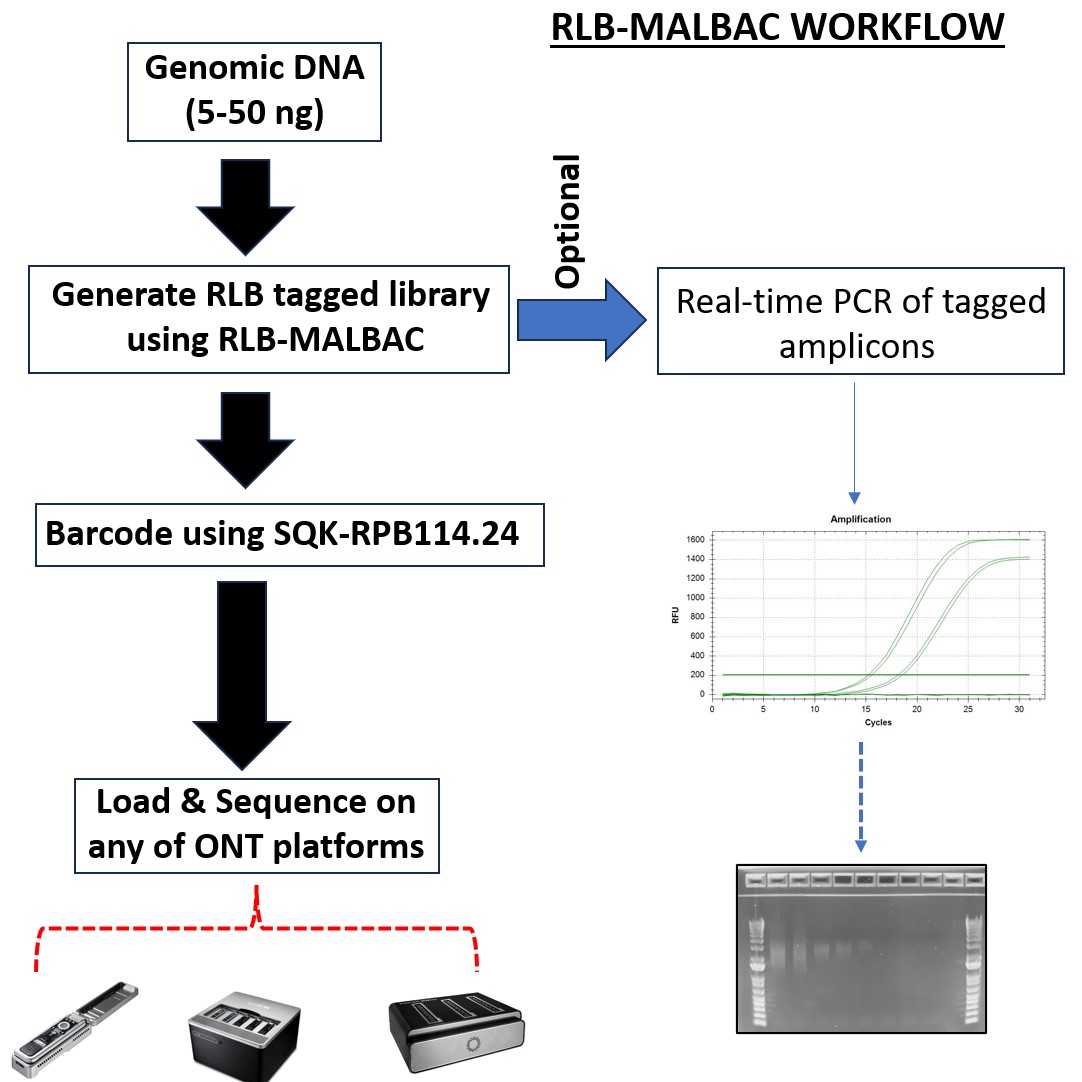
Low amounts of starting material can be a major bottleneck in preparing libraries for Oxford Nanopore sequencing. The only recourse till date has been to implement an amplification step, to “pre-amplify” the low amount of input DNA, with the tagmentation-PCR (SQK-RPB114.24 kit) or multiple displacement amplification (MDA) based workflow, being the two commonly used options. We here demonstrate a novel, PCR based amplification protocol, based on Multiple Annealing and Looping Based Amplification Cycles (MALBAC), a quasilinear whole genome amplification (WGA) method (Zhong et al., 2012). The libraries synthesized with this workflow, are compatible with the SQK-RPB114.24 kits barcoding system only. This protocol, henceforth, referred to as the RLB-MALBAC protocol, can representatively amplify low amounts (2-5ng) of genomic DNA and has successfully been used to sequence bacterial and viral genomes. The versatility of our RLB-MALBAC method, could have broad application in many other areas of biology for e.g., forensics, single cells genomics or transcriptomics, where sample limitation is an major bottleneck. We here present a working protocol of the RLB-MALBAC procedure, using the genomic DNA of Escherichia coli bacteriophage lambda DNA, as an example target template for sequencing.
Materials
Tris-EDTA buffer:
| A | B | |
| Tris-Cl | 10mM | |
| EDTA | 1 mM | |
| pH | 8.0 |
RLB-MALBAC Mastermix reaction volumes:
| A | B | C | |
| Component | Volume (1 reaction) | ||
| 1 | 10X ThermoPol Buffer | 2.5 µL | |
| 2 | 10mM dNTP | 1.0 µL | |
| 3 | 25 µM RLB-N9 primer | 1.0 µL | |
| 4 | 100 mM MgSO4 | 0.5 µL | |
| 5 | Vent (exo-) DNA polymerase | 0.6 µL | |
| 6 | Nuclease Free Water | 15.2 µL |
MALBAC amplification reaction using the following cycling conditions:
| A | B | C | D | E | |
| Cycle | Step | Temperature | Time | ||
| 1 | Denaturation | 94 °C | 5 minutes | ||
| 10 | Annealing | 20 °C | 50 seconds | ||
| Annealing | 30 °C | 50 seconds | |||
| Annealing | 40 °C | 45 seconds | |||
| Annealing | 50 °C | 45 seconds | |||
| Annealing | 60 °C | 30 seconds | |||
| Extension | 65 °C | 2 minutes | |||
| Extension | 70 °C | 2 minutes | |||
| Denaturation | 95 °C | 20 seconds | |||
| Annealing | 60 °C | 30 seconds | Slow ramping rate (0.1 C/ sec) | ||
| Annealing | 55 °C | 30 seconds | |||
| End | 4 °C | ∞ |
RLB-MALBAC mastermix using the reaction volumes :
| A | B | C | |
| Component | Volume (1 reaction) | ||
| 1 | 2X PowerTrack™ SYBR Green Master Mix (Cat No: A46012; ThermoFisher Scientific) | 12.5 µL | |
| 2 | 25 µM RLB-Control primer | 0.2 µL | |
| 3 | RLB-MALBAC library template | 1-2 µL | |
| 4 | Nuclease Free Water | 11.5 µL |
Run the Real-time PCR machine using the cycling paramaters:
| A | B | C | D | E | |
| A | B | C | D | ||
| Cycle | Step | Temperature | Time | ||
| 1 | Denaturation | 95°C | 5 minutes | ||
| 30 cycles | Denaturation | 95 °C | 15 sec | ||
| Anneal | 62°C | 15 sec | |||
| Extend | 65°C | 5 min | Data Acquire | ||
| Final Extension | 65°C | 10 min | |||
| End | 4°C | ∞ |
Components to be added in PCR tube:
| A | B | C | |
| Component | Volume (1 reaction) | ||
| 1 | 2X LongAmp Taq 2X Mastermix (Cat No: M0287S; New England Biolabs) | 25 µL | |
| 2 | RLB (BC01-24) primers (SQK-RPB114.24 kit) | 1.0 µL | |
| 3 | RLB-MALBAC library template | 10 µL | |
| 4 | Nuclease Free Water | 14.0 µL |
Amplify using the following cycling conditions:
| A | B | C | D | E | |
| A | B | C | D | ||
| 1 | Cycle | Step | Temperature(°C) | Time | |
| 2 | 1 | Initial Denaturation | 95°C | 3 minutes | |
| 3 | 15-17 cycles | Denaturation | 95°C | 15 sec | |
| 4 | Anneal | 56°C | 15 sec | ||
| 5 | Extend | 65°C | 6 min | ||
| 6 | Final Extension | 65°C | 6 min | ||
| 7 | End | 4°C | ∞ |
Reagents required
- Nuclease-free WaterThermofisherCatalog #AM9920
- Vent (exo-) DNA Polymerase - 200 unitsNew England BiolabsCatalog #M0257S
- PowerTrack™ SYBR Green Master Mix for qPCRThermo Fisher ScientificCatalog #A46012
- ThermoPol Reaction Buffer Pack - 6.0 mlNew England BiolabsCatalog #B9004S
- Magnesium Sulfate (MgSO4) Solution - 6.0 mlNew England BiolabsCatalog #B1003S
- Deoxynucleotide (dNTP) Solution MixNew England BiolabsCatalog #N0447S /Deoxynucleotide (dNTP) Solution MixNew England BiolabsCatalog #N0447L
- PCRClean DXAline BiosciencesCatalog #C-1003-5
Oligo required:
- RLB-N9 oligo (order from IDT): TTTTTCGTGCGCCGCTTCAACNNNNNNNNN (Claro et al 2023)
- RLB-CTRL oligo: TTTTTCGTGCGCCGCTTCAAC (Claro et al 2023)
- Buffer EBQiagenCatalog #19086
- LongAmp Taq 2X Master Mix - 100 rxnsNew England BiolabsCatalog #M0287S
- RLB (BC01-24) primers (SQK-RPB114.24 kit; Oxford Nanopore)
Troubleshooting
Before start
* Purified genomic DNA from E. coli bacteriophage lambda (Cat No: N3011S; New England Biolabs, Mississauga, ON).
* Prepare 5 mL of fresh 80% ethanol
* Thaw 10X ThermoPol Reaction buffer (B9004S)
* Thaw 100 mM Magnesium Sulfate (MgSO4) solution (B1003S).
* Thaw 10 mM dNTP solution (N0447S/N0447L)
* Thaw 25 µM RLB-N9 primer
* Allow Aline PCRClean DX beads (Cat No: ALI-C1003-5) to warm at room temperature (Aline Biosciences, Waltham, MA).
1.) Preparing input DNA
The E. coli bacteriophage lambda is supplied at 500 ng/µL. Thaw the stock tube and spin the content.
Dilute the stock concentration to 5 ng/µL using 1X Tris-EDTA (pH=8) buffer.
Tris-EDTA buffer:
| A | B | |
| Tris-Cl | 10 mM | |
| EDTA | 1 mM | |
| pH | 8.0 |
Store the diluted DNA at 4 °C till further use.
2.) Preparing working stock of primers
RLB-N9 primer:
- Dissolve the RLB-N9 primer (TTTTTCGTGCGCCGCTTCAACNNNNNNNNN) obtained from the supplier in 1X TE Buffer.
- Prepare a 25 micromolar (µM) working stock by diluting the 100 micromolar (µM) stock.
- Store aliquots at -20 °C .
TE Buffer:
| A | B | |
| Tris-Cl | 10 mM | |
| EDTA | 1 mM | |
| pH | 8.0 |
RLB-CTRL primer:
- Dissolve the RLB-Ctrl primer (TTTTTCGTGCGCCGCTTCAAC) obtained from the supplier in 1X TE Buffer.
- Prepare a 25 micromolar (µM) working stock by diluting the 100 micromolar (µM) stock.
- Store aliquots at -20 °C .
3.) RLB-MALBAC Reaction Setup
3s
In a 200 µL PCR tube, prepare the following RLB-MALBAC mastermix using the reaction volumes detailed in the table below. For more than one reaction multiply the volumes specified for 1 reaction plus one negative control.
RLB-MALBAC mastermix using the reaction volumes:
| A | B | C | |
| Component | Volume (1 reaction) | ||
| 1 | 10X ThermoPol Buffer | 2.5 µL | |
| 2 | 10 mM dNTP | 1.0 µL | |
| 3 | 25 µM RLB-N9 primer | 1.0 µL | |
| 4 | 100 mM MgSO4 | 0.5 µL | |
| 5 | Vent (exo-) DNA polymerase (2U/µL) | 0.6 µL | |
| 6 | Nuclease Free Water | 14.4 µL |
Mix the contents of the RLB-MALBAC mastermix by vortexing it for 00:00:03 and spin down the contents at the bottom of the tube.
3s
Dispense 20 µL of the RLB-MALBAC mastermix into individual PCR tube and to each tube add 5 µL of lambda DNA (5 ng/µL)
Vortex the reaction contents and spin down to collect it bottom of the tube.
Transfer the tubes to a PCR Thermocycler and start the MALBAC amplification reaction using the following cycling conditions:
| A | B | C | D | E | |
| A | B | C | D | ||
| Cycle | Step | Temperature | Time | ||
| 1 | Denaturation | 94 °C | 5 minutes | ||
| 10 | Annealing | 20 °C | 50 seconds | ||
| Annealing | 30 °C | 50 seconds | |||
| Annealing | 40 °C | 45 seconds | |||
| Annealing | 50 °C | 45 seconds | |||
| Annealing | 60 °C | 30 seconds | |||
| Extension | 65 °C | 2 minutes | |||
| Extension | 70 °C | 2 minutes | |||
| Denaturation | 95 °C | 20 seconds | |||
| Annealing | 60 °C | 30 seconds | Slow ramping rate (0.1 C/ sec) | ||
| Annealing | 55 °C | 30 seconds | |||
| End | 4 °C | ∞ |
The reaction samples can be stored at 4 °C for 168:00:00 or-20 °C for long term storage.
1w
4.) RLB-MALBAC Library Clean-up
22m 15s
Prepare Aline PCRClean DX beads (Cat No: ALI-C1003-5) for use; resuspend by vortexing.
Add 25 µL of the Aline PCRClean DX beads to the RLB-MALBAC PCR reaction.
Mix thoroughly by pipetting up and down 5-10 times.
Incubate at Room temperature for00:05:00 .
5m
Spin down the tube for 00:00:05 and place the tube on the magnetic separator for 00:05:00 or until the solution clears. Beads should be on the side of the tube.
5m 5s
With the tube still on the magnet, remove the liquid from the tube and discard.
Note
Be sure not to disturb the beads.
With the tubes still on the magnet, add 500 µL of 80% ethanol to the tube and let it sit for 00:02:00 . Try to minimize disturbing the beads.
2m
Remove the ethanol by pipetting and discard.
Repeat the 80% Ethanol wash step.
Dry the beads by incubating the tube for 00:10:00 at Room temperature . Ensure all the ethanol has evaporated from the tube.
Note
If there is visible ethanol in the tube, remove it by spinning the tube for 00:00:10 , putting the tube back on the magnet and removing the excess ethanol. Make sure the beads don't over dry
10m
When the beads are dry and there is no visible sign of ethanol, suspend the beads in 10 µL of EB buffer (Cat No: 19086; Qiagen). Proceed to next step 6 (Barcoding of RLB-tagged amplicons with SQK-RPB114.24 kit system) or perform the optional step 5 (Check incorporation of RLB-tag sequence).
5.) Check Incorporation of RLB-tag sequence (optional)
To test the incorporation of the RLB tag in the target DNA, real-time PCR reaction can be setup to amplify the RLB-MALBAC amplicons using the RLB-PCR control primer.
- In a 200 µL PCR tube, prepare the following RLB-MALBAC mastermix using the reaction volumes detailed in the table below. For more than one reaction multiply the volumes specified for 1 reaction plus at least two negative controls.
RLB-MALBAC mastermix using the reaction volumes :
| A | B | C | |
| Component | Volume (1 reaction) | ||
| 1 | 2X PowerTrack™ SYBR Green Master Mix (Cat No: A46012; ThermoFisher Scientific) | 12.5 µL | |
| 2 | 25 µM RLB-Control primer | 0.2 µL | |
| 3 | RLB-MALBAC library template | 1-2 µL | |
| 4 | Nuclease Free Water | 11.5 µL |
Transfer the tube to Real-time PCR machine and run it using the following cycling parameters:
| A | B | C | D | E | |
| A | B | C | D | ||
| Cycle | Step | Temperature | Time | ||
| 1 | Denaturation | 95°C | 5 min | ||
| 30 cycles | Denaturation | 95 °C | 15 sec | � | |
| Anneal | 62°C | 15 sec | |||
| Extend | 65°C | 5 min | Data Acquire | ||
| Final Extension | 65°C | 10 min | |||
| End | 4°C | ∞ |
If the RLB-MALBAC step was successful, the amplicons will have the RLB tags at both its ends. As a result, the RLB control PCR will give an amplification curve for the tagged template. No amplification will be observed for water controls.
6.) Barcoding of RLB-tagged amplicons with SQK-RPB114.24 kit system
Setup barcoding reaction using the RLB-BC 01 to 24 series barcodes, provided in the SQK-RPB114.24 kit. The purified RLB-tagged amplicons from step 4 (RLB-MALBAC Library Clean-up).
In a clean 200 µL PCR tube, add the following components:
| A | B | C | |
| Component | Volume (1 reaction) | ||
| 1 | RLB (BC01-24) primers (SQK-RPB114.24 kit) | 1.0 µL | |
| 2 | RLB-MALBAC library template | 10 µL | |
| 3 | Nuclease Free Water | 14.0 µL | |
| 4 | 2X LongAmp Taq 2X Mastermix (Cat No: M0287S; New England Biolabs) | 25 µL |
Mix gently by flicking the tube and spin down the contents.
Transfer the PCR tube to a thermocycler and amplify using the following cycling conditions:
| A | B | C | D | |
| Cycle | Step | Temperature ( °C) | Time | |
| 1 | Initial Denaturation | 95 °C | 3 minutes | |
| 15-17 cycles | Denaturation | 95 °C | 15 sec | |
| Anneal | 56 °C | 15 sec | ||
| Extend | 65 °C | 6 min | ||
| Final Extension | 65 °C | 6 min | ||
| End | 4 °C | ∞ |
Procced with the purification and library pooling (if multiple samples) protocol as outlined in the Oxford Nanopore's SQK-RPB114.24 kit.
Protocol references
Citations:
1. Zong, C.; Lu, S.; Chapman, A.R.; Xie, S. (2012). "Genome-wide detection of single-nucleotide and copy-number variations of a single human cell." Science 338, 1622.
2. Claro IM, Ramundo MS, Coletti TM, da Silva CAM, Valenca IN, Candido DS, Sales FCS, Manuli ER, de Jesus JG, de Paula A, Felix AC, Andrade PDS, Pinho MC, Souza WM, Amorim MR, Proenca-Modena JL, Kallas EG, Levi JE, Faria NR, Sabino EC, Loman NJ, Quick J. (2023) Rapid viral metagenomics using SMART-9N amplification and nanopore sequencing. Wellcome Open Res. 24; 6:241.
