Sep 08, 2025
REVA #4: Labeling of Vagus Nerve Subjects, Samples, and Paraffin Blocks
- Nicole A Pelot1,
- Noa B Nuzov2,
- Jennifer J Coleman2,
- Andrew J. Shoffstall2,3
- 1Department of Biomedical Engineering, Duke University, Durham, NC, USA, 27708;
- 2Department of Biomedical Engineering, Case Western Reserve University, Cleveland, OH, USA, 44106;
- 3APT Center, Louis Stokes Cleveland Department of Veterans Affairs Medical Center, Cleveland, OH
- Nicole A Pelot: ORCID: 0000-0003-2844-0190;
- Noa B Nuzov: ORCID: 0000-0001-8187-2115;
- Jennifer J Coleman: 0000-0001-6702-729X
- Andrew J. Shoffstall: ORCID: 0000-0002-0881-2180

Protocol Citation: Nicole A Pelot, Noa B Nuzov, Jennifer J Coleman, Andrew J. Shoffstall 2025. REVA #4: Labeling of Vagus Nerve Subjects, Samples, and Paraffin Blocks. protocols.io https://dx.doi.org/10.17504/protocols.io.n92ld6r3xg5b/v1
Manuscript citation:
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: July 15, 2025
Last Modified: September 08, 2025
Protocol Integer ID: 225835
Keywords: Human anatomy, Gross anatomy, Vagus nerve, Cranial nerves, Peripheral nervous system, Autonomic nervous system, Neuroanatomy, Grossing, reva tissue labels with the subject, reva standards for anatomical labeling, defining reva tissue label, labeling of vagus nerve subject, anatomical labeling, reva standard, vagus nerve subject, vagus nerve, label, labeling, reva, histology slide, paraffin block, paraffin blocks this protocol, sample
Funders Acknowledgements:
NIH SPARC REVA
Grant ID: 75N98022C00018
Disclaimer
DISCLAIMER – FOR INFORMATIONAL PURPOSES ONLY; USE AT YOUR OWN RISK
The protocol content here is for informational purposes only and does not constitute legal, medical, clinical, or safety advice, or otherwise; content added to protocols.io is not peer reviewed and may not have undergone a formal approval of any kind. Information presented in this protocol should not substitute for independent professional judgment, advice, diagnosis, or treatment. Any action you take or refrain from taking using or relying upon the information presented here is strictly at your own risk. You agree that neither the Company nor any of the authors, contributors, administrators, or anyone else associated with protocols.io, can be held responsible for your use of the information contained in or linked to this protocol or any of our Sites/Apps and Services.
Abstract
This protocol describes the rules for defining REVA tissue labels with the subject, sample, and block. These labels are used on containers, paraffin blocks, histology slides, data folder and file names across modalities, and metadata. This protocol does not include the REVA standards for anatomical labeling of nerves and branches.
Image Attribution
Diagram by Nicole Pelot.
Guidelines
N/A
Materials
N/A
Troubleshooting
Safety warnings
This protocol might include items and/or substances that may pose hazards (e.g., chemical, physical, biological, or otherwise) to your health upon use or exposure. Before engaging in the processes described in this protocol, familiarize yourself with and follow the safety data sheets, manufacturer safety recommendations, and local regulations.
Ethics statement
Be sure to seek approval for or an exemption from human subjects research from your local regulatory body(ies) as required by local and/or institutional regulations before initiating studies.
This study was determined to be exempt from IRB oversight by the Case Western Reserve University Institutional Review Board (IRB) because it involved de-identified cadaveric tissue and no protected health information was collected from the donors.
Section 1: Vagus Nerve Tissue Labeling
The label for a piece of vagus nerve tissue has up to four elements, and their definitions are detailed in the rest of the protocol. Briefly:
Project prefix
Cadaver number
Nerve “sample”, labeled with two letters for the anatomical region (superior-inferior division and transverse division (right/left, anterior/posterior)) and a number (one digit: 1, 2, 3, 4). Each sample is glued to an acrylic board.
Battleship label to label subdivisions of a given sample, based on the grid etched into the acrylic board, with two digits based on the row (01, 02, 03, ...) and one lowercase letter based on the column (a, b, c, d, ...).
Use dashes (not underscores) to separate label elements (except the project prefix and cadaver number, which are written sequentially).
The tissue label should be used for traceability: to label tissue containers, to define folder and file names, to name tissue/data in the metadata, to map and create linkages across datasets (i.e., different imaging modalities), etc.
Project prefix: The prefix of each label is “SR” for “SPARC REVA”; non-REVA cadavers (used for pilot testing, protocol development, and other projects) have the prefix “NR” for “non-REVA”.
Cadaver number: Each subject is assigned a three-digit identifier (e.g., 001, 002, 003, ...).
Superior-inferior regions: The nerve is divided into four superior-inferior sections (Figure 1).
Figure 1: Superior-inferior and transverse (right-left and anterior-posterior) regions to define sample labeling.
C: Cervical; from the medulla to the superior border of the clavicle.
T: upper Thoracic; from the superior border of the clavicle to 1 cm superior to the superior end of the esophageal plexus.
E: Esophageal plexus; from 1 cm superior to the superior end of the esophageal plexus to the esophageal hiatus.
S: Sub-esophageal, i.e., inferior (“sub”) to the esophageal plexus (which might end in the lower thoracic region or in the abdomen); from 1 cm inferior to where the anterior and posterior abdominal trunks form to the most distal ends of the abdominal vagal trunks and branches (distal end may vary).
Transverse regions: The nerve is divided into transverse sections (right/left halves for cervical and upper thoracic divisions; anterior/posterior halves for esophageal plexus and sub-esophageal divisions) (Figure 1).
R: right
L: left
A: anterior
P: posterior
Anatomical regions: There are eight anatomical regions per cadaver, and each is labeled with two letters—the superior-inferior nerve section label followed by the transverse label (Figure 2): CR, TR, CL, TL, EA, EP, SA, SP.
Figure 2: Anatomical regions to define sample labeling. See definitions for acronyms in the protocol text.
Sample label: Each anatomical region is cut into ≤9 cm-long samples to fit in the microCT scanner. Each sample label has two letters (based on the anatomical region described above) and one digit (to ensure uniqueness with multiple samples in a given anatomical region for a given cadaver). If a given region is already
9 cm, it will still be given a number (e.g., SA1 remains SA1 and does not become SA2 if it is alone, i.e., there is no SA2). The numerical order for CR, CL, TR, TL, EA, and EP will always be increasing in the direction of superior to inferior (e.g., CL1 and CR1 will always contain the proximal-most segment that leaves the medulla). The numerical order for the SA and SP regions will not be in a consistent orientation for ordering (e.g., can be vertically stacked or side-by-side); consult the labeled photo from the anatomy data of the excised nerve to determine the orientation of the SA and SP sections relative to each other and the rest of the nerve. There are 8 to 24 samples per nerve (with 2 to 4 samples per anatomical region):
CR1, CR2, CR3
TR1, TR2, TR3
CL1, CL2, CL3
TL1, TL2, TL3
EA1, EA2
EP1, EP2
SA1, SA2, SA3, SA4
SP1, SP2, SP3, SP4
Sample cutting
The vagal complex of each cadaver—spanning skull to abdomen, left and right—is removed intact, in one piece, and laid on a series of adjacent acrylic boards.
Cut the tissue (between acrylic boards) between the thoracic and esophageal anatomical regions on the right and left (i.e., two cuts) before staining with phosphotungstic acid (PTA) because the cervical/thoracic samples and the esophageal/abdominal samples require different PTA staining times.
After PTA staining, place all acrylic boards back in their correct places to reassemble the nerve for low-resolution computed tomography (CT) imaging.
After CT, cut the tissue to separate all samples (i.e., all acrylic boards) before microCT imaging.
Use a new razor blade for every single cut to reduce tissue deformation.
Battleship label (Figure 3): After removal from the cadaver, the vagal complex is glued to a series of acrylic boards; each board is 3 cm wide and up to 9 cm long. Each acrylic board is etched with a 5 mm x 5 mm grid and the sample label (CL1, CL2, ...; as defined above) on both the front and back. The grid is oriented with the diagonal notch in the top left, and the nerve is placed vertically in the center of the grid. We aim to align the vertical (superior-inferior) grid lines across boards for a given nerve. The nerve on the gridded acrylic boards is captured in post-dissection photos.
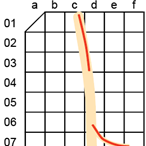
Figure 3: Schematic of a nerve (beige) glued to an acrylic backing board, etched with a 5 mm x 5 mm grid. The acrylic backing boards are 3 cm wide and up to 9 cm long, constrained by the tubes used to hold the samples for microCT imaging. The red lines and nearby labels provide example nerve segments that are grossed for embedding into one paraffin block; each block is labeled according to the “battleship” coordinates of the segments’ location on the grid.
The columns of the grid of each acrylic board are labeled with a lowercase letter (a, b, c, d, e, ...) and the rows are labeled with a two-digit number (01, 02, 03, 04, 05, ...).
Therefore, a given grid square is identified by two digits followed by one lowercase letter (e.g., 01b).
Histology grossing:
Gross each trunk and each branch into 15 mm segments (minimum length of 5 mm) as possible. Each segment is destined for one paraffin block.
Cut each 15 mm segment into three 5 mm pieces, as possible, that are embedded side-by-side in one paraffin block, oriented such that the superior nerve surface of each piece is sliced first.
Block labeling for vagal and sympathetic trunks:
Vagal and sympathetic trunks course approximately vertically (superiorly-inferiorly) along the gridded acrylic boards. The vagus is centered (right-left) on the acrylic board, and the sympathetic trunk is placed off-center.
Label the paraffin block using the grid squares spanned by the nerve segment, specifically, three characters of the top-left square followed by the three characters of the bottom-right square (e.g., SR002-CL1-01b05d; or SR002-CL1-01b03b if spanning only one column (b)).
Block labeling for vagal branches and non-vagal nerves:
Vagal branches and non-vagal nerves (e.g., cranial and spinal nerves that have connecting branches with the vagus nerve) typically course diagonally on the gridded acrylic boards.
Label the paraffin block using the grid square where the segment connects to the vagus trunk, followed by the string “side” (e.g., SR002-CL1-18cside).
Section 2: Example Labels
MicroCT
SR019-CL1: SPARC REVA, cadaver/subject 019, most superior (#1) sample of left cervical (CL) anatomical region.
Histology
SR005-EA1-01c03c: SPARC REVA, cadaver/subject 005, esophageal plexus, anterior, first sample, nerve segment spans 15 mm of one column that goes from 01c to 03c.
SR001-TR1-20d23e: SPARC REVA, cadaver/subject 001, upper thoracic, right side; nerve segment spans from grid square 20d (top left) to 23e (bottom right).
SR004-CL2-03cside: SPARC REVA, cadaver/subject 004, lower cervical left, nerve segment is a branch (or non-vagal nerve) that comes off the vagal trunk at the grid point 03 column c.
Acknowledgements
The authors thank the donors and staff of the Case Western Reserve University Anatomical Gift Program. Without the selfless donations of the donors and the tireless efforts of the staff of this program, the methods described in this protocol and the insights and advancements that result from studies conducted according to this protocol would not be possible.
