Mar 31, 2026
Quantification of P. salmonis Intracellular Replication by Genome Equivalents
- Pamela Aravena1,
- Javiera Ortiz Severín2,
- Verónica Cambiazo2
- 1Universidad de Chile;
- 2University of Chile

Protocol Citation: Pamela Aravena, Javiera Ortiz Severín, Verónica Cambiazo 2026. Quantification of P. salmonis Intracellular Replication by Genome Equivalents. protocols.io https://dx.doi.org/10.17504/protocols.io.36wgqq82ogk5/v1
Manuscript citation:
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: June 03, 2025
Last Modified: March 31, 2026
Protocol Integer ID: 219442
Keywords: Piscirickettsia salmonis, Intracellular infection, qPCR, Taqman probe, genome equivalents this protocol, genome equivalent, genome, qpcr, gene
Funders Acknowledgements:
ANID FONDECYT
Grant ID: 1211893
Millennium Science Initiative Program
Grant ID: ICN2021_044
Abstract
This protocol describes a method for accurate quantification of P. salmonis cells based on the number of copies of its genome identified by qPCR, using specific primers and probes targeting the single copy gene recF.
Image Attribution
Pamela Aravena
Materials
Leibovitz's L15 mediumLeibovitz's L-15 Medium, powderThermo FisherCatalog #41300039
Gentamicin Gentamicin SolutionMerck MilliporeSigma (Sigma-Aldrich)Catalog #G1397
Tryptic Soy BrothTryptic Soy BrothMerck MilliporeCatalog #1.05459
NaCl Sodium Chloride (NaCl)Merck MilliporeSigma (Sigma-Aldrich)Catalog #S3014
L-cysteine L-CysteineMerck MilliporeSigma (Sigma-Aldrich)Catalog #C7352
FeCl3Iron(III) chloride hexahydrateMerck MilliporeSigma (Sigma-Aldrich)Catalog #44944
PBSPhosphate-Buffered Saline, 1X without calcium and magnesium, PH 7.4 ± 0.1CorningCatalog #21-040-CV
FBS Fetal Bovine SerumThermo ScientificCatalog #A5209502
trypsin-EDTA Trypsin-EDTA (0.25%), phenol redGibco - Thermo Fisher ScientificCatalog #25200056
Blood & Tissue kit Qiagen DNeasy blood and tissue genomic DNA isolation kitQiagenCatalog #69504
TaqMan Gene expression assay kit Invitrogen TaqMan Universal PCR Master Mix (2X)Life TechnologiesCatalog #4304437
GoTaq Green Master Mix Promega GoTaq Green Master MixPromegaCatalog #M7122
nuclease free water Nuclease free water
Wizard SV Gel and PCR Clean-Up System Kit Promega Wizard SV Gel and PCR Clean-Up SystemPromegaCatalog #A9281
Troubleshooting
Before start
Grow a pure culture of P. salmonis for infecting SHK-1 monolayers
Infection of SHK-1 cells
3d
Infect SHK-1 cell cultures at 80% confluence with stationary state bacterial cultures using MOI 100:1 (bacteria:cell).
After 3 days of infection, wash the cultures with PBS and incubate 40 mins with Leibovitz's L15 medium with gentamicin (100 µg/mL) at 18 ºC.
3d
After incubation, wash the cells with PBS and incubate at 18 ºC with fresh Leibovitz's L15 medium supplemented with 5% of FBS, without antibiotics.
Purification of total genomic DNA (gDNA)
Select appropriate infection times before harvesting infected SHK-1 cells.
To harvest infected cells, wash the cell monolayers with cold PBS 1X and incubate with gentamicin (100 µg/mL) for 40 min to kill extracellular bacteria.
Wash the cells with cold PBS and then treat them with 0.25% trypsin-EDTA solution for 30 min at room temperature (RT).
Disrupt the monolayer by pipetting and collect the cells in a 15 mL sterile tube.
Centrifuge at 12,000 x g for 15 min and resuspend the pellets in 200 µL of PBS.
Isolate total genomic DNA (gDNA) from three independent cultures of infected SHK-1 cells per time, using the Blood & Tissue kit (Qiagen, USA) as recommended by the manufacturer.
Evaluate the concentration and the quality of the DNA by measuring the absorbance (260/280 nm) in each sample using a NanoQuant Spectrophotometer (Tecan Technologies).
Adjust the gDNA samples to a concentration of 10 ng/µL.
qPCR quantification using Taqman probes
qPCR assays are conducted using the TaqMan Gene expression assay kit (Invitrogen), and a TaqMan probe and primers (Table 1) targeting the single copy recF gene (GenBank Accession Number NZ_CP039032.1).
| A | B | C | |
| Name | Sequence 5´- 3´ | Function | |
| PS-recF-Fw_Complete | CTTTCGCTCTTCAGGTTGGC | For PCR. To amplify the complete sequence of recF gene | |
| PS-recF-Rv_Complete | ACTCTTATTTTCATCCCAACCAGCAT | For PCR. To amplify the complete sequence of recF gene | |
| recF_TP_Fwd | AAAATTTTGTTTAGCAATATTAGCGTTGTTTTT | For qPCR. To amplify a fragment of recF gene | |
| recF_TP_Rv | ACTCTTATTTTCATCCCAACCAGCAT | For qPCR. To amplify a fragment of recF gene | |
| recF_TP_Probe | FAM-ACGCCCAATTTTCA-NFQ | For qPCR. To hybridize with the recF fragment |
Table 1. Primers and probes used for conventional PCR and qPCR, used in this protocol.
Create a calibration curve using the gel-purified PCR product of the gene recF as templates for the qPCR reaction.
PCR product is obtained after PCR amplification of recF from purified P. salmonis CGR02 genome using GoTaq Green Master Mix (Promega), containing 200 ng of bacterial DNA, 12.5 µL of GoTaq mix (Promega, Wisconsin, United States), 5.5 µL of nuclease free water and 1 µL of each primer recF-complete (10mM – Table 1). The PCR amplification was performed in T100 (Bio-Rad) Thermal cycling controller with the following protocol: 10 min at 95 °C, 30 cycles of 95 °C for 60 s, 60 °C for 30 s and 72 °C for 50 s, and a final extension at 72 °C for 10 min. The PCR product (884 bp) was purified using Wizard SV Gel and PCR Clean-Up System Kit (Promega).
Quantify the product concentration by Qubit fluorometric quantitation and serially dilute to 10^6 and 10^0 copies of recF (see equation 1) before using as a template to generate standard curves for qPCR assays.
To convert to ng of DNA to copy number, the formula used was:
Equation 1
Set up the qPCR reactions containing 0.9 μM of each primer, 0.25 μM of probe, 10 µL of TaqMan Gene expression assays Master Mix and 10 ng of recF PCR product as template, in a total volume of 20 µL.
Use the following thermal conditions in an AriaMx Pro thermal cycler (Agilent Technologies): initial incubation at 50 °C for 2 min, followed by polymerase activation at 95 °C for 10 min, and 40 amplification/extension cycles of 95 °C for 15 s and 60 °C for 60 s.
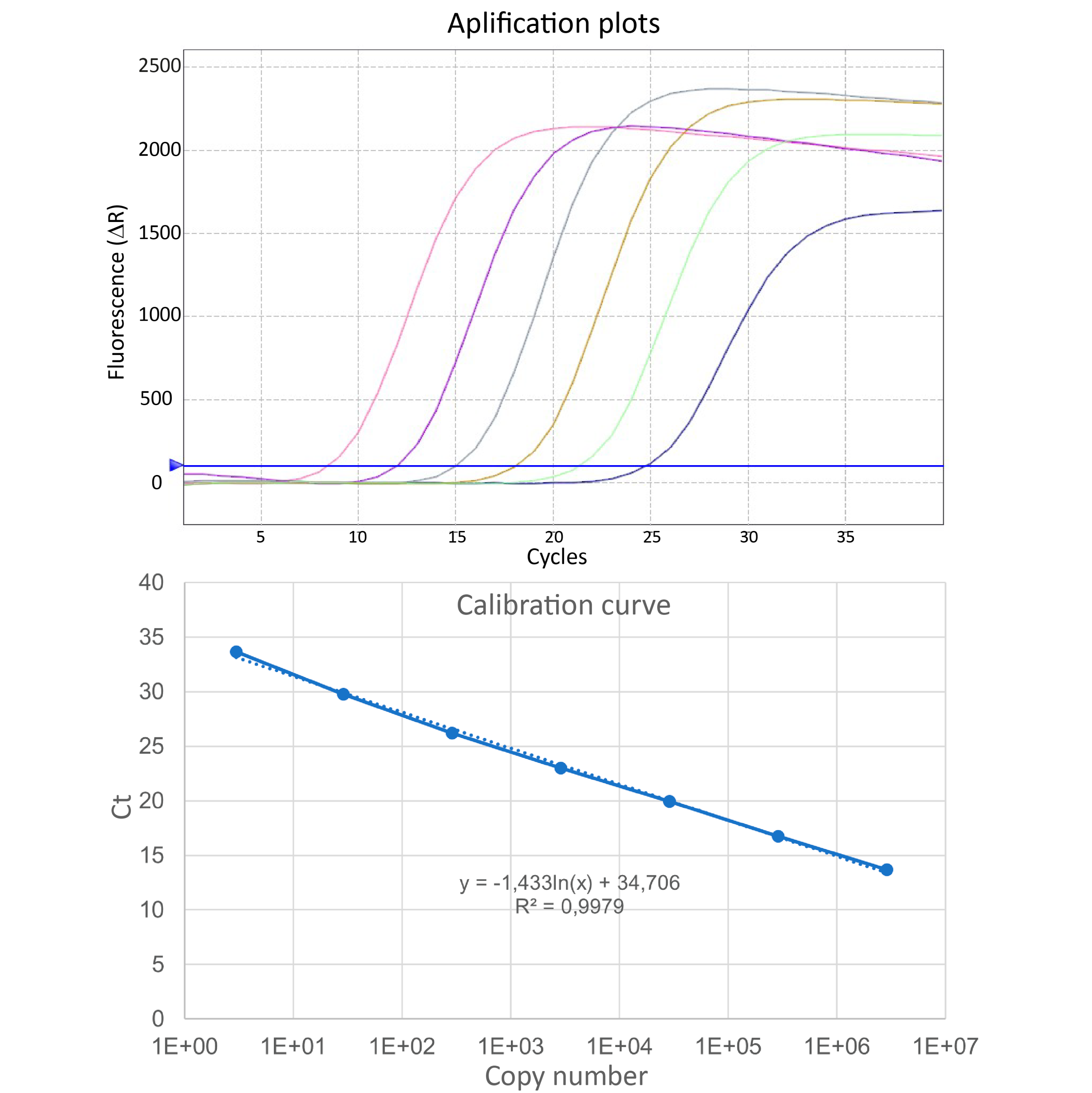
Use the resulting fluorescent plots to create the calibration curve (Ct values vs. recF copy numbers) and estimate numbers of P. salmonis genomes in the experimental samples.
Figure 1. Calibration curve of recF gene with Taqman Probe.
To calculate the Genome Equivalents (GE) in the infected SHK-1 cells, set up qPCR reactions as mentioned above but replace the recF PCR product with the gDNA from the infections.
Interpolate the Ct values in the calibration curve. The conversion from copy number to genome equivalent is based on the presence of only one copy of recF gene in the P. salmonis genome.
