Jul 27, 2025
Version 1
Quantification and Limitations of PSD95-NR2B Clustering using Proximity Ligation Assay in Primary Neuronal Cultures V.1
- Suba Soundarya Santhi Anandhan1
- 1King Abdullah University of Science and Technology

Protocol Citation: Suba Soundarya Santhi Anandhan 2025. Quantification and Limitations of PSD95-NR2B Clustering using Proximity Ligation Assay in Primary Neuronal Cultures. protocols.io https://dx.doi.org/10.17504/protocols.io.q26g79orqvwz/v2Version created by Suba Soundarya Santhi Anandhan
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: June 12, 2025
Last Modified: July 27, 2025
Protocol Integer ID: 222917
Keywords: Proximity Ligation Assay, PSD95, NR2B, high-throughput screening, protein-protein interactions, confocal microscopy, primary mouse neurons, optical plate, imaging, synapses, synaptic protein, reliable tool for synaptic protein interaction, primary neuronal cultures synaptic clustering, synaptic protein interaction, throughput proximity ligation assay, nr2b receptor stabilisation, nr2b clustering, psd95 interaction, synaptic connection, protein interaction, using proximity ligation assay, molecular interactions in the primary mouse, synaptic connections for plasticity study, crucial role in neuronal function, protein, limitations of psd95, quantification of clustering, inconclusiveness highlights biological limitation, neuronal cell, psd95, molecular interaction
Disclaimer
This protocol and its interpretations were independently developed and written by the author based on own experimental work and observations. All included images are unpublished, generated solely by the author, and are not intended for future publication. This document reflects the author’s own methodological optimizations and analysis and is intended purely as a technical reference. It does not represent the official views or findings of any affiliated institution or research group.
Abstract
Synaptic clustering of proteins plays a crucial role in neuronal functions. These protein interactions are not only technically challenging to quantify but time-consuming even with advanced super-resolution tools. To study the NR2B receptor stabilisation and clustering with PSD95, we employed an optimised high-throughput Proximity Ligation Assay (PLA) protocol for the 96-well optical plate. This is a reproducible protocol with consistent results across replicates. It also enables us to quantify molecular interactions in the primary mouse neuronal cells, with punctate analysed in Opera Phenix using its integrated quantification software. Key improvements in fixing, permeabilisation and nuclear counterstaining for high-throughput compatibility have been made. This method is technically robust, time efficient and provides a reliable tool for synaptic protein interactions. While applied to study NR2B-PSD95 interactions, the quantification of clustering was non-significant throughtout different treatments. These results are from iterated experiments (n=3), where inconclusiveness highlights biological limitations rather than technical. This document includes plausible interpretations of the equivocal data and outlines the use of this protocol to quantify pre and post synaptic connections for plasticity studies.
Attachments
Image Attribution
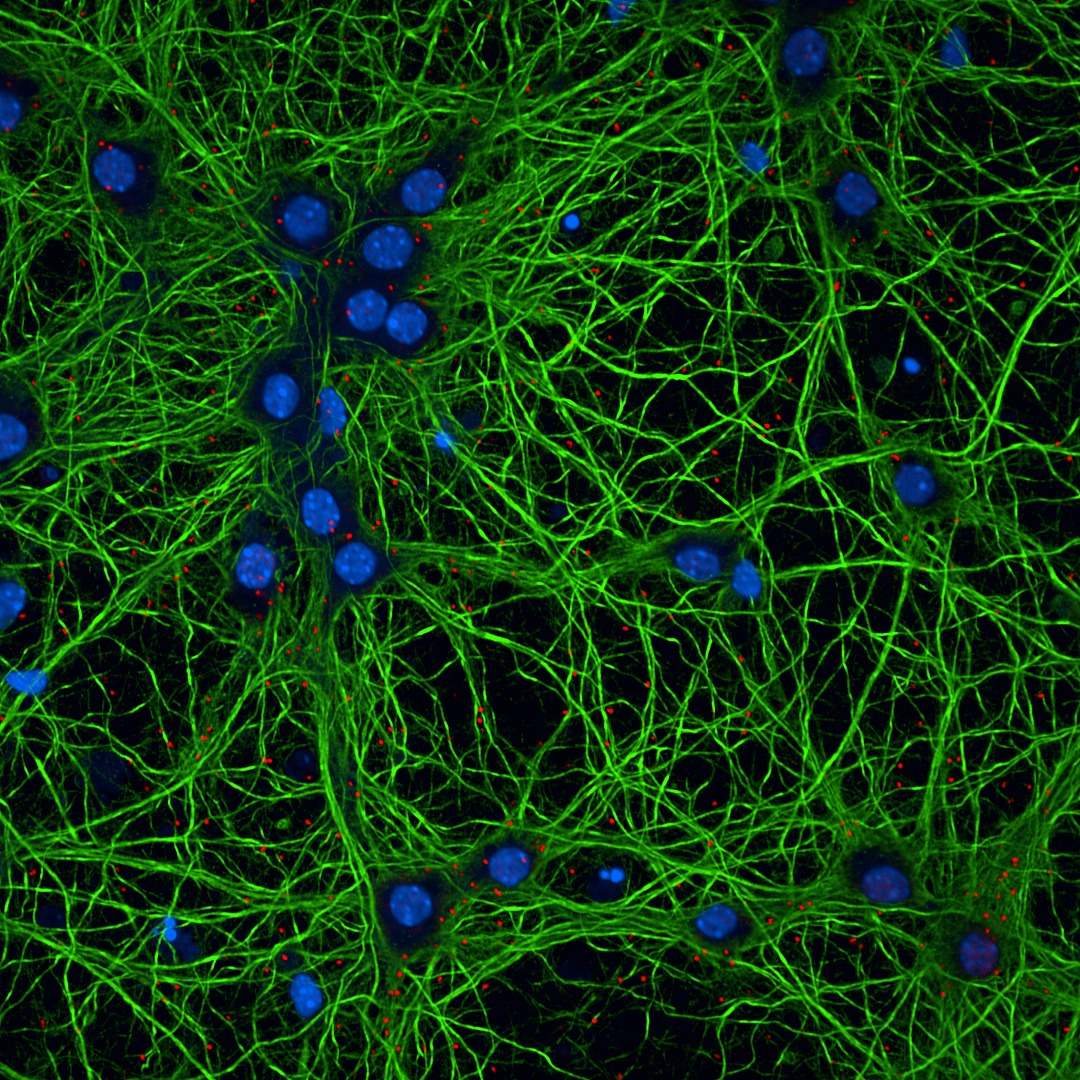
Representative image of primary mouse cortical neurons fixed and stained for PLA (red), MAP2 (green), and nuclei (blue). Puncta indicate protein–protein interactions analyzed using the described PLA protocol.
Guidelines
Principle of the experiment:
Proximity ligation assay enables the study of protein-protein interactions with spatial resolution, typically within 40nm distance. This method employs a secondary probe conjugated with unique DNA sequence that ligates to form a circular DNA with its complementary strand within proximity. When two proteins are brought together, the circular DNA is amplified using rolling circle replication, generating multiple copies of the DNA. Fluorescently, labelled oligonucleotides hybridise to the amplified product, producing a signal, visualised as puncta in confocal microscope.
Cell culture guidelines:
(Below mentioned are optimised and practiced protocol)
Primary neurons were dissociated from E17.5, CD1 embryos.
PLATING:
Prepare the NB plus (Cat. A#3582901) supplemented with B27+ (Cat. A#3582801), Pen/strep and Glutamax (Cat. #35050-038).
1) Coat the 96-well Griener senso plate (Cat. #655892) with 0.2mg/ml of Poly L-ornithine (Cat. P#3655-100mg) overnight at room temperature or in the incubator.
2) Wash the plates with autoclaved milliQ water twice. Perform the third wash using the media (150µl) that is used for plating. Aspirate the media just before the plating and avoid drying. This helped me achieve primary cell attachment to the glass bottom.
3) Plate 10,000 cells/well using the eppendorf multi pipette hand dispenser. Gently mix the cells before plating for even distribution.
4) Leave the plate in the hood with the lid, unhandled for 5min before moving it to the incubator. Immediate handling and placing in the incubator may not help with even surface attachment.
Note
In the 96-well, fill 200µl of the milliQ or cell culture grade water in the outer wells, and fill 100µl of water in the chimney wells to prevent evaporation of the media. Do not handle the plate frequently except for media change or a quick check.
MEDIA CHANGE:
For experiments conducted on DIV 14, change 50% of the media on DIV 7.
Neurons looked consistently unhealthy on DIV 17, after media change on DIV 7 and DIV 14.
Expected result
This PLA protocol was performed following the treatment of DIV14 primary mouse neurons with the exogenous compounds. When target proteins, PSD95-NR2B are within the proximity, a punctate is formed, visualised and quantified. Data were normalised using either, punctate/nuclei or punctate/area ratios. While using the puncta/nuclei, try to eliminate non-neuronal cell types that's devoid of MAP2 stain to ensure the neuronal specificity.
The normalised results from the independent biological replicates were analysed using One-way ANOVA. There was no statistical significance (p>0.05) across different treatment conditions. Random fields at identical positions were imaged for each treatment and technical replicates. The number of nuclei, punctate and the area of MAP2 compared within the biological replicates had no statistical significance.
These ambiguous results were concluded after multiple attempts of different experiments. This suggests a possible biological limitations rather than technical inconsistencies. The potential interpretations are explained below.
Limitations to study PSD95-NR2B interactions in primary neuronal cultures:
1) In vitro primary neuronal cultures lack the synaptic and structural organisations unlike brain slices. The lack of spatial arrangement and synaptic architecture may lead to variability in proximity based assays. Notably, this specific interactions have been successfully studied using PLA in brain tissues cited below.
Citation
LINK
2) The overlap of synaptic zones and dendritic shaft region can result in mixed signals. This introduces variability in data interpretation.
3) Choosing a protein non-native to the sub organelle, selectively or sporadically localised is imperative. In this experiment, NR2B is ubiquitously present which introduces variability making it difficult to distinguish synaptic versus extra synaptic interactions with PSD95.
4)Technically, PLA detects proteins present within the proximity. However, the presence of a PLA signal does not necessarily confirm, whether it is a functional or spatial interaction. This spatial interaction does not reflect biological relevance.
Citation
LINK
Citation
LINK
Conclusion:
Proximity Ligation Assay (PLA) is a powerful and sensitive method to study protein-protein interactions at sub cellular level. However, the current study highlights the limitations of assessing PSD95-NR2B in primary neuronal culture. In vitro system lacks synaptic architecture, extra-synaptic presence of NR2B makes it difficult to distinguish synaptic connections and its functional interaction. Furthermore, addition of synaptic markers such as homer or synapsin for normalisation may not fully resolve this issue. In cultured neurons, overlap of neurite network can lead to false positives. It is therefore advisable to confirm PLA findings with additional methods like co-immunoprecipitation, before drawing a definitive conclusion. This protocol is well suited for pre and post-synaptic interaction rather than receptor trafficking studies. Although, PLA is claimed to have a single molecule resolution, it is not suitable for receptor dynamics in primary neuronal culture. Advanced techniques like single molecule localisation microscopy or super-resolution microscopy are better suited to investigate nano domains and receptor organisation at synapses.
Materials
Requirements:
- Chicken Anti-MAP2 - Invitrogen PA1-10005
- Mouse Anti-PSD95 - Invitrogen MA1-046
- Paraformaldehyde - EMS Cat. #1571
- 10xPBS - Thermo scientific #70011069
- Rabbit Anti-NR2B - Merck #06-600
- Saponin - Sigma #84510
- Donkey anti-rabbit PLA (+) - DUO9200
- Donkey anti-mouse PLA (-) - DUO92004
- Goat anti-chicken Alexa 488 - ab #150173
- HOECHST - Thermo scientific #62249
- Neurobasal plus medium - Thermo scientific A#3582901
- B27 plus - Thermo scientific A#3582801
- Glutamax - Thermo scientific #35050-038
- 96-well Griener senso plate - #655892
- Poly L-ornithine - Sigma P#3655-100mg
- DUOLINK Proximity Ligation Assay - DUO92008
Troubleshooting
Safety warnings
1) Choosing the right and highly specific primary antibody is crucial.
2) Ensure the blocking reagent is stored properly. Prolonged storage at recommended 4°C, may reduce the effectiveness of the solution. Its inefficiency can be visualised by improper staining and many unusually faint PLA puncta.
3) Counterstaining with MAP2 after ligation and polymerisation results in smooth dendritic staining but may lead to the loss of PLA puncta during subsequent washing steps. To minimize this issue, MAP2 staining is performed together with the primary antibody incubation step, although this may slightly compromise the quality of MAP2 labelling.
Ethics statement
Experiments were conducted with the prior approval from IACUC.
Before start
Make sure to include appropriate negative controls (omission of one primary antibody). Prepare all the buffers including wash A and B in a sterile and filtered water. Kindly make a note of the pre-preparation before each step.
DAY 1 - Fixation
35m
PRE-PREP:
Prepare 4% Paraformaldehyde [PFA, EMS Cat. #1571]: Add 4ml of 10xPBS + 26ml of
milliQH2O, then add 10ml of 16% PFA.
Rapid fix the cells in 4% PFA at room temperature (RT) for 15-20 mins.
Note
Note: Perform it in the fume hood. I used 1ml pipette to remove the media from each well and promptly added approximate volume of 200 - 250ul of PFA to each well using transfer pipette. Washing is not required before fixing.
20m
Wash 3 times in 1xPBS, 5min/wash at room temperature.
Note
Note: First wash is critical and must not allow the PFA to dry in the well. Drying of PFA might cause background issues while imaging.
15m
Permeabilisation
25m
PRE-PREP:
Add 1ml of 1% of saponin [Stock solution stored at 4°C, Cat. Sigma #84510] to 9ml of 1xPBS [no calcium, magnesium] to a final concentration of 0.1% saponin.
Add 150µl of permeabilisation buffer 1xPBS-0.1% saponin and incubate for 10 mins at room temperature.
Note
Note: There are many types of detergent including nonidet, NP-40, Triton X-100, Tween-20, digitonin and various origins of saponin. I used saponin, it removes only cholesterol molecules in the membrane. Since my study focuses on receptor dynamics saponin was used.
10m
Wash 3 times in 1xPBS 5min/wash at room temperature.
15m
Blocking
1h
PRE-PREP:
Turn on the incubator to 37°C. To prepare a pre-humidity chamber, use a small laboratory tray that fits your 96 well plate. Place the pre-soaked blotting paper and add a few millilitres of water to retain the moisture of the paper. Cover the tray with aluminium foil and pre-warm at 37°C.
Add 60µl of DUOLINK blocking buffer to each well and incubate in the pre-humidity chamber for 60min at 37°C.
Note
Note: Test the blocking buffer before running the experiment. Refer attachment for more information.
1h
Primary Antibody
16h
PRE-PREP:
Primary antibodies in DUOLINK antibody diluent
Mouse Anti-PSD95 [Cat. MA1-046] - 1:500
Rabbit Anti-NR2B [Cat. 06-600] - 1:2000
Chicken Anti-MAP2 [Cat. PA1-10005] - 1:2000
DUOLINK antibody diluent - up to 40μl
After blocking no washes required.
Add 40μl of primary antibodies, incubate overnight 4°C in the pre-humidity chamber.
16h
DAY 2 - Secondary Antibody
1h 10m
PRE-PREP:
a) Bring the wash buffers A and B to the room temperature.
b) Turn on the incubator to 37°C.
c) Pre-warm the humidity chamber to 37°C and bring the plate to room temperature.
d) Secondary antibodies in DUOLINK antibody diluent
Donkey anti-rabbit PLA (+) probe [Cat. DUO92002] - 1:5
Donkey anti-mouse PLA (-) probe [Cat. DUO92004] - 1:5
Goat anti-chicken Alexa 488 [Cat. ab150173] - 1:1000
DUOLINK antibody diluent - up to 40μl
Wash the wells 2 times in buffer A for 5min/wash at RT.
Note
Note: During washes leave the pre-humidity chamber in the incubator.
10m
Add 40µl of secondary antibodies, incubate in the pre-humidity chamber for 1hr at 37°C.
1h
Ligation
30m
PRE-PREP:
Prepare the ligation mix based on the number of samples. For a 40μl reaction,
High purity or PCR grade water - 31μl
5x Ligation buffer [1:5] - 8μl
Ligase enzyme [1:40] - 1μl
Wash the wells 2 times in buffer A for 5min/wash at RT.
Note
Note: During washes leave the pre-humidity chamber in the incubator.
Add 40μl of ligation mixture, incubate in the pre-humidity chamber at 37°C for 30mins.
30m
Polymerisation (Rolling Circle Replication reaction)
1h 50m
PRE-PREP: Light sensitive reagents
DUOLINK polymerisation assay for 40ul reaction,
High purity or PCR grade water - 31.5μl
5x Polymerisation buffer [1:5] - 8μl
Polymerase enzyme [1:80] - 0.5μl
Wash the wells 2 times in buffer A for 5min/wash at RT.
Note
Note: During washes leave the pre-humidity chamber in the incubator.
10m
Add 40μl of amplification mixture, incubate in the pre-humidity chamber at 37°C for 100mins.
1h 40m
HOECHST - Nuclei staining and Imaging
46m
PRE-PREP:
Prepare required volume of HOECHST (1μg/ml, Cat. Thermo scientific #62249) in wash buffer B.
Wash once in buffer B for 10min at RT.
Note
Note: Pre-humidity chamber is not required from this step.
10m
Incubate with HOECHST for 15min at RT.
15m
Wash 2 times in buffer B for 10min at RT.
20m
Wash in 0.01x buffer B for 1min at RT.
1m
Add 150μl of 1xPBS and perform the imaging.
Note
Imaged using Opera Phenix High Content Screening equipped with 40x water immersion objective (Numerical Aperture 1.1). Pixel size = 0.32µm per pixel.
Protocol references
Citations
Verstraelen P, Garcia-Diaz Barriga G, Verschuuren M, Asselbergh B, Nuydens R, Larsen PH, Timmermans JP, De Vos WH. Systematic Quantification of Synapses in Primary Neuronal Culture.
https://doi.org/10.1016/j.isci.2020.101542Putra M, Rao NS, Gardner C, Liu G, Trommater J, Bunney M, Gage M, Bassuk AG, Hefti M, Lee G, Thippeswamy T. Enhanced Fyn-tau and NR2B-PSD95 interactions in epileptic foci in experimental models and human epilepsy.
https://doi.org/10.1093/braincomms/fcae327Verstraelen P, Van Dyck M, Verschuuren M, Kashikar ND, Nuydens R, Timmermans JP, De Vos WH. Image-Based Profiling of Synaptic Connectivity in Primary Neuronal Cell Culture.
https://doi.org/10.3389/fnins.2018.00389Acknowledgements
The author acknowledges the use of laboratory infrastructure and materials provided.

