Nov 19, 2025
Pushing boundaries: overcoming technical barriers in spatial RNA-seq analysis of developing wheat grains
- Farhad Masoomi-Aladizgeh1,2,
- Thomas M. Ashhurst3,4,
- James Fraser5,
- Samson Dowland3,6,
- Thomas H. Roberts1,2
- 1School of Life and Environmental Sciences, University of Sydney, Camperdown, NSW 2006, Australia;
- 2Sydney Institute of Agriculture, University of Sydney, Camperdown, NSW 2006, Australia;
- 3School of Medical Sciences, University of Sydney, Camperdown, NSW 2006, Australia;
- 4Sydney Cytometry Core Research Facility, University of Sydney and Centenary Institute, Camperdown, NSW 2006, Australia;
- 510X Genomics Pty Ltd, NSW 2000, Australia;
- 6Histology Facility, University of Sydney, Camperdown, NSW 2006, Australia
- Spatial Omics

Protocol Citation: Farhad Masoomi-Aladizgeh, Thomas M. Ashhurst, James Fraser, Samson Dowland, Thomas H. Roberts 2025. Pushing boundaries: overcoming technical barriers in spatial RNA-seq analysis of developing wheat grains. protocols.io https://dx.doi.org/10.17504/protocols.io.5qpvodq47g4o/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: September 23, 2025
Last Modified: November 19, 2025
Protocol Integer ID: 227964
Keywords: developing wheat grains spatial rna, wheat grains spatial rna, technical barriers in spatial rna, broader application of spatial rna, spatial rna, comprehensive transcriptome analysis at cellular resolution, rna, developing wheat grain, rnase inhibitor, spatial context of gene expression, tissue adherence during the preparation, analysis in immature wheat grain, spatial gene expression, comprehensive transcriptome analysis, immature wheat grain, wheat grain, seq technology in plant research, gene expression, intact tissue architecture, complex tissue structure
Abstract
Spatial RNA sequencing enables comprehensive transcriptome analysis at cellular resolution while preserving the spatial context of gene expression within the intact tissue architecture. However, its application to plant tissues presents unique challenges due to high cellular heterogeneity and complex tissue structure. The Visium HD 3’ protocol was used to analyse spatial gene expression in developing wheat grains. The protocol, optimised for mammalian tissues, was not inherently compatible with sections of developing wheat grains, because complete tissue detachment occurred during early processing stages. To address this issue, the protocol was extensively refined by introducing a 30% sucrose solution containing an RNase inhibitor, which markedly improved tissue sectioning and mounting. Further optimisation included initial fixation with cold methanol and isopropanol, reducing PBS concentration during post-fixation washes, post-enhancer washes and destaining steps, and lowering the temperature during preparation of the enhancer mix. These modifications improved tissue adherence during the preparation of fresh‑frozen sections and the quality of cross-sections, enabling the capture of poly(A) RNA and synthesis of high-quality cDNA suitable for library preparation and spatial RNA sequencing analysis in immature wheat grains. These findings should facilitate the broader application of spatial RNA-seq technology in plant research.
Materials
- Visium HD 3' Reagents - Kit A Small, PN-1000854
- Visium HD 3' Reagents - Kit B Small, PN-1000855
- Visium Slide Cassettes S3 6.5 mm, 2 pk, PN-1000847
- Visium HD Slide, 6.5 mm, 2 rxns, PN-1000670
- Visium CytAssist Reagent Accessory Kit, PN-1000499
- Visium CytAssist Alignment Aid, 6.5 mm, PN-1000866
- Glycerol, G5516
- Nuclease-free water, W4502
- Dulbecco’s Phosphate Buffered Saline (PBS), D1408
- Acetone, 179124
- Methanol, 34860
- Alcoholic Eosin, HT110116
- Gill II Hematoxylin, GHS232
- Dako Bluing Buffer, CS702
Troubleshooting
1. Fresh-frozen Tissue Preparation
Tissue collection
Immature wheat grains were harvested from primary or secondary florets of the middle spike 20 days after anthesis. The immature grains were immediately frozen in liquid nitrogen using an isopentane bath and stored at -80°C.
Sucrose infiltration
Sucrose treatment was applied after snap-freezing and before cryosectioning to improve the sectioning quality and adhesion of developing wheat grains at the milky stage (20 days after anthesis), which have high water content. These grains are highly hydrated, making them challenging to section and prone to detachment when preparing the tissue sections for the Visium HD 3’ workflow. Pre-cooled isopentane was used to minimise the formation of large ice crystals and to preserve tissue morphology during the collection of developing grains. The sucrose infiltration in this workflow served as an osmotic stabiliser to firm the tissue and enhance sectioning quality. The need for this step became evident only after processing immature grains for Visium HD 3’ as complete tissue detachment occurred during fixation and H&E staining when using the standard 10X Genomics workflow recommended for mammalian cells (10X Genomics, 2025a), highlighting the need for improved adhesion strategies for plant tissues. This reflects that while sucrose treatment may not be necessary for many plant tissues before snap freezing, it is critical for water-rich tissues, such as developing grains and should be considered in future protocols to prevent any potential issues in downstream analysis.
CRITICAL STEP 1: For sucrose infiltration, the frozen tissues were incubated in 200 µl of 30% sucrose in 1X PBS, containing 1 U/µl RNase inhibitor, for 2 hours at 4°C. This amount is sufficient to submerge two immature grains in 2-ml Eppendorf tubes.
Important notes:
1. Depending on the developmental stages of the tissue of interest, it is important to adjust the duration of sucrose infiltration. A longer sucrose infiltration period may result in drier, harder tissue sections that are more difficult to mount on slides. In contrast, a shorter infiltration period may not fully resolve the issues described above.
2. It is recommended that tissue sections be tested with different incubation periods without adding RNase inhibitor to the sucrose solution until the treatment for the specific tissue section is optimised.
3. After sucrose infiltration, the immature grains should be rinsed in cold 1X PBS to remove any remaining sucrose. Then, the tissues should be gently wiped with Kimwipes and transferred to new Eppendorf tubes.
4. Except for the time when the tissues are incubated in 30% sucrose in 1X PBS on ice, dry ice must be used to transfer the immature grains from and to a -80°C freezer.
Cryosectioning
The sucrose-infiltrated grains were embedded in OCT (Optimal Cutting Temperature) compound in the cryostat. Simply, a developing grain was placed on a drop of OCT to hold it, followed by the addition of more OCT to support the tissue during sectioning. The cryostat temperature was set to -21°C, while the specimen temperature ranged from -15°C to -20°C, depending on the sectioning quality. After obtaining appropriate tissue sections, a cold slide (SuperFrost Plus Adhesion Slides) that has been maintained in the cryostat for at least 5 minutes should be used to mount the section. To do so, the positively charged slides should be gently pressed down onto the knife stage to pick up the tissue section. There is no need to place a finger on the backside of the slide to allow the tissue section to adhere.
CRITICAL STEP 2: the specimen temperature should be increased up to -15°C if the tissue sections crack, but if the tissue sections curl, the specimen temperature should be decreased up to -20°C.
CRITICAL STEP 3: The morphology of tissues affects the outcome of sucrose infiltration and sectioning. In tissues like developing grains, which exhibit a tapered apex and a rounded midsection, the distribution of infiltrated sucrose can differ significantly, with sucrose penetrating more quickly into the narrowed apex than into the broader midsection, which contains a larger mass, within a given time. As a result, sections cut from the narrowed apex may be dry and brittle due to excessive sucrose infiltration, making them prone to tearing or curling during mounting. To address this, two strategies can be considered: 1) optimising the duration of sucrose exposure for specific regions of interest through time-course infiltration experiments, or 2) preparing slightly thicker tissue sections to reduce dehydration artefacts, provided it does not interfere with the Visium HD 3' platform. This protocol was successfully tested on tissue sections with thicknesses of 10 and 15 µm. The latter was used throughout the workflow to account for variations in immature grain size due to heat and the resulting differences in sucrose infiltration.
Important notes:
1. Before working on the actual specimen, it is recommended to prepare a dummy specimen to get comfortable with sectioning. This provides a chance to fine-tune the cryostat conditions, especially the temperature needed to produce high-quality sections, without risking the actual specimen.
2. If the official aligner tool from 10X Genomics is not available, it is recommended to draw a 6.5 mm² reference area (or 12.5 mm², depending on the capture area) on a microscopy slide to ensure tissue sections are correctly positioned within the Visium HD 3' capture area. This helps capture RNA from the entire tissue section, especially when multiple tissue sections are placed in the capture area.
Slide Preparation, Fixation and H&E Staining
Fixation is the most crucial step in preparing tissue sections for spatial RNA-seq analysis. Initial attempts using the 10X Genomics Visium HD 3' protocol, optimised for mammalian tissues, resulted in the complete detachment of developing grain sections from slides. This was observed when the developing grains were immersed in cold methanol:acetone (1:1) for 1 minute, followed by post-fixation washes, the application of enhancer mix, post-enhancer washes, and H&E staining. To address this, a series of trials were conducted to optimise the fixation and slide preparation protocol for immature wheat grains.
Table 1. Summary of experimental trials conducted to modify slide preparation, fixation and H&E staining for Visium HD 3' workflow.
Final optimised protocol for slide preparation, fixation and H&E staining:
- Sucrose infiltration: the developing grains were incubated in 200 µl of 30% sucrose in 1X PBS with 1 U/µl RNase inhibitor for 2 hours at 4°C.
- Thermal cycler step: the tissue sections were incubated at 37°C for up to 2 minutes to promote dehydration (10X Genomics, 2025b).
- Fixation: the tissue sections were immersed in cold 100% methanol for 30 minutes at -30°C, followed by incubation in 0.5 ml cold 100% isopropanol for 1 minute and air-dry for 5 minutes (10X Genomics, 2025b).
- Post-fixation and enhancer wash: the tissue sections were washed with 0.1X PBS (not 1X PBS).
- Buffer optimisation: the enhancer mix was prepared using pre-warmed 0.1X PBS at 30°C, rather than 37°C, and maintained at this temperature until use.
Important notes:
1. Preliminary trials without Visium HD 3' reagents might be necessary to optimise the workflow to achieve high-quality tissue sections before full protocol implementation.
2. When 0.1X PBS was used at room temperature, it produced similar results as when 0.1X PBS was warmed to 30°c. However, because the enhancer mix might precipitate at room temperature, 0.1X PBS was preheated to 30°C to prepare the enhancer mix.
Destaining
To maintain consistency, 0.1X PBS was used instead of 1X PBS during the destaining process, too.
Workflow for Fresh-frozen Tissue Preparation of Wheat Immature Grain for Visium HD 3'
The workflow for the preparation of fresh-frozen tissue includes tissue collection and sucrose infiltration, crosectioning, fixation and H&E staining, and destaining.
Figure 1. The workflow for preparing tissue sections for Visium HD 3' runs. After collecting developing wheat grains (1a), the samples were incubated in liquid nitrogen with isopentane (2a). Due to tissue detachment observed during the Visium HD 3’ workflow, sucrose infiltration was used as an osmotic stabiliser to firm the tissue and improve sectioning quality (3a). It is recommended to consider sucrose treatment before freezing water-rich plant tissues in isopentane, if feasible. The treated developing grains were then embedded in OCT compound (1b), followed by optimisation of sectioning quality in the cryostat (2b). If the tissue was not treated with sucrose before freezing, this is the stage at which the treatment can be applied to the developing grains. Additionally, the thickness of the tissue sections can be adjusted, provided it aligns with the workflow. The tissue sections prepared in the cryostat underwent dehydration for 1-2 minutes (1c), followed by fixation, which is a critical stage in this workflow, especially when working on plant tissues and can be
extended if necessary (2c). Then, the fixed tissue was treated as instructed in the Visium HD 3’ protocol with the following modifications: 0.1X PBS was used instead of 1X PBS during the staining and destaining processes, with the temperature decreased to 30°C instead of 37°C.
Expected Results
For spatial gene expression analysis, a custom reference genome for wheat was created from IWGSC RefSeq v2.1 and its corresponding annotation (Zhu et al., 2021) using the mkref function in spaceranger v4.0.1. The fastq reads were then mapped to the custom reference genome using the count pipeline in spacersanger v4.0.1. The output file was then imported into Loupe Browser ver 9.0.0 for analysis.
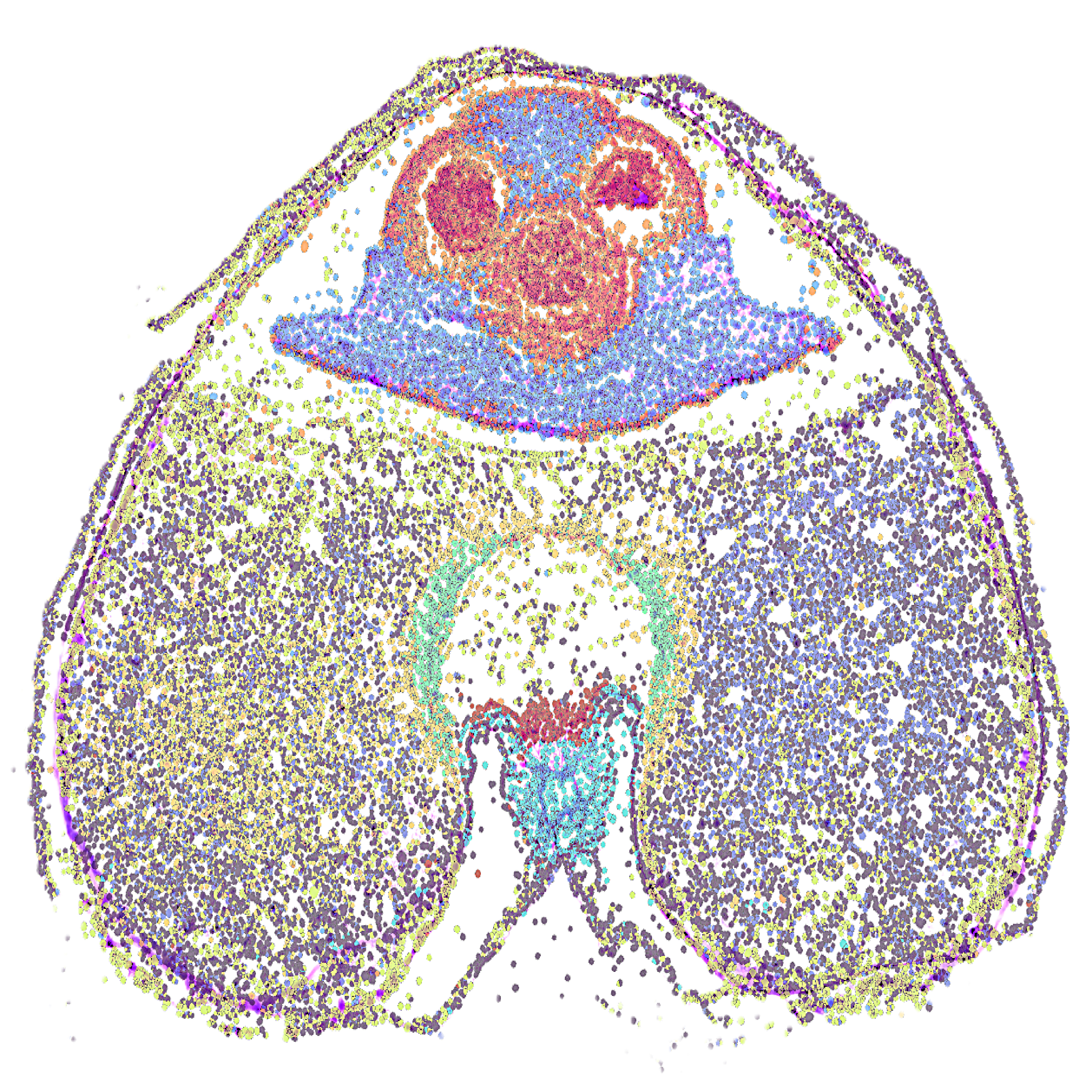
Figure 2. Visium HD 3' gene expression analysis using tissue cross-sections prepared from developing wheat grains. (a) tissue detachment observed after slide preparation, fixation, and H&E staining, prior to method optimisation; (b) high-quality cross-section after slide preparation, fixation, and H&E staining, following method optimisation; (c) electropherogram displaying the average size and concentration of the pooled library after cDNA synthesis from RNA captured from the tissue; (d) spatial gene expression of tissue cross-section at bin-level resolution and cell segmentation. The scale bars in figures a and b indicate 400 and 200 µm, respectively.
Protocol references
10X Genomics. 2025a. Visium HD 3’ Fresh Frozen
Tissue Preparation Handbook (CG000804, Rev B). Retrieved from https://www.10xgenomics.com/support/spatial-gene-expression-hd-three-prime/documentation/steps/tissue-prep-for-fresh-frozen/visium-hd-3-prime-fresh-frozen-tissue-preparation-handbook
10X Genomics. 2025b. Are plant samples compatible with the Visium HD 3’ assay? Retrieved from https://kb.10xgenomics.com/hc/en-us/articles/38948485416461-Are-plant-samples-compatible-with-the-Visium-HD-3-assay
Zhu, T. et al., 2021. Optical maps refine the bread wheat Triticum aestivum cv. Chinese Spring genome assembly. Plant J, 107(1): 303-314.
