Apr 07, 2025
Version 2
Processing human brain tissue for population-scale SQK-LSK114 Oxford Nanopore long-read DNA sequencing SOP V.2

- Jackson Mingle1,
- Kimberly Paquette1,
- Breeana Baker1,
- Laksh Malik1,
- Maysa Abdelhalim1,
- Pilar Alvarez Jerez1,
- Kimberley J Billingsley1,
- on behalf of the CARD Long-read Team1
- 1Center for Alzheimer's and Related Dementias, National Institute on Aging, Bethesda, Maryland, USA
- NIH Center for Alzheimer's and Related Dementias

Protocol Citation: Jackson Mingle, Kimberly Paquette, Breeana Baker, Laksh Malik, Maysa Abdelhalim, Pilar Alvarez Jerez, Kimberley J Billingsley, on behalf of the CARD Long-read Team 2025. Processing human brain tissue for population-scale SQK-LSK114 Oxford Nanopore long-read DNA sequencing SOP . protocols.io https://dx.doi.org/10.17504/protocols.io.kxygx3zzog8j/v2Version created by Jackson Mingle
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: In development
We are still developing and optimizing this protocol
Created: March 04, 2025
Last Modified: April 07, 2025
Protocol Integer ID: 123802
Keywords: Long-read sequencing, Oxford Nanopore sequencing, Human tissue disruption, High molecular weight DNA extraction, DNA extraction, Brain tissue extraction, DNA size selection, DNA shearing, read sequencing of human brain tissue, lsk114 oxford nanopore, genome sequencing, nanopore team, read sequencing, sequencing data, genome, human brain tissue for population, nih center for alzheimer, read dna, sequencing, sequencing sop, human brain tissue, alzheimer disease, processing human brain tissue, patients with alzheimer, alzheimer, lewy body dementia, related dementia, dna, frontotemporal dementia, dementia
Disclaimer
Still in development.
Abstract
At the NIH Center for Alzheimer's and Related Dementias (CARD) (https://card.nih.gov/research-programs/long-read-sequencing), we will generate long-read sequencing data from thousands of patients with Alzheimer's disease, frontotemporal dementia, and Lewy body dementia, as well as healthy subjects. With this research, we will build a public resource consisting of long-read genome sequencing data from a large number of confirmed patients with Alzheimer's disease and related dementias and healthy individuals. To generate this large-scale nanopore sequencing data, we have developed a protocol for processing and long-read sequencing of human brain tissue, targeting an N50 of ~25-30 kb and ~30X coverage.
Acknowledgements:
We would like to thank the Nanopore team (Jade Bartolo, Olivor Holman, Androo Markham, and Jessica Anderson), PacBio team (Jeffrey Burke, Michelle Kim, Duncan Kilburn, and Kelvin Liu), and the whole CARD long-read team. Workflow graphic by Paige Jarreau.
†Correspondence to: Kimberley Billingsley [email protected]

Figure 1. Overview of the HMW DNA extraction and ONT sequencing protocol
Materials
Materials:
| A | B | |
| Material | Vendor (Part Number) | |
| Sterile weigh boats | Any major lab supplier (MLS) | |
| Single edge razor blades | Any MLS | |
| Laboratory spatulas | Any MLS | |
| Tweezers | Any MLS | |
| Plastic wrap | Any MLS | |
| 2 mL Protein LoBind tubes | Eppendorf (022431102) | |
| Ethanol (96-100%) | Any MLS | |
| RNaseZap | Thermo Fisher Scientific (AM9780) | |
| Nanobind HT CBB kit | PacBio (102-301-900) | |
| Nanobind PanDNA kit | PacBio (103-260-000) | |
| Stainless Steel Beads, 5 mm | Qiagen (69989) | |
| Wide-bore P1000 tips | Thermo Fisher Scientific (2079G) | |
| Tris-EDTA, 1X Solution pH 8.0 | Fisher Scientific (BP2473) | |
| Wide-bore P200 tips | Thermo Fisher Scientific (2069G) | |
| KingFisher 96 deep-well plates, barcoded | Thermo Fisher Scientific (95040450B) | |
| KingFisher 96 tip comb, barcoded | Thermo Fisher Scientific (97002534B) | |
| Isopropanol (100%) | Any MLS | |
| 1.5 mL DNA LoBind tubes | Eppendorf (022431021) | |
| 1 mL Luer-Lok Syringe | BD (309628) | |
| 26G 1.5" Blunt Needle, Capped | SAI Infusion Technologies (B26-150) | |
| Qubit 1X dsDNA BR assay kit | Thermo Fisher Scientific (Q33266) | |
| Qubit Flex Assay Tube Strips | Thermo Fisher Scientific (Q33252) | |
| Femto Pulse gDNA 165kb analysis kit | Agilent Technologies, Inc. (FP-1002-0275) | |
| Megaruptor 3 DNAFluid+ kit | Diagenode (07020001) | |
| TE Buffer | Thermo Fisher Scientific (12090) | |
| 2X Loading Solution | Sage Science (190192) | |
| High Pass Plus Cassette | Sage Science (BPLUS10) | |
| Ligation Sequencing Kit XL V14 | Oxford Nanopore Technologies (SQK-LSK114-XL) | |
| NEBNext Companion Module for Oxford Nanopore Technologies Ligation Sequencing | New England Biolabs (E7180L) | |
| Nuclease-free water | Any MLS | |
| 0.2 mL thin-walled PCR tubes | Thermo Fisher Scientific (AB-0620B) | |
| AMPure XP reagent | Beckman Coulter (A63882) | |
| Qubit 1X dsDNA HS assay kit | Thermo Fisher Scientific (Q33231) | |
| PromethION flow cell | Oxford Nanopore Technologies (FLO-PRO114M) | |
| Flow Cell Wash Kit | Oxford Nanopore Technologies (EXP-WSH004-XL) |
Table 1. Required materials
Equipment:
| A | B | |
| Equipment | Vendor (Part Number) | |
| 1.5/2 mL microcentrifuge tube cooling block | Any major lab supplier (MLS) | |
| Analytical balance | Any MLS | |
| TissueLyser Single-Bead Dispenser, 5 mm | Qiagen (69965) | |
| TissueLyser III | Qiagen (9003240) | |
| TissueLyser Adapter Set 2 x 24 | Qiagen (69982) | |
| DynaMag-2 Magnet | Thermo Fisher Scientific (12321D) | |
| Microcentrifuge | Eppendorf (5404000413) | |
| ThermoMixer | Eppendorf (5382000023) | |
| KingFisher Apex system | Thermo Fisher Scientific (5400940) | |
| KingFisher Apex 96 DW head | Thermo Fisher Scientific (24079930) | |
| KingFisher Apex 96 DW heating block | Thermo Fisher Scientific (24075930) | |
| Qubit Flex fluorometer | Thermo Fisher Scientific (Q33327) | |
| Vortex mixer | Any MLS | |
| Minicentrifuge | Any MLS | |
| Femto Pulse system | Agilent Technologies, Inc (M5330AA) | |
| Megaruptor 3 system | Diagenode (B06010003) | |
| BluePippin Size Selection System | Sage Science (BLU0001) | |
| Thermal cycler | Bio-Rad (1851197) | |
| Platform rocker | Any MLS | |
| PromethION 24 or 48 sequencing unit | Oxford Nanopore Technologies (PRO-SEQ024 or PRO-SEQ-048) |
Table 2. Required equipment
Materials for NGS STAR:
| A | B | |
| Material | Vendor (Part Number) | |
| 2 mL tubes | Sarstedt (72.693.005) | |
| PCR ComfortLid | Hamilton (814300) | |
| MIDI plate | Thermo Fisher Scientific (AB-0859) | |
| HSP plate | Bio-Rad (HSP9601) | |
| 50 µL tips | Hamilton (235979) | |
| 300 µL tips | Hamilton (235903) | |
| 1000 µL tips | Hamilton (235940) | |
| 60 mL reservoir | Hamilton (56694-01) | |
| 20 mL reservoir | Roche (3004058001) |
Table 3. Additional materials required for automated library preparation on Hamilton NGS STAR
Equipment for NGS STAR:
| A | B | |
| Equipment | Vendor (Part Number) | |
| NGS STAR | Hamilton (STAR) | |
| Magnetic Stand-96 | Thermo Fisher Scientific (AM10027) |
Table 4. Additional equipment required for automated library preparation on Hamilton NGS STAR
Troubleshooting
Part 1: Human brain tissue disruption with Qiagen TissueLyser III
Place the following supplies on dry ice and allow to chill for 00:10:00 prior to use:
- Sterile weigh boats
- Razor blades
- Spatulas
- Tweezers
- Empty 2 mL Eppendorf Protein LoBind tubes - labeled with sample ID
- Cooling block
Note: Reusable metal tools (spatulas, tweezers) must be cleaned with 70% ethanol, RNaseZap, and distilled water prior to use. Apply a layer of plastic wrap to the dry ice and place metal tools on the plastic wrap to chill them prior to use.
Obtain tissue samples from -80 °C freezer and place them on dry ice.
Don all necessary protective equipment prior to setting up the biosafety cabinet:
- Disposable lab coat
- Gloves (2 pairs)
- Face shield or mask
Wipe down biosafety cabinet with 70% ethanol, RNaseZap, and distilled water. Ensure scale is set up properly and level. Place dry ice buckets with supplies and samples in biosafety cabinet.
Remove cooling block from dry ice. Place sterile weigh boat onto cooling block.
Weigh labeled empty 2 mL Eppendorf Protein LoBind tube and tare the scale, ensuring the tube is centered on the scale.
Note: If there's ice surrounding the tube, wipe it off before taring. When weighing the sample, work quickly. As the dry ice surrounding the tube evaporates, the weight may fluctuate.
Pour frozen brain tissue onto sterile weigh boat on cooling block. With the razor blade in one hand, and with the other hand shielding the weigh boat to prevent frozen tissue from flying onto the surface, cut the brain tissue firmly.
Pick up the cut piece of brain tissue with the razor blade and transfer to the empty labeled tube. Weigh the tube immediately and record the weight. If necessary, continue to cut the brain tissue to reach the target weight. Return to dry ice immediately.
Note: For most brain regions, 35-45 mg is required for DNA extraction for long-read sequencing. Input requirements will vary per brain region (e.g. 8-10 mg is recommended for DNA extraction from cerebellum samples). DNA recovery from brain tissue extractions will vary based on amount of gray matter and white matter per sample.
Note: It is crucial to prevent brain tissue from thawing during the cutting process. Once the brain tissue is removed from dry ice, cut and weigh the tissue quickly and return to dry ice immediately. Thawing of brain tissue during cutting could lead to DNA extractions with reduced quality.
Note: For flaky or powdery samples, clean metal tweezers or spatulas can be used to transfer tissue to the tube.
Return any unused brain tissue on the weigh boat to a tube for permanent storage. Return to dry ice immediately.
Dispose of the razor blade in the sharps container. Dispose of the weigh boat, top layer of gloves, and empty tube formerly containing brain tissue (if applicable) in Medical Pathological Waste (MPW) box. Return the cooling block to dry ice.
Repeat steps 4 - 10 for additional brain tissue samples. Wipe down biosafety cabinet with 70% ethanol, RNaseZap, and distilled water, and put on a new top layer of gloves between each sample to prevent cross-contamination. Keep all samples on dry ice when not in use.
When finished, wipe down biosafety cabinet with 70% ethanol, RNaseZap, and distilled water. Clean reusable metal tools with 10% bleach and water.
Complete the following preparation steps:
- Place cooling block for 2 mL tubes on dry ice for 00:10:00
- Place TissueLyser adapters in -20 °C freezer
- Place Buffer CT on ice
- Chill centrifuge to 4 °C
- Set ThermoMixer to 55 °C
- Turn on the UV light in the KingFisher Apex system for 01:00:00
- Wipe down all surfaces, pipettes, and magnetic racks with 70% ethanol and RNaseZap
Obtain tissue samples and place in pre-chilled cooling block.
Using the Qiagen bead dispenser, add one 5mm stainless steel bead to each sample.
Note: To prevent brain tissue from flying out of the tube, dispense bead into tube cap, then close the tube.
Add 750 µL cold Buffer CT. Ensure all brain tissue chunks are submerged in Buffer CT using the pipette tip. Keep on wet ice during processing of additional samples.
Remove the TissueLyser adapters from -20 °C freezer. Load samples into the middle rows of the TissueLyser adapters. Ensure TissueLyser adapters are balanced.
Note: Samples loaded into the top and bottom rows of the TissueLyser adapter tend to be unequally homogenized.
Place the lid onto the TissueLyser adapters and load onto TissueLyser. Do not over-tighten the adapters in the TissueLyser.
Run the TissueLyser with a frequency of 25 Hz for 00:00:40 .
Remove samples from the TissueLyser adapters. Place on wet ice.
Remove the stainless steel bead by sliding the sample tube vertically along the magnetic rack with the cap open. The bead will stick to the magnet.
Note: Wipe down magnetic rack with 70% ethanol and RNaseZap before and after removing stainless steel beads from sample tubes.
Centrifuge at 6,000 x g for 00:05:00 at 4 °C to pellet homogenate.
Note: Insert samples into centrifuge with the hinge facing out.
Remove and discard supernatant, taking care to avoid disturbing the pellet.
Note: Pellet may not be visible, so avoid touching the bottom of the tube with the pipette tip.
Add 1 mL cold Buffer CT. Pipette-mix ~10x with a wide-bore P1000 pipette tip to resuspend the pellet. If necessary, continue pipetting until the pellet is fully resuspended.
Centrifuge at 6,000 x g for 00:05:00 at 4 °C to pellet homogenate.
Remove and discard supernatant, taking care to avoid disturbing the pellet.
Note: Pellet may not be visible, so avoid touching the bottom of the tube with the pipette tip.
Add 20 µL Proteinase K.
Add 50 µL TE pH 8.
Add 60 µL Buffer CS.
Add 100 µL Buffer CLE3.
Pipette-mix ~15x with a wide-bore P200 pipette tip.
Note: Often, the pellet does not fully resuspend following pipette-mixing. Incubation in the next step should ensure adequate sample lysis.
Incubate for 01:00:00 at 55 °C and 900 rpm in Thermomixer. Once complete, spin down.
Add 20 µL RNase A and pipette-mix 5x with a wide-bore P200 pipette.
Incubate on ThermoMixer for 00:30:00 at 55 °C and 900 rpm. Once complete, spin down.
Note: During this time, begin preparing the plates for the KingFisher Apex as described in step 36.
Add 50 µL Buffer SB and vortex for 00:00:10 at maximum speed. Spin down.
Part 2: Extracting HMW DNA using the Nanobind HT CBB kit and Nanobind PanDNA kit for human brain tissue on the KingFisher Apex system
Prepare the KingFisher Apex 96 deep-well plates:
- Plate 1: Lysis/Binding Plate: 50 µL Buffer BL3
- Plate 2: Nanobind Storage Plate: One 3 mm Nanobind disk per well
- Plate 3: Wash Plate 1: 600 µL Buffer CW1 per well
- Plate 4: Wash Plate 2: 600 µL Buffer CW1 per well
- Plate 5: Wash Plate 3: 600 µL Buffer CW2 per well
- Plate 6: Wash Plate 4: 600 µL Buffer CW2 per well
- Plate 7: Elution Plate : 100 µL Buffer EB per well
- Plate 8: Tip Plate: KingFisher Apex 96 deep-well tip comb
Note: Nanobind disks do not need to be perfectly centered in the wells, but ensure they are at the bottom of the wells and not stuck to the side walls.
Note: Buffer CW1 and CW2 are supplied as concentrates. This kit uses CW1 with a 60% final ethanol concentration and CW2 with a 60% final ethanol concentration. Before using, add the appropriate amount of ethanol (96–100%) to Buffer CW1 and Buffer CW2 as indicated on the bottles.
Transfer samples from step 35 into the Lysis/Binding Plate, pre-filled with 50 µL Buffer BL3. Sample volume should be ~250 µL .
Ensure the KingFisher Apex instrument is set up with the 96 deep-well magnetic head and the 96 deep-well heating block.
Select the Brain_Tissue_Nanobind_HT_Apex script on the KingFisher Apex instrument. Insert plates into the KingFisher Apex instrument as indicated on the display and press 'Next' after every plate to confirm position. The protocol will start when the final plate is loaded and the 'Next' button is pressed.
When prompted by the instrument (~12 minutes after start), remove the Lysis/Binding Plate from the instrument and add 300 µL of isopropanol to each well. Re-insert the plate and press 'Next' to resume the protocol.
Note: Add isopropanol gently against the side of the well into the Lysis/Binding solution. Adding isopropanol directly to the Lysis/Binding solution may affect extraction purity.
When prompted by the instrument, remove the Elution Plate from the instrument. Transfer eluate from each well to a new 1.5 mL Eppendorf DNA LoBind tube.
Note: Residual sample volume may be present in the tip comb plate. Transfer residual eluate to the 1.5 mL Eppendorf DNA LoBind tube.
Let eluate rest overnight at room temperature to allow DNA to solubilize.
For any samples with remaining tissue debris, centrifuge at 10,000 x g for 00:05:00 at room temperature. Transfer the supernatant to a new 1.5 mL Eppendorf DNA LoBind tube.
Following overnight rest, hand-shear 10x with a 1 mL Luer-Lock syringe and a 1.5" needle. Ensure no sample is left behind in the syringe by removing the plunger.
Quantify by taking a single measurement on the Qubit Flex Fluorometer with the Qubit 1x dsDNA BR Assay. Use 1 µL of DNA per measurement.
Note: Triplicate measurements (top, middle, and bottom) are advised for new cohorts or brain regions. Additional hand-shearing with a syringe may be necessary for samples with exceptionally variable measurements.
Optional: Quantify by taking a single measurement on the NanoDrop 8000 spectrophotometer. Use 2 µL of DNA per measurement.
Note: Measurement on NanoDrop 8000 is advised for new cohorts or brain regions. Additional hand-shearing with a syringe may be necessary for samples with exceptionally variable measurements.
Optional: Size on the Agilent Femto Pulse System with the Genomic DNA 165 kb kit. The expected size range for samples post-extraction is >60kb.

Figure 2. Femto Pulse trace post-extraction
Note: Sizing on Agilent Femto Pulse System is advised for new cohorts or brain regions. Additional hand-shearing with a syringe may be necessary for samples with exceptionally variable measurements.
Store at 4 °C until ready for shearing.
Part 3: Shearing with Diagenode Megaruptor 3 DNAFluid+ Kit
Prepare DNA in 100 µL Buffer EB, at a concentration of up to 150 ng/µL in a Megaruptor 3 shearing tube.
Note: Shearing conditions are optimized for sequencing N50s ~25-35 kb. Shearing with Megaruptor 3 is concentration- and volume-dependent. Samples with a concentration > 150 ng/µL may be susceptible to under-shearing. Do not shear at a concentration > 150 ng/µL.
Note: For samples with concentrations > 150 ng/µL, prepare DNA in > 100 µL, and ensure sample concentration is under 150 ng/µL. Ensure the volume is input on MegaRuptor 3 accurately in step 52.
Note: For samples with low yields, shear the entire sample in 100 µL, then pool with additional samples after shearing to reach the required input for size selection, if desired.
Remove the Megaruptor 3 DNA Fluid+ shearing syringe from the package. Tighten the assembly and ensure the plunger is completely compressed. Attach the syringe to the shearing tube and ensure the syringe is snug on the cap of the tube.
Load samples and syringes onto the Megaruptor 3. If running an odd number of samples, samples can be balanced with an empty corresponding tube.
Run samples on the Megaruptor 3 at speed 45. Enter the appropriate sample volume (100 µL) and sample concentration.
Note: Sample volume entered into MegaRuptor 3 protocol must accurately reflect the sample volume. If running samples with different volumes (due to high concentrations), samples must be run on separate MegaRuptor 3 protocols.
Note: Ensure samples of different volumes and concentrations are run on separate MegaRuptor 3 protocols. For different volumes, keep samples within +/- 10 µL of each other. For different concentrations, keep samples within +/- 15 ng/µL of each other.
Note: Shearing speed may need to be optimized for new cohorts or brain regions to achieve targeted size.
Repeat the previous step for a total of 2 runs on the Megaruptor 3 at speed 45.
Remove the samples from the Megaruptor 3. Remove the syringe from the tube. Make sure the plunger is fully depressed to avoid losing sample volume. Disassemble the needle and remove residual sample from the needle and syringe to avoid losing sample volume.
Quantify by taking a single measurement on the Qubit Flex Fluorometer with the Qubit 1x dsDNA BR Assay. Use 1 µL of DNA per measurement.
Size on the Agilent Femto Pulse System with the Genomic DNA 165 kb kit. The expected size range for samples post-shearing is ~25-40 kb.

Figure 3. Femto Pulse trace post-shearing
Store at 4 °C until ready for size selection. For long-term storage, store at -80 °C .
Part 4: Size Selection with Sage Science BluePippin
Prepare up to 10.5 µg sheared DNA in 77 µL TE buffer or Buffer EB.
Note: If samples are low volume, low concentration, or limited due to lack of starting material, samples can be prepared in 32 µL or 62 µL TE buffer.
Remove 2X Loading Solution from 4 °C fridge and allow it to reach room temperature.
Note: If preparing samples at a lower volume of 32 µL or 62 µL , standard (1X) Loading Solution should be used and allowed to come to room temperature.
Create a new protocol using the BluePippin software and select the cassette type:
- Click the "Protocol Editor" tab
- Click the "Cassette Type" folder icon
- Click "+" icon next to "0.75% Agarose Dye-Free"
- Select "10kb High Pass Plus Marker U1"
- Click "SELECT"
Note: The standard protocol used is "10kb High Pass Plus Marker U1". If samples require more aggressive size selection following analysis of post-shearing Femto Pulse traces, "15kb High Pass Plus Marker U1" or "20kb High Pass Plus Marker U1" protocols can be run instead.
Enter the sample IDs into the Sample ID fields. Enter Marker for the Marker well in Lane 4.
Note: Lane 4 is the preferred lane for the Marker, but other lanes can be used.
Make sure the check box for "End Run when Elution is Completed" is selected.
In the "Range" column, ensure the rectangle boxes next to sample lanes are light gray, or selected. Ensure the rectangle box in the Marker lane is dark gray, or not selected.

Figure 4. Range mode selection
Enter the lane number to which the DNA marker will be loaded into the "Reference Lane" field, and click the "APPLY REFERENCE TO ALL LANES" button.
Enter the "BP Start" value of 10000 for all sample wells.
Note: The "BP End" and "BP Target" values are automatically populated.
Note: If using the "15kb High Pass Plus Marker U1" or "20kb High Pass Plus Marker U1" protocols, "BP Start" values should be entered as 15000 and 20000, respectively.
Click "Save As" to name and save the run.
Remove the gel cassette from the foil packaging and inspect the levels of buffer in all buffer reservoirs.
Note: Reservoirs should be nearly full. If the buffer level in any reservoir appears lower, or less than 50% full, fill with spare electrophoresis buffer.

Figure 5. Buffer reservoir volume inspection
Inspect for bubbles due to delamination of agarose from the bottom of the cassette in the region used for optical detection of DNA. To inspect, turn the cassette upside down and gently tilt the cassette under a light source. If a bubble is detected, do not use the affected lane to run the DNA reference marker. Affected lanes can be used for size selection of sample DNA.
Note: Sometimes, flat bubbles can occur between the top surface of the gel column and the plastic top of the channel. This will not affect run quality or optical detection and can be used for markers or samples without any adverse effects. If it is difficult to tell if a bubble will be problematic, it may be better to not use the affected lane for the marker. If a gel piece is detached, do not use that particular lane at all. Sage Science may be able to provide a replacement cassette.

Figure 6. High Pass Plus cassette inspection
Dislodge air bubbles from behind the elution wells. Tilt the cassette sample well side down, with the lower buffer chambers facing up, and tap gently to release the bubbles.
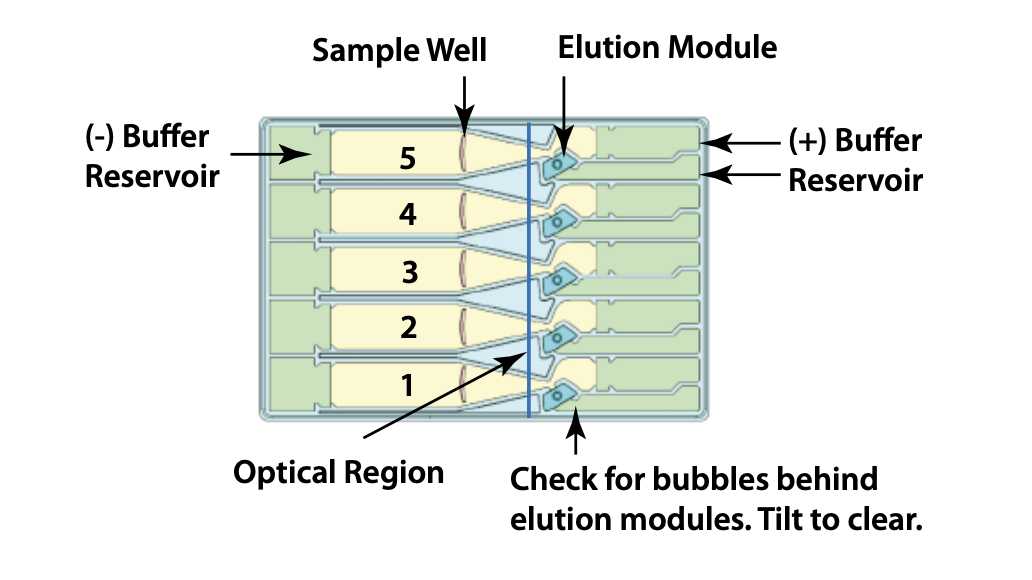
Figure 7. High Pass Plus cassette diagram
Place the cassette into the optical nest. Keep the cassette slightly tilted down so that the air bubbles do not return to the area behind the elution modules. Make sure the cassette is fully seated into the bottom of the nest to ensure proper optical alignment.
Remove the white tabbed adhesive strips from the cassette. Place one hand on the cassette, and hold it firmly in the nest. Grab the white tabs and pull the strips firmly and slowly toward the front of the instrument until they are removed.
If necessary, fill buffer reservoirs with spare electrophoresis buffer if any reservoir appears low or less than 50% full. For any buffer reservoirs that appear full, remove 750 µL buffer to reduce leakage at the end of the run.
Completely remove all buffer from each elution module using a P100 pipette tip. Move the pipette tip slowly to avoid piercing the gel at the corner of the elution modules.
Add 80 µL of fresh electrophoresis buffer to each elution module. Change pipette tips between elution modules. When refilling, dispense the fresh buffer slowly, raising the pipette tip slowly from the bottom of the elution module to avoid trapping air bubbles in the module.
Seal the elution modules with an adhesive tape strip. Place the tape over the elution wells and rub firmly to seal the elution ports using a straight edge. The port should be tightly sealed without any wrinkles around the edges of the port. If necessary, use two overlapping tape strips.
Add 11 µL 2X Loading Solution to the prepared DNA samples. Pipette-mix or flick mix, and spin down.
Note: The 2X Loading Solution is very viscous, so care should be taken to aspirate and dispense fully.
Note: If preparing samples at 32 µL or 62 µL DNA, 10 µL or 20 µL Standard (1X) Loading Solution should be added, respectively.
Gently remove the cassette from the optical nest.
Calibrate the optical sensors:
- Click the "Main" tab
- Place the calibration fixture onto the optical nest — the dark side of the fixture must be down, and completely cover all LED detectors
- Close the lid of the BluePippin instrument
- Press the "CALIBRATE" button; a calibration sub-window will launch
- Check that the calibration target box ("Target I pH, mA") at the top of the window reads "0.60"; if not, manually change the target value to "0.60"
- Press the "CALIBRATE" button within the calibration window
- If the status window reads "Calibration OK", press "EXIT" to return to the main screen
Note: Calibration should be performed no more than 30 minutes before the start of a run. If more than 30 minutes elapses between calibration and the start of the run, a new calibration should be performed.

Figure 8. Calibration fixture placement
Place the cassette into the optical nest. Keep the cassette slightly tilted down so that the air bubbles do not return to the area behind the elution modules. Make sure the cassette is fully seated into the bottom of the nest to ensure proper optical alignment.
Close the lid and perform the Continuity Test:
- Press the "Test" button — the test sub-window will open, and the test routine will automatically measure the current in each separation and elution channel
- A successful Continuity Test will return a "PASS" indication
- The cassette temperature must be above 17 °C (62 °F); if a lane fails due to low elution current (the affected lane will be highlighted with an orange color), refilling the elution module will usually resolve the problem
- Press "Return" to close the Continuity Test sub-window
Remove 85 µL of buffer from sample wells and load 85 µL of sample into the sample well. Take care not to pierce the gel with the pipette tip.
Note: When removing buffer, it is useful to immerse the pipette tip just below the surface of the buffer and follow the liquid level down with the tip as the buffer is removed. When buffer removal is completed, there will be ~70 µL of buffer left in the well. When adding sample, place the tip of the pipette just below the surface of the buffer, and slowly eject the sample. Don't be concerned if the sample well slightly overfills. The density of the sample will allow it to sink before it can flow out of the well.
Note: Cassettes may be reused if 2-3 lanes are used (1-2 samples). Reseal the cassette with the adhesive tape and store at room temperature. However, one remaining lane must again be loaded with the DNA marker.
Note: If preparing samples at 32 µL or 62 µL DNA, remove 40 µL or 80 µL of buffer from sample wells and load 40 µL or 80 µL of sample into the sample well, respectively.
Remove DNA marker from 4 °C fridge. Do not keep out any longer than needed. Return to 4 °C fridge promptly after use.
Remove 40 µL of buffer from the marker well and load 40 µL of marker into the marker well. Take care not to pierce the gel with the pipette tip.
Note: When removing buffer, it is useful to immerse the pipette tip just below the surface of the buffer and follow the liquid level down with the tip as the buffer is removed. When buffer removal is completed, there will be ~70 µL of buffer left in the well. When adding sample, place the tip of the pipette just below the surface of the buffer, and slowly eject the sample. Don't be concerned if the sample well slightly overfills. The density of the sample will allow it to sink before it can flow out of the well.
Close the lid. Ensure the proper protocol is loaded in the "Protocol Name" field in the "Main" tab.
Press "START". The run will automatically stop when every collection is complete.
Note: Samples can be left in the BluePippin following elution overnight. Remove samples from the elution modules first thing the following morning.
For best recovery, wait at least 30-45 minutes after the end of the run before removing samples from the elution modules.
Remove samples from the elution modules using a standard P200 pipette. Aspirate slowly to minimize breakage of large DNA fragments. Samples will be ~80 µL.
Note: Additional 10-30% yield can often be achieved by rinsing the elution well with TE+0.1% Tween 20 (provided by Sage Sciences). Add 80 µL Tween solution to the elution well, wait 1 minute, then remove solution and pool with original extracted sample, or process separately.
Remove the cassette and dispose of properly. Do not keep used cassettes in the BluePippin with the cover closed. Humidity from the cassette may cause damage to the electrodes.
Quantify by taking a single measurement on the Qubit Flex Fluorometer with the Qubit 1x dsDNA BR Assay. Use 1 µL of DNA per measurement.
Size on the Agilent Femto Pulse System with the Genomic DNA 165 kb kit. The expected size range for samples post-size selection is ~24-40 kb, with a DQN > 9.8.

Figure 9. Femto Pulse trace post-shearing and size selection
Store at 4 °C until ready for library preparation. At least 2.5 µg DNA are required for library preparation. If necessary, repeat shearing and/or size selection to achieve required mass for library preparation.
Part 5a: Ligation Sequencing DNA V14 (SQK-LSK114) library preparation
Note: For automated library preparation, see Part 5b: Ligation sequencing DNA V14 (SQK-LSK114) library preparation on Hamilton NGS STAR.
Thaw the following reagents at room temperature, then store on ice:
- NEBNext FFPE DNA Repair Buffer
- Ultra II End-Prep Reaction Buffer
- Ligation Buffer (LNB)
- Elution Buffer (EB)
Place the following reagents at room temperature:
- AMPure XP beads
- Short Fragment Buffer (SFB)
- Qubit 1X HS dsDNA working solution and standards NOTE: Qubit 1X HS dsDNA working solution must be stored away from light
Prepare 2.5 µg DNA in 48 µL nuclease-free water in a 0.2 mL thin-walled PCR tube.
Note: If necessary, input volume may be greater than 48 µL to accommodate 2.5 µg DNA input mass. Input volume should not exceed 168 µL.
Prepare the following DNA repair enzyme master mix, adjusting the reagent volumes for the number of samples being processed:
- 3.5 µL NEBNext FFPE DNA Repair Buffer (vortex and spin down)
- 3.5 µL Ultra II End-Prep Reaction Buffer (vortex and spin down)
- 2 µL NEBNext FFPE DNA Repair Mix (do not vortex, spin down)
- 3 µL Ultra II End-Prep Enzyme Mix (do not vortex, spin down)
Pipette gently 10x to mix. Avoid pipetting bubbles.
Note: Keep NEB enzyme mixes in the freezer until use and return promptly.
Note: Prepare the DNA repair enzyme master mix with a 10% overage.
Add 12 µL DNA repair enzyme master mix. Pipette-mix or flick mix 10x, and spin down.
Incubate samples at 20 °C for 00:30:00 , followed by65 °C for 00:05:00 in a thermocycler.
Note: Start and pause thermocycler to allow lid to come to 85 °C before putting samples in.
Transfer samples to new 1.5mL Eppendorf DNA LoBind tubes.
Resuspend AMPure XP beads by vortexing.
Add 60 µL or equal sample volume of resuspended AMPure XP beads. Flick mix 10x and spin down.
Note: AMPure XP bead volume should be adjusted to match the input sample volume and enzyme mix post-incubation.
Incubate on platform rocker for00:05:00 at room temperature.
Prepare 500 µL of fresh 80% ethanol per sample in nuclease-free water.
Spin down samples and place on magnetic rack. Wait until supernatant is clear and colorless, about 00:02:00 .
Keep the tube on the magnetic rack and pipette off the supernatant.
With the samples remaining on the magnetic rack, add 200 µL of 80% ethanol. Pipette on the opposite wall to avoid disturbing the pellet. After 00:00:05 , remove the ethanol. Do not resuspend the beads in ethanol.
Note: If initial volume of beads was significantly higher than 60 µL, more ethanol may be required to keep the beads fully covered.
Repeat the previous step.
Spin down and place tubes on the magnetic rack. Pipette off any residual ethanol.
Allow to dry for ~00:00:30 , but do not overdry to the point of cracking.
Remove tubes from the magnetic rack. Add 60 µL nuclease-free water. Flick mix 10x and spin down.
Incubate for00:03:00 at 37 °C and 450 rpm in Thermomixer.
Spin down and place the samples on the magnetic rack until eluate is clear and colorless, about 00:02:00 .
Transfer 60 µL of eluate into a new 1.5 mL Eppendorf DNA LoBind tube.
Note: It is possible to store samples at 4 °C overnight at this step.
Prepare the following ligation enzyme master mix, adjusting the reagent volumes for the number of samples being processed:
- 25 µL Ligation Buffer (pipette-mix, spin down)
- 10 µL Quick T4 DNA Ligase (do not vortex, spin down)
- 5 µL Ligation Adapter (do not vortex, spin down)
Pipette gently 10x to mix. Avoid pipetting bubbles.
Note: Keep Quick T4 DNA Ligase and Ligation Adapter in the freezer until use and return promptly.
Note: Prepare the ligation enzyme master mix with a 10% overage.
Add 40 µL ligation enzyme master mix. Pipette-mix or flick mix 10x, and spin down.
Incubate the samples for 00:30:00 at room temperature.
Resuspend AMPure XP beads by vortexing.
Add 45 µL of resuspended AMPure XP beads. Flick mix 10x and spin down.
Incubate on platform rocker for 00:05:00 at room temperature.
Spin down samples and place on magnetic rack. Wait until supernatant is clear and colorless, about 00:02:00 .
Keep the tube on the magnetic rack and pipette off the supernatant.
With the samples remaining on the magnetic rack, add 250 µL of Short Fragment Buffer. Remove samples from the magnetic rack, flick mix until beads are fully resuspended, and spin down.
Wait until supernatant is clear and colorless, about 00:02:00 , then remove the supernatant.
Repeat the previous step.
Spin down and place tubes on the magnetic rack. Pipette off any residual Short Fragment Buffer.
Allow to dry for ~00:00:30 , but do not overdry to the point of cracking.
Remove tubes from the magnetic rack. Add 26 µL Elution Buffer. Flick mix 10x and spin down.
Incubate for 00:20:00 at 37 °C and 450 rpm in Thermomixer.
Spin down and place the samples on the magnetic rack until eluate is clear and colorless, about 00:02:00 .
Transfer 26 µL of eluate into a new 1.5 mL Eppendorf DNA LoBind tube.
Quantify by taking a single measurement on the Qubit Flex Fluorometer with the Qubit 1X dsDNA HS Assay. Use 1 µL of DNA per measurement.
Store at 4 °C until ready for loading. At least 750 ng DNA is required for loading for a 72 hour run. If necessary, repeat library preparation with additional sheared and size selected DNA to achieve required mass for loading. For long-term storage, store libraries at -80 °C .
Note: Recommended DNA input per load is 20 fmol, calculated to be 300 ng of 24kb dsDNA using https://nebiocalculator.neb.com/#!/dsdnaamt. For 3 full loads, 900 ng DNA is required. At a minimum, 750 ng is required for 2 full loads of 300 ng each, followed by an incomplete load comprised of the recovered DNA library from the first load and the remaining DNA library under 300 ng.
Note: If greater than 900 ng DNA library is available, PromethION flow cell may be loaded 4 times in a 96 hour run for improved data output.
Part 5b: Ligation Sequencing DNA V14 (SQK-LSK114) library preparation on Hamilton NGS STAR
Note: For manual library preparation, see Part 5a: Ligation sequencing DNA V14 (SQK-LSK114) library preparation.
Thaw the following reagents at room temperature, then store on ice:
- NEBNext FFPE DNA Repair Buffer
- Ultra II End-Prep Reaction Buffer
- Ligation Buffer (LNB)
- Elution Buffer (EB)
Place the following reagents at room temperature:
- AMPure XP beads
- Short Fragment Buffer (SFB)
- Qubit 1X HS dsDNA working solution and standards NOTE: Qubit 1X HS dsDNA working solution must be stored away from light
Prepare 2.5 µg DNA in 48 µL nuclease-free water in a BioRad Hard-Shell PCR Plate.
Note: DNA input mass should be 2 µg at a minimum. Input volume may not be greater than 48 µL .
Note: Samples must be processed in multiples of 8. If preparing less samples than a multiple of 8, fill remaining wells with 48 µL nuclease-free water.
Launch the LSK-114 protocol in the Hamilton Method Manager.
Note: Ensure the deck layout of the NGS STAR matches the protocol being run.
Note: Daily and weekly maintenance must be performed prior to beginning the protocol. Follow manufacturer guidelines for preventive maintenance and cleaning procedures.
Select "Process01: DNA repair and end-prep" as the start process.

Figure 10. Select start process
Select "Process02: DNA repair and end-prep clean-up" as the stop process.

Figure 11. Select stop process
Select the input file worklist.

Figure 12. Select input file worklist
Note: Samples must be processed in multiples of 8. If preparing less samples than a multiple of 8, fill remaining wells with 48 µL nuclease-free water.
Prepare the DNA repair enzyme mix in 2 mL Sarstedt tubes according to the instructions in the method. Avoid pipetting bubbles.

Figure 13. Prepare DNA repair enzyme mix
Load PCR ComfortLids onto the NGS STAR deck according to the instructions in the method.

Figure 14. Load PCR ComfortLids
Load MIDI plates and HSP plates onto the NGS STAR deck according to the instructions in the method.

Figure 15. Load MIDI and HSP plates
Load 50 µL tips onto the NGS STAR tip carriers according to the instructions in the method. Use the tip count screen to input loaded 50 µL tips.

Figure 16. Load 50 µL tips

Figure 17. Select loaded 50 µL tips
Note: Method should be started with full tip racks to avoid unnecessary errors.
Load 300 µL tips onto the NGS STAR tip carriers according to the instructions in the method. Use the tip count screen to input loaded 300 µL tips.

Figure 18. Load 300 µL tips

Figure 19. Select loaded 300 µL tips
Note: Method should be started with full tip racks to avoid unnecessary errors.
Load 80% ethanol in a 60 mL reagent trough in the reagent trough carrier according to the instructions in the method.

Figure 20. Load 80% ethanol
Load nuclease-free water in a 60 mL reagent trough and AMPure XP beads in a 20 mL reagent trough in the reagent trough carrier according to the instructions in the method.

Figure 21. Load nuclease-free water and AMPure XP beads
Note: AMPure XP beads must be resuspended by vortexing prior to use.
Load 1000 µL tips and the prepared sample plate with 48 µL DNA per well onto the NGS STAR deck according to the instructions in the method. Use the tip count screen to input loaded 1000 µL tips.

Figure 22. Load 1000 µL tips and DNA sample plate

Figure 23. Select loaded 1000 µL tips
Note: Method should be started with full tip racks to avoid unnecessary errors.
Note: Ensure the magnetic stand is placed on the carrier in the 5th position.
Load the DNA repair enzyme mix tubes onto the CPAC cooler according to the instructions in the method.

Figure 24. Load DNA repair enzyme mix
Note: Enzyme mixes should be spun down and placed on the NGS STAR deck with the caps removed. Take care to avoid pipetting bubbles in the tubes to avoid inaccurate dispensing.
Close the NGS STAR door and press 'Ok.' The protocol will now begin.
When prompted at the end of the run, remove the sample plate from the NGS STAR. Discard the reagent troughs and enzyme mix tubes.

Figure 25. Run completion dialog
Note: It is possible to store samples at 4 °C overnight at this step.
Launch the LSK-114 protocol in the Hamilton Method Manager.
Note: Ensure the deck layout of the NGS STAR matches the protocol being run.
Note: Daily and weekly maintenance must be performed prior to beginning the protocol. Follow manufacturer guidelines for preventive maintenance and cleaning procedures.
Select "Process03: Adapter ligation" as the start process.

Figure 26. Select start process
Select "Process04: Adapter ligation clean-up" as the stop process.

Figure 27. Select stop process
Select the input file worklist.

Figure 28. Select input file worklist
Note: Samples must be processed in multiples of 8. If preparing less samples than a multiple of 8, fill remaining wells with 60 µL nuclease-free water.
Prepare the ligation enzyme mix in 2 mL Sarstedt tubes according to the instructions in the method. Avoid pipetting bubbles.

Figure 29. Prepare ligation enzyme mix
Load MIDI plates and HSP plates onto the NGS STAR deck according to the instructions in the method.

Figure 30. Load MIDI and HSP plates
Load 50 µL tips onto the NGS STAR tip carriers according to the instructions in the method. Use the tip count screen to input loaded 50 µL tips.

Figure 31. Load 50 µL tips

Figure 32. Select loaded 50 µL tips
Note: Method should be started with full tip racks to avoid unnecessary errors.
Load 300 µL tips onto the NGS STAR tip carriers according to the instructions in the method. Use the tip count screen to input loaded 300 µL tips.

Figure 33. Load 300 µL tips

Figure 34. Select loaded 300 µL tips
Note: Method should be started with full tip racks to avoid unnecessary errors.
Load AMPure XP beads in a 20 mL reagent trough, Short Fragment Buffer in a 60 mL reagent trough, and Elution Buffer in a 60 mL reagent trough in the reagent trough carrier according to the instructions in the method.

Figure 35. Load AMPure XP beads, Short Fragment Buffer, and Elution Buffer
Note: AMPure XP beads must be resuspended by vortexing prior to use.
Load 1000 µL tips and the prepared sample plate with 60 µL DNA per well onto the NGS STAR deck according to the instructions in the method. Use the tip count screen to input loaded 1000 µL tips.

Figure 36. Load 1000 µL tips and DNA sample plate

Figure 37. Select loaded 1000 µL tips
Note: Method should be started with full tip racks to avoid unnecessary errors.
Note: Ensure the magnetic stand is placed on the carrier in the 5th position.
Load the ligation enzyme mix tubes onto the CPAC cooler according to the instructions in the method.

Figure 38. Load ligation enzyme mix
Note: Enzyme mixes should be spun down and placed on the NGS STAR deck with the caps removed. Take care to avoid pipetting bubbles in the tubes to avoid inaccurate dispensing.
Close the NGS STAR door and press 'Ok.' The protocol will now begin.
When prompted at the end of the run, remove the sample plate from the NGS STAR. Discard the reagent troughs and enzyme mix tubes.

Figure 39. Run completion dialog
Transfer26 µL of eluate into a new 1.5 mL Eppendorf DNA LoBind tube.
Quantify by taking a single measurement on the Qubit Flex Fluorometer with the Qubit 1X dsDNA HS Assay. Use 1 µL of DNA per measurement.
Store at 4 °C until ready for loading. At least 750 ng DNA is required for loading. If necessary, repeat library preparation with additional sheared and size selected DNA to achieve required mass for loading. For long-term storage, store libraries at -80 °C .
Note: Recommended DNA input per load is 20 fmol, calculated to be 300 ng of 24kb dsDNA using https://nebiocalculator.neb.com/#!/dsdnaamt. For 3 full loads,900 ng DNA is required. At a minimum, 750 ng DNA is required for 2 full loads of 300 ng each, followed by an incomplete load comprised of the recovered DNA library from the first load and the remaining DNA library under 300 ng.
Note: If greater than 900 ng DNA library is available, PromethION flow cell may be loaded 4 times in a 96 hour run for improved data output.
Part 6: Loading the PromethION Flow Cell
Thaw the following reagents at room temperature:
- Sequencing Buffer (SB)
- Library Solution (LIS)
- Flow Cell Tether (FCT)
- Flow Cell Flush (FCF)
- Elution Buffer (EB)
Mix by vortexing, spin down, and place on ice.
Take flow cells out of the fridge. Allow to sit at room temperature for 00:20:00 .
Note: Condensation can form on the flow cell in humid environments. Inspect the gold connector pins on the top and underside of the flow cell for condensation and wipe off with a lint-free wipe if any is observed. Ensure the heat pad (black pad) is present on the underside of the flow cell.
Load flow cells into the PromethION docking ports. Perform flow cell check prior to flow cell priming.
Note: Only flow cells with a starting pore count greater than 6500 pores should be used for sequencing runs. Flow cells with starting pore counts less than 6500 pores can be used to generate additional sequencing data after an initial run.
Prepare the flow cell priming mix by combining 1170 µL Flow Cell Flush and30 µL Flow Cell Tether per sample. Mix by vortexing.
Prepare 300 ng DNA library in 32 µL Elution Buffer in a 1.5 mL Eppendorf DNA LoBind tube.
Note: Recommended DNA input is 20 fmol, calculated to be 300 ng of 24kb dsDNA using https://nebiocalculator.neb.com/#!/dsdnaamt.
Add 100 µL Sequencing Buffer (SB).
Add 68 µL Library Solution (LIS).
Slide the flow cell inlet port cover clockwise to open. Draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µL
- Insert the tip into the inlet port
- Turn the wheel until dial shows 220-230 µL, until a small volume of buffer enters the pipette tip
Note: Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µL, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
Load 500 µL of the priming mix into the flow cell via the inlet port, avoiding the introduction of air bubbles.
Wait 00:05:00 .
Load 500 µL of the priming mix into the flow cell via the inlet port, avoiding the introduction of air bubbles.
Mix the prepared library gently by pipetting up and down 5x just prior to loading.
Load 200 µL of library into the inlet port using a P1000 pipette.
Close the valve to seal the inlet port. Install the light shield. Close the PromethION door.
Wait 00:10:00 before initiating sequencing run in MinKNOW:
- Navigate to the start page and click "Start sequencing"
- Fill in the experiment name and sample ID, select the flow cell position, and load run configuration preset (if applicable)
- Select the sequencing kit used in the library preparation (SQK-LSK114-XL) on the Kit page
- Configure sequencing and output parameters - Fast basecalling model, 72 or 96 hour run time, basecalled output off, raw reads output .POD5, and minimum Q score of 8
- Click "Start" to initiate the sequencing run
Within the first hour of sequencing, pay attention to pore occupancy and pore scan results. If necessary to resolve sequencing issues related to sample or flow cell quality:
- Perform pore scan on flow cell and note if it resolves pore count issues.
- Stop the sequencing run, remove the flow cell from the PromethION, and insert it again into a different position. Restart the run and note if it resolves pore count issues.
- Stop the sequencing run. Recover DNA library and load onto a new flow cell, following protocol from step 170.
Following 72 hours of sequencing and 3 loads of prepared DNA libraries, the sample should yield a data output ~90-100 Gb with an N50 ~20-30 kb.

Figure 40. Read length distribution for brain tissue sample with a 72 hour sequencing run

Figure 41. PromethION pore scan results for brain tissue sample with a 72 hour sequencing run and 3 loads of DNA library
Part 7: Washing and reloading the PromethION Flow Cell
Thaw the following reagents at room temperature:
- Sequencing Buffer (SB)
- Library Solution (LIS)
- Flow Cell Tether (FCT)
- Flow Cell Flush (FCF)
- Elution Buffer (EB)
- Wash Diluent (DIL)
Mix by vortexing, spin down, and place on ice.
Prepare the flow cell wash mix by combining 398 µL Wash Diluent and 2 µL Wash Mix per sample. Mix by pipetting. Do not vortex.
Note: Keep Wash Mix (WMX) in the freezer until use and return promptly. Do not vortex.
Note: Prepare the flow cell wash mix with a 10% overage.
Pause the sequencing run in MinKNOW.
If necessary, recover the already loaded DNA library:
- Slide the inlet port cover clockwise to open
- Set a P1000 pipette tip to 200 µL
- Insert the tip into the inlet port
- Turn the wheel until dial shows 400 µL
Note: Do not attempt to recover the loaded library after removing waste from the flow cell to prevent air from being drawn across the sensor array area, which would lead to a significant loss of sequencing channels.
Note: DNA library recovery is necessary for samples with less than 900 ng of available DNA library for loading, and can be used for the third load in place a complete 300 ng DNA library load.
Remove waste buffer from flow cell:
- Close the inlet port
- Insert a P1000 pipette into waste port 2 or 3 and remove the waste buffer
Note: It is vital that the inlet port is closed before removing waste to prevent air from being drawn across the sensor array area, which would lead to a significant loss of sequencing channels.
Slide the inlet port cover clockwise to open. Draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µL
- Insert the tip into the inlet port
- Turn the wheel until dial shows 220-230 µL, until a small volume of buffer enters the pipette tip
Note: Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µL, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
Load 400 µL of the flow cell wash mix into the flow cell via the inlet port, avoiding the introduction of air bubbles. Close the inlet port and ensure the light shield is installed.
Wait 01:00:00 .
Prepare the flow cell priming mix by combining 1170 µL Flow Cell Flush and30 µL Flow Cell Tether per sample. Mix by vortexing.
Prepare 300 ng DNA library in 32 µL Elution Buffer in a 1.5 mL Eppendorf DNA LoBind tube.
Note: Recommended DNA input is 20 fmol, calculated to be 300 ng of 24kb dsDNA using https://nebiocalculator.neb.com/#!/dsdnaamt.
Note: If available sample mass for the third load is under 300 ng, DNA library should be recovered for the first load and used for the third load. Additional DNA library can be spiked in to the recovered DNA library.
Add 100 µL Sequencing Buffer (SB).
Add 68 µL Library Solution (LIS).
Slide the inlet port cover clockwise to open. Draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µL
- Insert the tip into the inlet port
- Turn the wheel until dial shows 220-230 µL, until a small volume of buffer enters the pipette tip
Note: Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µL, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
Load 500 µL of the priming mix into the flow cell via the inlet port, avoiding the introduction of air bubbles.
Wait 00:05:00 .
Load 500 µL of the priming mix into the flow cell via the inlet port, avoiding the introduction of air bubbles.
Mix the prepared library gently by pipetting up and down 5x just prior to loading.
Load 200 µL of library into the inlet port using a P1000 pipette.
Close the valve to seal the inlet port. Install the light shield. Close the PromethION door.
Wait 00:10:00 before resuming sequencing run in MinKNOW.
Within the first hour of sequencing, pay attention to pore occupancy and pore scan results. If necessary to resolve sequencing issues related to sample or flow cell quality:
- Perform pore scan on flow cell and note if it resolves pore count issues.
- Stop the sequencing run, remove the flow cell from the PromethION, and insert it again into a different position. Note if it resolves pore count issues.
- Stop the sequencing run. Recover DNA library and load onto a new flow cell, following protocol from step 192.
Following 72 hours of sequencing and 3 loads of prepared DNA libraries, the sample should yield a data output ~90-100 Gb with an N50 ~20-30 kb.

Figure 42. Read length distribution for brain tissue sample with a 72 hour sequencing run

Figure 43. PromethION pore scan results for brain tissue sample with a 72 hour sequencing run and 3 loads of DNA library
Protocol references
TissueLyser III User Manual (Qiagen): https://www.qiagen.com/us/resources/resourcedetail?id=c41836ee-0782-486c-9f1e-61f21f955588&lang=en
Megaruptor 3 DNAFluid+ Kit Guide (Diagenode): https://www.diagenode.com/files/products/kits/protocol-DNAFluid-V1.pdf
High-Pass DNA Size Selection User Guide (Sage Science): https://sagescience.com/wp-content/uploads/2019/01/BPLUS10-user-guide-460045-Rev-E.pdf
Ligation sequencing DNA V14 (SQK-LSK114) library preparation (Oxford Nanopore): https://nanoporetech.com/document/genomic-dna-by-ligation-sqk-lsk114?device=PromethION
Washing and reloading the PromethION Flow Cell (Oxford Nanopore): https://nanoporetech.com/document/flow-cell-wash-kit-exp-wsh004
