Jul 28, 2022
Version 1
Preparing 1x PCR Master Mix V.1
This protocol is a draft, published without a DOI.
- Jenny Molloy1,
- Stephane Fadanka2,
- Nadine Mowoh2,
- Cordellia Cordellia Fulai3
- 1University of Cambridge, Beneficial Bio;
- 2Mboalab, Beneficial Bio;
- 3Mboalab
- Beneficial Bio
- Low-cost, high-quality ...

Protocol Citation: Jenny Molloy, Stephane Fadanka, Nadine Mowoh, Cordellia Cordellia Fulai 2022. Preparing 1x PCR Master Mix. protocols.io https://protocols.io/view/preparing-1x-pcr-master-mix-b5qbq5sn
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: February 28, 2022
Last Modified: July 28, 2022
Protocol Integer ID: 58851
Keywords: Preparing 1x PCR Master mix, 1x PCR Master mix formulation, production of benbio 1x pcr master mix, preparing 1x pcr master mix, benbio 1x pcr master mix, 1x pcr master mix this protocol document, pcr master mix, openvent cellular reagent, cellular reagent, subsequent pcr reaction, wet formulation, different colors of the wet formulation, pcr, reagent, formulation
Abstract
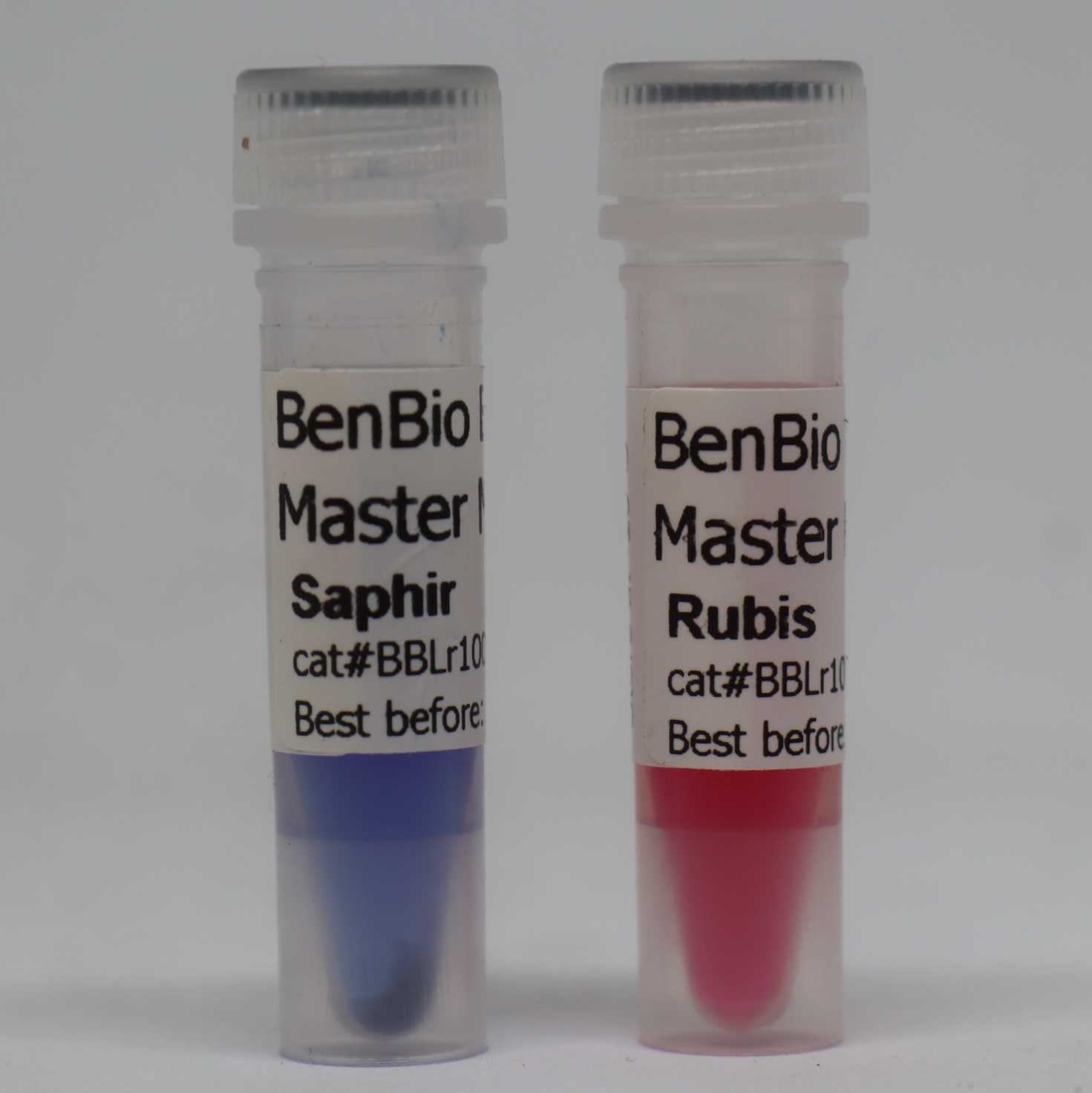
This protocol documents the production of BenBio 1X PCR Master Mix "Wet" and "Dry" formulations including ‘the different colors of the Wet formulations (Rubis or oink and Saphir or blue). This method uses OpenVent cellular reagents. The PCR master mixes are then stored for subsequent PCR reactions.
Cellular reagents are defined as common molecular biology enzymes expressed in E.coli but not subsequently purified before use i.e. dried E.coli cells are used as the reagent (Bhadra et al (2018)).
Guidelines
This protocol describes the steps in preparing and testing the functionality of 1X PCR master mix.
Materials
Equipment
Thermocycler
Micropipette
0.2ul PCR tubes
1.5ml Eppendorf tubes
Microwave
Gel casting tray
Well comb
UV transilluminator
Voltage source (Electrophoresis unit)
Reagents
OpenVent DNA polymerase
PCR Primers
Agarose (electrophoresis grade)
DNA template PCR amplicon ( Lambda 0.5 and 1kb or other)
DNA loading dye (6x NEB)
DNA ladder (Bioline 1kb)
DNA gel stain (SYBR Safe or other Ethidium bromide, EtBr stain)
Troubleshooting
Safety warnings
- Wear protective clothing like Lab coats, with gloves and face masks during the process and avoid dust formation from any powders.
- Take special care when handling the EtBr gel stain and the UV transilluminator
Before start
- Ensure that the cellular reagent (OpenVent enzyme) to be used is available and has been pre-tested for functionality.
- Ensure that all the other components needed to prepare the PCR mix are available and free from contaminants..
Cellular reagents preparation
Note
Before making Master mixes, we typically start by producing the cellular reagents which will be used. The cellular reagents then under go a quality control tests to ensure the enzymes are functional and free from contaminating nucleases.
- Prepare a fresh batch of cellular reagents (OpenVent) following BenBio protocol for plate protein expression on autoinduction media.
1d
Functionality test
Remove the enzymes from storage, reconstitute and test for functionality as described in the BenBio protocol using the specific test that apply for cellular reagents (OpenVent enzyme).
2h 30m
Preparation of 1x PCR Master mix formulations
Preparing the work surface and materials:
- Clean the working surface first with 1:10 dilution of Bleach, then70 % (v/v) Ethanol
- Clean the micropipettes with 70 % (v/v) alcohol then with Lookout DNA erase solution; keep on a clean surface throughout manipulation.
- Crush enough ice to fill the PCR box or a beaker bowl till it's 3/4 filled.
- Remove all reagents needed from -20 °C and 4 °C and thaw On ice
- Prepare a sterile labelled 1.5 mL microcentrifuge tubes or larger volume tubes depending on amount of PCR mix needed, and leave open On ice .
5m
Pipetting components to make up the "Wet" mix
- Pipette the corresponding amount of PCR grade water to reconstitute the enzyme as was determined during preparation (in this case 30 uL PCR water is used to reconstitute the enzyme) and place on ice.
- Pipette all reagents in the tube following the order stated in the table below:
Note
- We adopt 2 formulations of 1x PCR Master mix using 2 different dyes as trackers ( Azorubine and Bromophenol blue), and have 2 states of the mix formulations as "Wet" and "Dry".
- Also a 2x concentration of the PCR master mix can be done to reduce the volume of water used for the preparation to ease drying and enhance shelf life of the PCR master mix formulation.
| A | B | |
| Components | Amount in μL for 1x concentration of PCR Master mix | |
| PCR grade H2O | 8.8 | |
| dNTPs mix (25 mM) | 0.6 | |
| 10x Thermopol Buffer | 2 | |
| MgSO4 (100mM) | 0.6 | |
| Cellular reagents (Freshly prepared and heat treated) | 3 | |
| 20% (w/v) Trehalose stock solution | 4.5 | |
| Bromophenol Blue 0.25% (w/v)Azorubine 0.25% (w/v) | 0.5 | |
| 20 |
- Determine the total volume needed and make necessary calculations to adjust the amount of each component to be pipetted.
- Mix well by gentle agitations (avoid using vortex as the mixture will foam)
Note
- At this point the Wet master mix formulation is ready for use after performing functionality tests and can be aliquoted into desired size screw cap tubes in the desired volume for long term storage (avoid frequent freezing and thawing).
- If the dry formulation is desired, continue with the steps described below.
Aliquoting to make a "Dry" mix
- Carefully pipette 25 µL of Mix formulation into each 0.2ml PCR tubes
- Place PCR tubes (left Opened) in an Airtight container filled halfway with silica beads.
- Place the container in an incubator and dry at37 °C Overnight
- Store dried down tubes at4 °C in air-tied sachet or containers filled halfway with silica beads.
10m
Quality control - Functionality testing
2h 30m
Check the functionality of the produced PCR Master Mix by running control PCR reactions.
We generally use the BenBio internal protocol which can be modified depending on the PCR master mix formulation and concentration being tested.
Note
For functionality of 1x Wet mix formulation we make a 20 µL total volume PCR reaction as follows:
- Pipette 17 µL of the mix into a 0.2 mL PCR tube and add1 µL of each PCR primer and 1 µL of DNA template and run the thermocycling process following the specific protocol for the DNA template and primers used.
For 1x Dry formulation:
- Rehydrate the dried mix tube with 17 µL PCR water add1 µL of each PCR primer and 1 µL of DNA template and run the thermocycling process following the specific protocol for the DNA template and primers used.
2h
Running Agarose gel and Visualization
After running the PCR reaction, the samples are visualized to check for amplification as follows:
- Prepare 1.5% agarose gel and run the gel electrophoresis to completion.
- Visualize to check for DNA bands which signify amplification to show that the enzyme or PCR master mix are functional (able to amplify a specific region of a test DNA template).
30m
