Oct 14, 2025
Version 1
mtscMultiome ATAC + GEX V.1
- Marc Appis1,2,3,
- Yu-Hsin Hsieh1,2,
- Pauline Kautz1,2,4,
- Lena Nitsch1,2,3,
- Maren Salla1,2,3,
- Klara Penkert1,2,
- Caleb Lareau5,
- Patrick Maschmeyer1,2,
- Leif S. Ludwig1,2
- 1Berlin Institute of Health at Charité - Universitätsmedizin Berlin (BIH);
- 2Max‐Delbrück‐Center for Molecular Medicine in the Helmholtz Association (MDC), Berlin Institute for Medical Systems Biology (BIMSB);
- 3Freie Universität Berlin, Department of Biology/Chemistry/Pharmacy;
- 4Technische Universität Berlin, Institute of Biotechnology;
- 5Computational and Systems Biology Program, Memorial Sloan Kettering Cancer Center

Protocol Citation: Marc Appis, Yu-Hsin Hsieh, Pauline Kautz, Lena Nitsch, Maren Salla, Klara Penkert, Caleb Lareau, Patrick Maschmeyer, Leif S. Ludwig 2025. mtscMultiome ATAC + GEX. protocols.io https://dx.doi.org/10.17504/protocols.io.36wgqq2w3gk5/v1
Manuscript citation:
Mimitou EP, Lareau CA, Chen KY, Zorzetto-Fernandes AL, Hao Y, Takeshima Y, Luo W, Huang TS, Yeung
BZ, Papalexi E, Thakore PI, Kibayashi T, Wing JB, Hata M, Satija R, Nazor KL, Sakaguchi S, Ludwig LS, Sankaran VG, Regev A, Smibert P. Scalable, multimodal profiling of chromatin accessibility, gene
expression and protein levels in single cells. Nat Biotechnol. 2021 Oct;39(10):1246-1258. doi: 10.1038/s41587-021-00927-2. Epub 2021 Jun 3. PMID: 34083792; PMCID: PMC8763625.
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
This protocol is an implementation of the protocol published in Nature Biotechnology (see literature reference) and is successfully used in our lab on a regular basis. However, adaptions for application in another lab or for different target cell types might be necessary.
Created: June 26, 2025
Last Modified: October 14, 2025
Protocol Integer ID: 221118
Keywords: Single cell multiomics, Mitochondrial genomics, scATAC-seq, scRNA-seq, mitochondrial genome into the 10x genomics multiome atac, capture of mitochondrial dna, mitochondrial genome, mitochondrial dna, gex mtscmultiome atac, mtdna, 10x genomics multiome atac, mtscmultiome atac, epi multiome atac, cytometric enrichment, atac library generation, gex libraries ready for submission, human peripheral blood mononuclear cell, gex library, cryopreserved cell, desired cell population
Funders Acknowledgements:
National Human Genome Research Institute
Grant ID: UM1 HG012076
Disclaimer
The authors of this protocol don't take any legal responsibility for usage of this protocol, neither they give any guaranty for successful execution.
Abstract
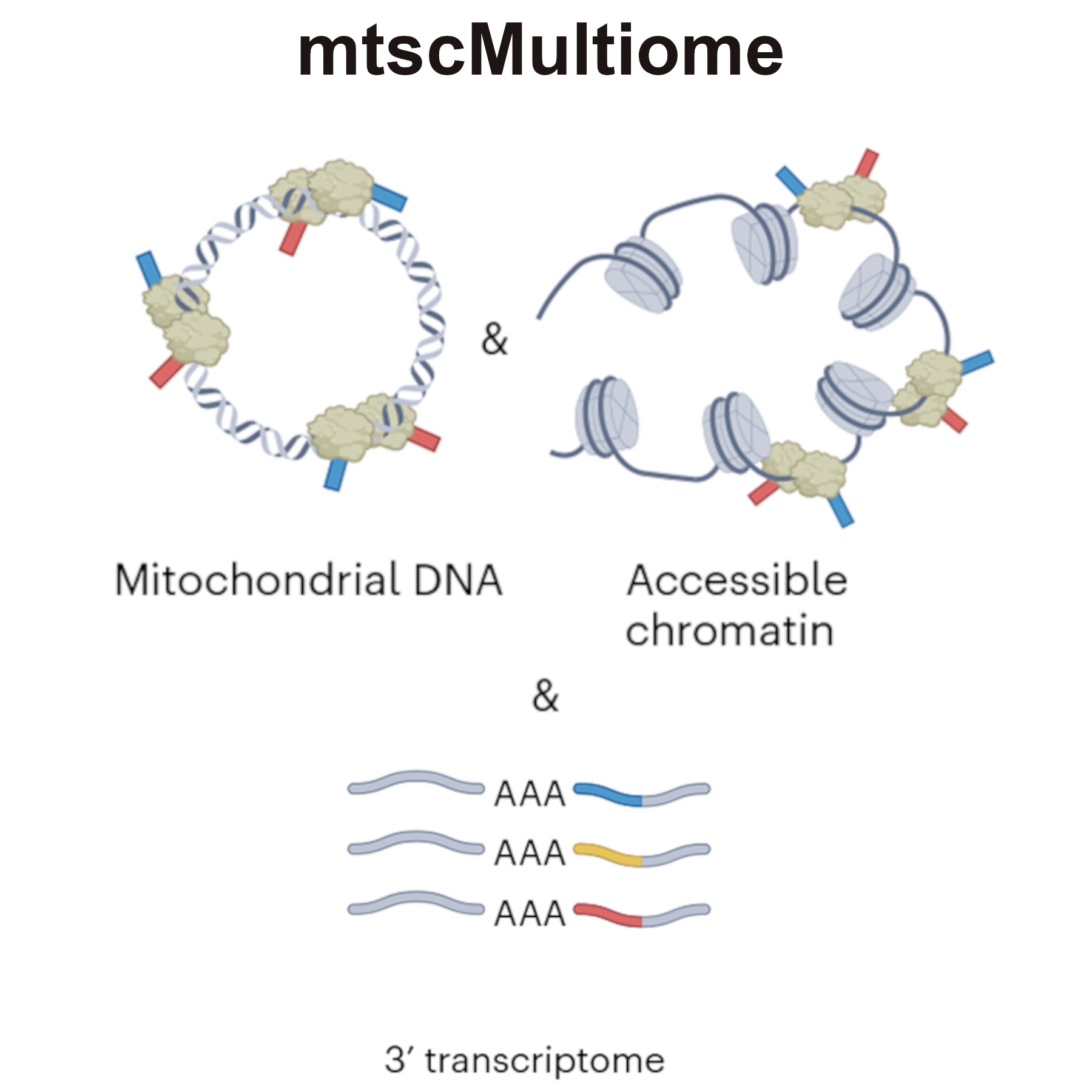
mtscMultiome ATAC + GEX integrates the capture of mitochondrial DNA (mtDNA) for variant calling from the mitochondrial genome into the 10x Genomics Multiome ATAC + GEX workflow (now Epi Multiome ATAC + Gene Expression). The protocol was developed with human peripheral blood mononuclear cells but can be adapted for other cell types, specimens, and species, which may require further optimization.
The protocol starts from cryopreserved cells and includes (1) thawing, washing, and staining of surface markers for flow cytometric enrichment of desired cell populations, (2) cell fixation, permeabilization, and tagmentation, (3) GEM generation and barcoding, (4) post-GEM incubation cleanup, (5) pre-amplification PCR, (6) ATAC library generation, (7) cDNA amplification, and final (8) GEX library construction to yield ATAC and GEX libraries ready for submission to NGS.
The original version of this protocol was published by Mimitou et al., Nat Biotechnol. 2021 (doi: 10.1038/s41587-021-00927-2).
Attachments
Image Attribution
Modified based on Nitsch L, Lareau CA, Ludwig LS. Mitochondrial genetics through the lens of single-cell multi-omics. Nat Genet. 2024 Jul;56(7):1355-1365. doi: 10.1038/s41588-024-01794-8. Epub 2024 Jul 1. PMID: 38951641; PMCID: PMC11260401.
Guidelines
This protocol follows the 10x Genomics Chromium Next GEM Single Cell Multiome ATAC + GEX workflow. Please consult the corresponding user guide before. The present protocol is based on the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression User Guide • Rev F (see supplementary information to this protocol).
Starting from the tagmentation, important deviations from the 10x workflow are highlighted in this Protocols.io version. However, it is particularly useful to check reagent handling instructions, troubleshooting, and Chip loading with the Chromium Multiome user guide.
Please note that initial sample preparation and handling depend very much on the desired starting material and thus, sample handling until tagmentation might be adapted. This version of mtscMultiome ATAC + GEX was adapted to analyze human primary peripheral blood mononuclear cells (PBMCs).
Materials
Equipment required for mtscMultiome ATAC + GEX
- Standard laboratory equipment (microliter pipets and tips, multichannel pipets (ideally Rainin), tubes (PCR tubes, 1.5 mL, 2 mL, 15 mL, 50 mL), table-top centrifuges, Vortexer/shaker, heating block, serological pipets & tips)
- Vortexer with 10X adapter
- FACS tubes with filter (e.g., Cell Strainer Snap Cap)
- Tissue Culture Hood (clean bench)
- Swinging bucket centrifuge (with inlets for 50 mL, 15 mL, FACS tubes)
- Water bath (up to 37°C)
- Heat block (for microreaction tubes, up to 65°C)
- Cell culture microscope
- Cell counting equipment: either manual (i.e., Neubauer counting chamber) or automated (e.g., Countess 3 FL Automated Cell Counter)
- FACS sorter (if L/D and negative selection of neutrophils shall be performed; panel needs to be adapted based on the used FACS sorter)
- Chromium iX/X (Controller)
- 10x Magnetic Separator
- Dry ice
- BioAnalyzer (Agilent) (recommended) or TapeStation (Agilent) + corresponding hsDNA kit
- Qubit hsDNA Kit (Invitrogen)
Reagents required for mtscMultiome ATAC + GEX
1) For Day 1 (Steps 1 - 3)
| Reagent | Manufacturer / Vendor | Product ID | Link | Can be replaced? | Amount per sample | |
| Penicillin-Streptomycin (10,000 U/ml; 10 mg/ml) | PAN Biotech | #P06-07100 | https://shop.pan-biotech.de/en/Penicillin-Streptomycin-10.000-U-ml-Penicillin-10-mg-ml-Streptomycin/P06-07100 | Optional supplement for thawing media, can be replaced by P/S from different manufacturer | 500 µL | |
| FBS Standard | PAN Biotech | #P30-3306 | https://shop.pan-biotech.de/en/FBS-Standard-South-America-origin-fetal-bovine-serum-0.2-m-sterile-filtered/P30-3306 | Necessary part of thawing media, can replaced by FBS from different manufacturer | 5 mL | |
| RPMI 1640 Glutamax | Thermo Fisher | #61870010 | https://www.thermofisher.com/order/catalog/product/61870010 | Necessary main part of the thawing media, can be replaced by RPMI from different manufacturer; Can be replaced by other full media (HPLM, DMEM etc.) | 50 mL | |
| BSA stock | Miltenyi Biotec | #130-091-376 | https://www.miltenyibiotec.com/DE-en/products/macs-bsa-stock-solution.html | Required for generation of protein-rich buffers; Can be replaced by BSA from different manufacturer | 440 µL | |
| PBS | Thermo Fisher | #14190094 | https://www.thermofisher.com/order/catalog/product/14190094 | Can be replaced by sterile, Nuclease-free PBS from different manufacturer | 13 mL | |
| Protector RNase Inhibitor (40 U/µl) | Merck Millipore | #3335399001 | https://www.sigmaaldrich.com/DE/de/product/roche/rnainhro | Could be replaced by RNase inhibitor from different manufacturer (with same activity/concentration); however, replacement is not recommended | 31 µL | |
| NP-40 (10%) | Merck Millipore | #11332473001 | https://www.fishersci.de/shop/products/nonidet-p40-substitute-ultrapure-thermo-scientific/15875388?srsltid=AfmBOoragyw-6maozyGoIM59qTAWc5CKth054ntYJf8m1BFAnau9Xb5V | Can be replaced by NP-40 from different manufacturer | 1.1 µL | |
| Dithiothreitol (DTT) (100 mM) | Thermo Fischer | #707265ML | https://www.thermofisher.com/order/catalog/product/707265ML | Can be replaced by DTT from different manufacturer | 12.1 µL | |
| UltraPure Nuclease-free water | Thermo Fisher | #10977-035 | https://www.thermofisher.com/order/catalog/product/10977035 | Can be replaced by nuclease-free water from different manufacturer | 460 µL | |
| Tris-HCl | SigmaAldrich | #T2194-100ML | https://www.sigmaaldrich.com/DE/de/product/sigma/t2194 | Can be replaced by Tris_HCl from different manufacturer | ||
| NaCl (5M) | SigmaAldrich | #59222C-500ML | https://www.sigmaaldrich.com/DE/de/product/sigma/59222c | Can be replaced by NaCl from different manufacturer | ||
| MgCl2 (1M) | SigmaAldrich | #M1028-100ML | https://www.sigmaaldrich.com/DE/de/product/sigma/m1028 | Can be replaced by MgCl2 from differemt manufacturer | ||
| Trypan Blue | ThermoFischer | #T10282 | https://www.thermofisher.com/order/catalog/product/T10282 | Can be replaced by Trypan Blue from different manufacturer or by similar L/D staining dye | 20-30 µL | |
| Pierce™ 16 % Formaldehyde (w/v), methanol-free | Thermo Fisher | #28908 | https://www.thermofisher.com/order/catalog/product/28908 | Can be replaced by PFA from different manufacturer; should have molecular biology grade and be methanol-free | 3 µL | |
| Glycine | SigmaAldrich | #50046-50G | https://www.sigmaaldrich.com/DE/de/product/sigma/50046 | Can be replaced with glycine (molecular biology grade) from a different manufacturer | 25 µL of a 2.5 M solution | |
| Glycerol (50% v/v) | Ricca Chemical Company | #3290-32 | https://www.riccachemical.com/products/general-use-reagents/other-aqueous-solutions/3290-(-r3290000-) | Can be replaced by Glycerol 50% (v/v) from a different manufacturer | ~5 mL |
Reagents required for day 1 of the mtscMultiome protocol
The following table includes the components of the recommended panel for FACS-based sorting of live CD66b- PBMCs. It can be adapted based on the target cell type and lab-specific requirements.
| Antigen | Colour / channel | Recommended product | Link | Notes | Amount per sample | |
| CD14 | FITC (= AF488) | BioLegend AlexaFluor 488 anti-human CD14 (clone 63D3) #367130 | https://www.biolegend.com/en-us/products/alexa-fluor-488-anti-human-cd14-antibody-15066 | Can be replaced by PE anti-human CD14 from another manufacturer (new titration might be necessary) | 1 µL | |
| CD66b | PE | Biolegend PE anti-human CD66b /clone G10F5) #305106 | https://www.biolegend.com/en-us/products/pe-anti-human-cd66b-antibody-6529 | Can be replaced by PE anti-humanCD66b from another manufacturer (new titration might be necessary) | 1 µL | |
| FcX block | n/a | Human TruStain FcX (BioLegend, #422301) | https://www.biolegend.com/en-us/products/human-trustain-fcx-fc-receptor-blocking-solution-6462 | Recommended to prevent non-specific binding of antibodies; can be replaced by other FcX-receptor block | 2.5 µL | |
| L/D | Sytox Blue (DAPI) | SYTOX Blue dead cell stain (Invitrogen, #S34857) | https://www.thermofisher.com/order/catalog/product/S34857 | Can be replaced by other L/D dye in the same channel | <1 µL |
Recommended FACS panel for sorting of viable non-CD66b PBMCs.
The following table gives an overview on the components of the 10X Chromium Next GEM Single Cell Multiome ATAC + GEX kit required for day 1 of the mtscMultiome ATAC + GEX protocol. We strongly suggest to follow the handling instructions as provided in the kit manual. Furthermore, these items should not be replaced.
| Item | 10x Product number | Kit Box | Amount per 1 sample + 10% | |
| 20X Nuclei Buffer | 2000207 | ATAC Kit A | 2.75 µL | |
| ATAC Buffer B | 2000193 | ATAC Kit A | 7 µL | |
| ATAC Enzyme B | 2000265 | ATAC Kit A | 3 µL | |
| Barcoding Reagent Mix | 2000267 | GEM Kit A | 54.45 µL | |
| Barcoding Enzyme Mix | 2000266 | GEM Kit A | 8.25 µL | |
| Template Switch Oligo | 3000228 | GEM Kit A (after resuspension to be stored at -80°C) | 1.21 µL | |
| Reducing Agent B | 2000087 | GEM Kit A | 2.1 µL | |
| Quenching Agent | 2000269 | GEM Kit A | 5 µL | |
| Single Cell Multiome Gel Beads A | 2000261 | Gel Bead Kit A | 50 µL | |
| Chromium Next GEM Chip J | 2000264 | Next GEM Chip J & Gaskets | 1 piece (up to 8 samples) | |
| Gasket | 370017 | Next GEM Chip J & Gaskets | 1 piece (1 per Chip J) | |
| Partitioning Oil | 2000190 | Chromium Partitioning Oil | 2-3 mL |
Reagents from the 10x Chromium Next GEM Single Cell Multiome kit required for the first part of the mtscMultiome ATAC + GEX protocol (fixation, lysis, tagmentation, GEM generation, RT-PCR & quenching)
2) For day 2 (steps 4 - 5)
| Reagent | Manufacturer | Product ID | Link | Can be replaced | Amount per sample | |
| Dynabeads MyOne SILANE | 10x Genomics | 2000048 | https://cdn.10xgenomics.com/image/upload/v1660261285/support-documents/2000048_Dynabeads___MyOne___SILANE_sds.pdf | Can't be replaced | 14.3 µL | |
| Elution buffer (EB) | Qiagen | 19086 | https://www.qiagen.com/de/products/discovery-and-translational-research/lab-essentials/buffers-reagents/buffer-eb?catno=19086 | Might be replacable. However, replacement by a different elution buffer was not tested and is therefore not recommended. | 220 µL | |
| 10% Tween-20 | Can be replaced by 10% Tween-20 from a different manufacturer | <1 µL | ||||
| Ethanol (HPLC grade) | Carl Roth Chemie | 9065.1 | https://www.carlroth.com/com/en/alcohols/ethanol/p/9065.1 | Can be replaced by analytical/HPLC-grade ethanol from a different manufacturer | 2 mL | |
| UltraPure Nuclease-free water | ThermoFischer | 10977-035 | https://www.thermofisher.com/order/catalog/product/10977035 | Can be replaced by nuclease-free water from different manufacturer | <2 mL | |
| SPRIselect DNA Size Selection Reagent | Beckman Coulter Life Sciences | B23317 | https://www.beckman.de/reagents/genomic/cleanup-and-size-selection/size-selection/b23317 | Can't be replaced | 250 µL |
Reagents required for day 2 of the mtscMultiome protocol
The following table gives an overview on the components of the 10X Chromium Next GEM Single Cell Multiome ATAC + GEX kit required for day 2 of the mtscMultiome ATAC + GEX protocol. We strongly suggest to follow the handling instructions as provided in the kit manual. Furthermore, these items should not be replaced.
| Item | 10x Product number | Kit box | Amount per 1 sample | |
| Cleanup Buffer | 2000088 | GEM Kit A | 200.2 µL | |
| Reducing Agent B | 2000087 | GEM Kit A | <1 µL | |
| Recovery Agent | 220016 | Chromium Recovery Agent | 125 µL | |
| Pre-Amp Mix | 2000274 | Amp Kit A | 50 µL | |
| Pre-Amp Primers | 2000271 | Amp Kit A | 4 µL |
Reagents from the 10x Chromium Next GEM Single Cell Multiome kit required for the second part of the mtscMultiome ATAC + GEX protocol (Post GEM incubation cleanup and pre-amplification PCR & cleanup)
3) For day 3 (step 6)
For ATAC library construction, freshly prepared 80% ethanol (500 µL per sample), SPRIselect reagent (155 µL per sample), nuclease-free water (15 µL per sample) and Elution buffer (21 µL per sample) are required. For reagent details see table at day 2. Additionally, Qubit & BioAnalyzer reagents are required for library QC.
| Item | 10x Product Number | Kit box | Amount per 1 sample | |
| Amp Mix | 2000047 | Amp Kit A | 50 µL | |
| SI-PCR Primer B | 2000128 | Amp Kit A | 7.5 µL | |
| Single Index Plate N Set A | 3000427 | Single Index Kit N Set A | 2.5 µL |
Reagents from the 10x Chromium Next GEM Single Cell Multiome kit required for the third part of the mtscMultiome ATAC + GEX protocol (ATAC library construction)
4) For day 4 (step 7)
For cDNA amplification, the following materials are required (see table at day 2 for more details on the reagents):
- SPRIselect reagent: 60 µL per sample
- Freshly prepared 80% ethanol: 400 µL per sample
- Elution buffer: 40.5 µL per sample
- BioAnalyzer reagents
| Item | 10x Product number | Kit box | Amount per 1 sample | |
| Amp mix | 2000047 | Amp Kit A | 55 µL | |
| cDNA Primers | 2000089 | Amp Kit A | 16.5 µL |
Reagents from the 10x Chromium Next GEM Single Cell Multiome kit required for the fourth part of the mtscMultiome ATAC + GEX protocol (cDNA amplification)
5) Step 8
This step is identical to the 10x Chromium Next GEM Single Cell Multiome ATAC + GEX kit step 7 (Gene Expression Library Construction. Please refer to the user guide for required materials.
Protocol materials
SYTOX™ Blue Dead Cell Stain, for flow cytometryThermo FisherCatalog #S34857
Human TruStain FcX™BioLegendCatalog #422301
NP-40 10% Merck MilliporeSigma (Sigma-Aldrich)Catalog #11332473001
MACS BSA Stock SolutionMiltenyi BiotecCatalog # 130-091-376
DTT (100mM Solution)Thermo Fisher ScientificCatalog #707265ML
Protector RNase Inhibitor (40 U/ul) 2000 unitsMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335399001
Nuclease-free Water
1X PBS
Pierce 16% Formaldehyde (w/v), Methanol-freeThermo Fisher ScientificCatalog #28908
Dynabeads MyOne Silane10x GenomicsCatalog #2000048
Sodium Chloride Solution 5 M Merck MilliporeSigma (Sigma-Aldrich)Catalog #59222C
RPMI 1640 Medium, GlutaMAX™ SupplementThermo FisherCatalog #61870010
FBS Standard, South America origin, fetal bovine serum, 0.2 µm sterile filteredPAN BiotechCatalog #P30-3306
Tris-HCl (ph7.4)Merck MilliporeSigma (Sigma-Aldrich)Catalog #T2194-100ML
GlycineMerck MilliporeSigma (Sigma-Aldrich)Catalog #50046
Penicillin-Streptomycin, 10.000 U/ml Penicillin, 10 mg/ml StreptomycinPAN BiotechCatalog #P06-07100
DPBS, no calcium, no magnesiumThermo FisherCatalog #14190094
MgCl2Merck MilliporeSigma (Sigma-Aldrich)Catalog #M1028-100mL
BSA Stock SolutionMiltenyi BiotecCatalog #130-091-376
Troubleshooting
Safety warnings
Please be careful when handling (potentially) infectious biological material. Stick to local regulations when working with biohazards.
This protocol includes the use of the following hazardous goods. Check safety instructions and hazardous goods sheets before starting to work with them (see the "Materials" section for links to the manufacturer websites):
- Paraformaldehyde
- Ethanol
- Penicillin/Streptomycin
- Dithiothreitol (DTT)
- NP-40
Ethics statement
Please follow all the ethical regulations for the use of human donor and patient samples. It is the responsibility of the user to ensure that all legal obligations are followed.
Before start
It is highly recommended to follow the checklist outlined below before starting with the experiment:
- Book tissue culture hood, FACS sorter, centrifuges, Chromium X/iX, and PCR cyclers in advance
- Check the availability of all materials and equipment as outlined in the "Materials" section of this protocol
- At least one day before starting with sample handling, the buffers & reagents as listed in step 2.1 of this protocol should be prepared. They can be stored for multiple weeks to months at indicated temperatures.
- The buffers as listed in steps 2.2-2.5 must be prepared freshly on the same day when the experiment is conducted.
- 80% ethanol must be prepared freshly for steps 4-8.
Step 0: Preparations of materials and equipment
55m
Prepare cell thawing
- Cool down a swinging-bucket centrifuge to 4 °C .
- Have dry ice ready.
- Pre-warm thawing media to 37 °C .
15m
Prepare buffers
General buffers and media
- The buffers and media in this step can be stored for several weeks at 4 °C .
1) Prepare cell thawing media: RPMI + 10% FBS + 1% P/S
- Requires: FBS Standard, South America origin, fetal bovine serum, 0.2 µm sterile filteredPAN BiotechCatalog #P30-3306 RPMI 1640 Medium, GlutaMAX™ SupplementThermo FisherCatalog #61870010
- Optional: Penicillin-Streptomycin, 10.000 U/ml Penicillin, 10 mg/ml StreptomycinPAN BiotechCatalog #P06-07100
- Add 50 mL FBS (and optionally 5 mL P/S) to 500 mL RPMI
- 50 mL per sample required
- Filter media through a 0.2 µm filter
- Store at 4 °C
2) Prepare recovery buffer for FACS: PBS + 0.04% BSA (PBS/BSA)
- Requires: BSA Stock SolutionMiltenyi BiotecCatalog #130-091-376 DPBS, no calcium, no magnesiumThermo FisherCatalog #14190094
- Add 2 mL 10 % volume BSA to 500 mL PBS
- 5-7 mL per sample required
- Store at 4 °C
3) Prepare 2X ATAC-Resuspension buffer (2X ATAC-RSB)
- Requires: Nuclease-free Water Tris-HCl (ph7.4)Merck MilliporeSigma (Sigma-Aldrich)Catalog #T2194-100ML Sodium Chloride Solution 5 M Merck MilliporeSigma (Sigma-Aldrich)Catalog #59222C MgCl2Merck MilliporeSigma (Sigma-Aldrich)Catalog #M1028-100mL
- Add per 116.46 mL Nuclease-free water: 720 µL 1 Molarity (M) MgCl2, 2399 µL 1 Molarity (M) Tris-HCl and 479 µL 5 Molarity (M) NaCl
- 620 µL per sample required
- Store at 4 °C
4) Prepare 2.5 M Glycine stock
- 2.5 Molarity (M) glycine (75.07 g/mol)
- Requires 1X PBS and GlycineMerck MilliporeSigma (Sigma-Aldrich)Catalog #50046
- Dissolve 4.7 g glycine in 25 mL 1X PBS
- Filtrate sterile (0.2 µm)
- 25 µL per sample required
- Store at Room temperature
Optional (in case a new Template Switch Oligo (TSO) aliquot must be prepared):
5) Prepare Template Switch Oligo (TSO)
Centrifuge a new tube of TSO (in the GEM Kit A, stored at -20 °C ) briefly, resuspend in 80 µL Low TE Buffer. Vortex 00:00:15 at maximum speed, centrifuge briefly, leave at room temperature for a minimum of 00:30:00 . After resuspension, store at -80 °C .
20m
PBS/BSA/RI (PBR) buffer
(Prepare freshly, keep on ice until use)
Requires:
- 1X PBS
- MACS BSA Stock SolutionMiltenyi BiotecCatalog # 130-091-376
- Protector RNase Inhibitor (40 U/ul) 2000 unitsMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335399001
| Reagent | Stock concentration | Final concentration | per 1 sample + 10% (X * 1.1) | |
| PBS | - | - | 2460.7 µL | |
| BSA | 10% | 1% | 275 µL | |
| RNase inhibitor | 40 U/µl | 0.2 U/µl | 14.3 µL | |
| Total | 2750 µL |
Pipetting scheme for the PBR buffer (required for washing of cells after sorting)
5m
Lysis buffer (LLL buffer)
(Prepare freshly, keep on ice until use)
Requires:
- 2X ATAC-RSB (prepared in step 2.1)
- NP-40 10% Merck MilliporeSigma (Sigma-Aldrich)Catalog #11332473001
- MACS BSA Stock SolutionMiltenyi BiotecCatalog # 130-091-376
- DTT (100mM Solution)Thermo Fisher ScientificCatalog #707265ML
- Protector RNase Inhibitor (40 U/ul) 2000 unitsMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335399001
- Nuclease-free Water
| Reagent | Stock conc. | Final conc. | Per 1 sample + 10% (X * 1.1) | |
| 2x ATAC-RSB | 2x | 1x | 55 µL | |
| NP-40 | 10% | 0.1% | 1.1 µL | |
| BSA | 10% | 1% | 11 µL | |
| DTT | 100 mM | 1 mM | 1.1 µL | |
| RNase inhibitor | 40 U/µL | 1 U/µL | 2.75 µL | |
| Nuclease-free water | - | - | 39.05 µL | |
| Total | 110 µL |
Pipetting scheme for the lysis buffer (LLL), mild lysis to retrieve intact but permeabilized mitochondria
5m
Wash buffer (WB)
(Prepare freshly, keep on ice until use)
Requires:
- 2X ATAC-RSB (prepared in step 2.1)
- MACS BSA Stock SolutionMiltenyi BiotecCatalog # 130-091-376
- DTT (100mM Solution)Thermo Fisher ScientificCatalog #707265ML
- Protector RNase Inhibitor (40 U/ul) 2000 unitsMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335399001
- Nuclease-free Water
| Reagent | Stock conc. | Final conc. | Per 1 sample + 10% (X * 1.1) | |
| 2x ATAC-RSB | 2x | 1x | 550 µL | |
| BSA | 10% | 1% | 110 µL | |
| DTT | 100 mM | 1 mM | 11 µL | |
| RNase inhibitor | 40 U/µL | 1 U/µL | 13.75 µL | |
| Nuclease-free water | - | - | 415.25 µL | |
| Total | 1100 µL |
Pipetting scheme for the wash buffer, used to wash cells post-sorting.
5m
1x Nuclei buffer (1x NB)
(Prepare freshly, keep On ice until use)
Requires:
- 20X Nuclei Buffer (from the 10x Genomics Epi Multiome ATAC + Gene Expression Kit, 10x-Product number: 2000207, box "ATAC Kit A"
- Protector RNase Inhibitor (40 U/ul) 2000 unitsMerck MilliporeSigma (Sigma-Aldrich)Catalog #3335399001
- DTT (100mM Solution)Thermo Fisher ScientificCatalog #707265ML
- Nuclease-free Water
| Reagent | Stock conc. | Final conc. | Per 1 sample | |
| 20x Nuclei buffer | 20x | 1x | 2.5 µL | |
| RNase inhibitor | 40 U/µL | 1 U/µL | 1.3 µL | |
| DTT | 100 mM | 1 mM | 0.5 µL | |
| Nuclease-free water | - | - | 45.7 µL | |
| Total | 50 µL |
Pipetting scheme for the 1X Nuclei buffer, used for recovery of fixed, permeabilized cells for subsequent tagmentation and barcoding
5m
Step 1: Cell Preparation - Thawing, washing, staining, FACS
3h 5m 30s
Thawing of cryopreserved PBMCs from liquid nitrogen stocks.
For cryopreserved cells, retrieve the cryovial from liquid nitrogen and transport it on dry ice.
Quickly thaw in a water bath at 37 °C until only a small ice crystal remains. Remove from the water bath, spray with ethanol and transfer to a clean tissue culture hood.
3m
Carefully transfer the thawed cells from the cryovial to a 50 mL conical tube.
1m
Rinse the cryovial with 1 mL pre-warmed thawing media (RPMI + 10% FBS) and add it dropwise to the 50 mL conical tube while gently shaking the tube.
2m
Sequentially dilute cells in the 50 mL conical tube by incremental 1:1 volume additions of pre-warmed media at a speed of 1 mL/3-5 sec to the tube and swirl. Repeat for a total of 5 times with ~1 min pauses between additions.
7m
Centrifuge 400 x g, 4°C, 00:05:00
5m 30s
Carefully remove the supernatant and resuspend the cell pellet in 10 mL PBS or FACS buffer.
2m
Centrifuge 400 x g, 4°C, 00:05:00
6m
Carefully remove the supernatant and resuspend the cell pellet in 1 mL PBS or FACS buffer. Transfer the cell suspension to a new 15 mL tube and top up to a 5 or 10 mL total volume.
2m
Determine the cell concentration and viability using a
Equipment
Countess 3 FL Automated Cell Counter
NAME
Automated Cell Counter
TYPE
Thermofisher scientific
BRAND
AMQAF2000
SKU
LINK
or a hemocytometer. Write down cell counts and cell suspension volume.
3m
Centrifuge 400 x g, 4°C, 00:05:00
6m
Remove the supernatant without disrupting the cell pellet and resuspend in 1 mL PBS/BSA. Transfer to a 1.5 mL microcentrifuge tube. Rinse the 15 mL tube with 0.4 mL PBS/BSA and transfer the rinse to the 1.5 mL tube containing the cells.
2m
FACS staining
Centrifuge 400 x g, 4°C, 00:05:00 and remove the supernatant.
6m
Resuspend cells in 100 µL PBS/BSA (max. 5 million cells per 100 µL ).
1m
Add 2.5 µL Fc blocking reagent Human TruStain FcX™BioLegendCatalog #422301 (= 1:40, V:V dilution) and incubate for00:05:00 at 4 °C .
6m
Add 1 µL CD66b-PE (1 % (v/v) ), 1 µL CD14-FITC (1 % (v/v) ).
Note
Optional co-staining of CD66b and CD14 for FACS:
Fluorescence-activated cell sorting (FACS) ensures reducing the fraction of dead cells and excluding residual neutrophils from the sample (e.g., after Ficoll), as both can negatively impact sequencing data quality. To remove neutrophils, the exclusion of CD66b+ cells is sufficient; however, because monocytes may express basal levels of CD66b, co-staining with CD14 is recommended to accurately exclude CD66b+CD14- neutrophils while retaining all CD14+ monocytes.
1m
Incubate for 00:20:00 On ice .
20m
Wash 2x with 1 mL PBS/BSA (spin 400 x g, 4°C, 00:05:00 ).
15m
Resuspend in 400 µL PBS/BSA with 0.1 % (v/v) SYTOX™ Blue Dead Cell Stain, for flow cytometryThermo FisherCatalog #S34857 .
2m
Sorting
Pass the cell suspension through a 5 mL FACS Tube with Cell Strainer Snap Cap to generate a single cell suspension right before sorting.
5m
Sort live CD66b- cells into a 15 mL conical tube with 5 mL On ice PBS + 0.04% BSA (prepared in step 1.2 (FACS recovery buffer) of the preparations section "step 0").
On a BD FACS Aria III, the following settings are recommended:
- Collect cells into a cooled 15 mL tube, rinsed with protein-rich buffer (see above; e.g., FACS buffer or PBS + BSA)
- ND filter: ND 1.5 filter
- Nozzle: 100 µm
Note that those settings might be adapted if another FACS sorter is used.
Note
Optional in case of sensitive cells:
In case of sensitive cell types, the cells can also be sorted into media to reduce stress. To minimize adhesion of cells to the tube walls, the media should contain at least 0.4% BSA, FBS or any other protein source.
Expected result
Required cell number post-sort:
It is recommended to retrieve at least 500k viable cells (of the population of interest) post-sort to have sufficient starting material for fixation and lysis to end up with 40'000 cells to load onto the Chromium Chip.
1h 30m
Step 2: Fixation, Lysis/Permeabilization, and Tagmentation
2h 38m 45s
Fixation (~0.5hr)
Centrifuge the conical tubes with sorted cells for 400 x g, 4°C, 00:05:00 .
6m
Remove the supernatant without disrupting the pellet.
1m
Resuspend cells in 450 µL Room temperature PBS/BSA/RI (PBR buffer) (prepared in step 2.2 of the preparations stection ("Step 0").
1m
(Optional, in case of not having sorted the cells): Pass the cell suspension through a 5 mL FACS Tube with a Cell Strainer Snap Cap to generate a single cell suspension.
Transfer to a 1.5 mL tube and immediately proceed with cell fixation.
30s
Add 3 µL Pierce 16% Formaldehyde (w/v), Methanol-freeThermo Fisher ScientificCatalog #28908 on top of the cells in 450 µL PBS/BSA/RI, quickly invert the tube several times to fix the cells in 0.1% formaldehyde. Incubate for00:05:00 at room temperature, swirl and invert occasionally.
6m
Quench with glycine solution to a final concentration of 0.125 Molarity (M) . Add 25 µL 2.5 Molarity (M) glycine solution into the 453 µL fixed cells. Quickly invert the tube several times.
1m
Wash 2x with 1 mL On ice PBS/BSA/RI, mix by inverting the tube 5 times and centrifuge at 400 x g, 4°C, 00:05:00 .
14m
Remove the supernatant carefully and proceed with cell lysis/permeabilization.
Cell lysis, permeabilization, and intracellular staining
Add 100 µL ice-cold lysis buffer to the cell pellet, pipet gently up and down 3 times. Recommended cellular concentrations are 300-500k cells/100 µl.
1m
Incubate on ice for 00:03:00 (primary cells).
3m
Wash 1x by adding 1 mL of chilled wash buffer and mix by inverting the tube 3 times. (Do NOT mix by pipetting up & down!). Centrifuge at 500 x g, 4°C, 00:05:00 (not 400g!).
7m
Discard the supernatant gently without disrupting the pellet.
1m
Dilute cells in chilled 1x Diluted Nuclei buffer with a volume depending on the number of cells to be recovered after sequencing (see table below). Taking into account the expected loss of material following the lysis and wash steps, it is recommended to initially resuspend in a small volume of 10-20 µL.
Depending on how many cells you expect, an initial recommendation is to resuspend the cells in 12.5 µL 1x Nuclei buffer from which 2.5 µL are taken for cell counting.
| Number of cells recovered | Cell stock concentration (cells/µL) | |
| 10,000 | 3,230 - 8,060 |
The final number of cells recovered at the end of the experimental workflow/detected in the sequencing data depends on the number of cells loaded. The mtscMultiome ATAC + GEX protocol loads 5 µL suspension of tagmented single cells onto the Chromium chip. Loading maximum 8k cells/µL is recommended to obtain an optimal number of cells post-sequencing.
Note
Tip in case of poorly singulated cells:
In case cells aggregated during processing and didn't yield a well separated single cell suspension, filtering through a cell strainer snap cap (35 µm pore size) is recommended.
10m
Count an aliquot using Trypan Blue and a cell counter, e.g.
Equipment
Countess 3 FL Automated Cell Counter
NAME
Automated Cell Counter
TYPE
Thermofisher scientific
BRAND
AMQAF2000
SKU
LINK
If necessary, adjust the cell count by diluting the cells in 1X Nuclei Buffer. Typically, we target 8000 cells/µL .
10m
Proceed immediately with the tagmentation reaction.
Tagmentation
Prepare the Transposition Mix on ice. For multiple samples, prepare the Transposition Mix per sample in an individual PCR tube of an 8-tube strip. Pipette mix 10 times, centrifuge briefly and maintain on ice.
| Transposition Mix | B | 1x (µL) | |
| Total | 10.0 | ||
| ATAC Enzyme | 3.0 | ||
| ATAC buffer B | 7.0 |
Composition of the Tagmentation/Transposition Mix
2m
Gently mix the cell stock by pipetting it slowly up and down. Add the 5 µL of the cell stock to the tube containing the Transposition Mix. Gently pipette mix 6 times (pipette set to 10 μL). DO NOT centrifuge.
5m
Incubate in a thermal cycler using the following protocol:
Lid Temperature 50 °C , Reaction Volume 15 µL , Run Time 01:00:00 .
| Step | Temperature | Time | |
| Incubate | 37°C | 00:60:00 | |
| Hold | 4°C | Hold |
PCR cycler program for the tagmentation.
1h
Immediately proceed to the next step: 30 mins after the initiation of tagmentation, thaw reagents needed for GEM Generation and Barcoding.
Cut one tube of beads (Single Cell Multiome Gel Beads, PN. 2000261, -80°C) from the strip per sample, equilibrate the beads to room temperature. Do not keep them on ice!
Thaw the Template Switch Oligo (TSO) (stored at -80 °C ), Reducing Agent B, Barcoding Reagent Mix at room temperature (upon thawing, place On ice ).
Step 3: GEM Generation and Barcoding
2h 16m
Preparations
Prepare the GEM Generation Master Mix On ice . Pipette mix 10x and centrifuge briefly.
| GEM Generation Master Mix | Per 1 sample (X * 1.1) in µL | |
| Barcoding Reagent Mix | 54.45 | |
| Template Switch Oligo (TSO) | 1.21 | |
| Reducing Agent B | 2.09 | |
| Barcoding Enzyme Mix | 8.25 | |
| Total | 66 |
Pipetting scheme for the GEM Generation and Barcoding Master Mix
5m
Prepare chip loading
Place On ice :
- Samples after tagmentation
- Master Mix
- Pre-chilled 8-strip emulsion-safe PCR tubes (1 tube per sample).
Room temperature :
- Single Cell Multiome Gel Beads
- Chromium Next GEM Chip J and 10x Gasket
- Partitioning oil and 50% glycerol
- Reagent reservoir with divider
- Two lids of tip boxes (to cover the chip and reservoir while loading)
- 200 µL emulsion-safe Low-Retention Filter tip (e.g., Rainin Filter Tip)
- 1 Emulsion-safe PCR tubes for the remaining beads.
15m
Chip assembly, loading of the Chromium Controller, and transfer of GEMs
35m
- Follow the instructions as outlined in the 10x Genomics Epi Multiome ATAC + Gene Expression user guide, CG000338 Rev E: Step 2.1-2.4.
General remarks
- Use a Rainin pipette
- While pouring 50% glycerol and partitioning oil into the two compartments of the reagent reservoir, do not mix the two reagents!
- Before taking the chip and gasket, spread your gloves with ethanol to prevent static electricity and only touch the side of chip/gasket!
- Cover the chip and reagent reservoir with the lid of the tip box between steps.
- Put the gel bead holder onto the 10x Genomics vortex adaptor before loading your sample.
Assemble the Chromium Next GEM Chip J according to the manufacturer’s instructions.
Load the Chromium Next GEM Chip J according to the manufacturer’s instructions.
Run the Chromium Controller according to the manufacturer’s instructions.
Transfer GEMs from the chip to the PCR strip On ice according to the manufacturer’s instructions.
GEM Incubation
Incubate in a Bio-Rad C1000 Touch Thermal cycler with a 96-Deep Well Reaction Module with the following protocol:
Cycler settings:
- Lid temperature 50 °C
- Reaction volume 100 µL
- Total run time 01:15:00
- Program:
| Step | Temperature | Time | |
| 1 | 37°C | 00:45:00 | |
| 2 | 25°C | 00:30:00 | |
| 3 | 4°C | Hold (do not leave over night!) |
PCR cycler program for GEM incubation (RT-PCR)
Equipment
C1000 Touch™ Thermal Cycler with 96–Deep Well Reaction Module
NAME
Thermal Cycler
TYPE
BioRad
BRAND
1851197
SKU
LINK
1h 15m
Note: After GEM incubation, proceed immediately to Quench!
Retrieve the Quenching Agent (PN-2000269) from -20 °C and equilibrate to Room temperature while the PCR program is running.
Add 5 µL of the Quenching Agent to each sample to stop the reaction.
1m
Slowly pipette mix 10 times (pipette set to 90 µl). The solution will be viscous. Ensure that no liquid remains along the tube walls and pipette tips. If necessary, aspirate the entire volume and dispense slowly back into the tube.
5m
Store at -80 °C for up to 4 weeks, or proceed to the next step.
Step 4: Post GEM Incubation Cleanup
1h 11m
Follow the instructions as outlined in the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression user guide, CG000338 Rev F: Step 3.
Material and pre-prep
Thaw the Cleanup Buffer at 65 °C for 00:10:00 at maximum speed on a Thermomixer with shaking feature. Afterwards, cool to Room temperature .
10m
Thaw the Reducing Agent B to Room temperature (Upon thawing, place On ice ).
Prepare the Dynabeads Cleanup Mix
- Vortex
- Maintain at Room temperature .
- Requires Dynabeads MyOne Silane10x GenomicsCatalog #2000048 *
| Component | Per 1 sample + 10% (X * 1.1) | |
| Cleanup Buffer | 200.2 µL | |
| Dynabeads MyOne SILANE* | 14.3 µL | |
| Reducing Agent B | 5.5 µL | |
| Total | 220 µL |
Pipetting scheme for the Dynabeads Cleanup Mix.
* Store at 4 °C . Vortex thoroughly (≥30 sec) before adding the Dynabeads to the mix. Aspirate the full liquid volume with a pipette tip to verify that the beads have not settled in the bottom of the tube. If clumps are present, pipette mix to resuspend completely. DO NOT centrifuge before use.
5m
Prepare the Elution Solution I
- Add reagents in the order listet
- Vortex, spin down
- Maintain at Room temperature
| Component | Per 1 sample + 10% (X * 1.1) | |
| Buffer EB | 59.25 µL | |
| 10% Tween 20 | 0.605 µL | |
| Reducing Agent B | 0.605 µL | |
| Total | 60.5 µL |
Pipetting scheme for the Elution Solution I
5m
Prepare 80 % (v/v) ethanol (10 mL for 5 reactions) freshly.
2m
Dynabeads cleanup
Add 125 µL Recovery Agent to each sample at room temperature. DO NOT pipette mix or vortex the biphasic mixture!
1m
Gently invert the tube 10 times to mix. Keep the lid actively closed with the fingers to prevent unintentional leaking of the tube during inversion!
Centrifuge briefly.
1m
Slowly remove and discard 125 µL Recovery Agent/Partitioning Oil (pink) from the bottom of each tube without aspirating any aqueous sample.
2m
Vortex Dynabeads Cleanup Mix and add 200 µL to each sample. Pipette mix 5x with the pipette set to 200 µl. Do NOT close the cap of the PCR tube, otherwise the sample will spill out!
2m
Cover the PCR tube with the lid of a tip box and incubate for 00:10:00 at Room temperature
10m
At the end of 10 min incubation, place on the 10x Magnetic Separator, high position (magnet•High) until the solution clears.
2m
Discard the supernatant.
1m
While on the magnet, add 300 µL freshly prepared 80 % (v/v) ethanol to each sample without disrupting the pellet.
30s
Wait 00:00:30 and then remove the ethanol.
1m
While the sample is on the magnet, add 200 µL 80 % (v/v) ethanol to each sample without disrupting the pellet.
30s
Wait 00:00:30 and then remove the ethanol.
30s
Remove tubes from the magnet and centrifuge briefly. Place again on the magnet•Low.
1m
Remove any remaining ethanol.
30s
Remove the tube strip from the magnet. Immediately add 50 µL Elution Solution I to each sample.
1m
Pipette mix (pipette set to 30 µl) without introducing bubbles.
1m
Incubate 00:01:00 at room temperature.
1m
Centrifuge briefly. Place on the magnet•Low until the solution clears.
1m
Transfer 50 µL sample to a new tube strip and proceed to the SPRI cleanup as per protocol.
1m
SPRIselect cleanup
Vortex the SPRIselect reagent until fully resuspended. Add 90 µL (1.8X) SPRIselect reagent to each sample. Pipette thoroughly.
1m
Incubate 00:05:00 at Room temperature .
5m
Centrifuge briefly. Place on the magnet•High until the solution clears.
2m
Discard the supernatant.
1m
While keeping the tube(s) on the magnet•High, add 200 µL 80 % (v/v) ethanol to each sample without disrupting the pellet. Wait 00:00:30 and then remove the ethanol.
1m
Repeat the wash for a total of 2 washes.
3m
Centrifuge briefly and place on the magnet•Low.
1m
Remove any remaining ethanol.
1m
Remove the tube strip from the magnet and immediately add 46.5 µL Buffer EB.
1m
Pipette mix (pipette set to 45 μL) without introducing bubbles.
1m
Incubate for 00:02:00 at Room temperature .
2m
Centrifuge briefly and place on the magnet•Low until the solution clears.
1m
Transfer 46 µL sample to a new tube strip.
1m
Step 5: Pre-Amplification PCR & Clean-up
53m 30s
Material and pre-prep
Equilibrate the Pre-Amp Primers to Room temperature . Upon thawing, vortex and centrifuge briefly. Maintain On ice .
Thaw the Pre-Amp Mix on ice. Pipette mix and centrifuge briefly.
ATAC/GEX Pre-amplification PCR
(as outlined in the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression user guide, CG000338 Rev F: Step 4)
Prepare the Pre-Amplification Mix On ice . Pipette mix 10X and centrifuge briefly.
| Component | 10X product number (PN) | Per sample (µL) | |
| Pre-Amp Mix | 2000274 | 50 | |
| Pre-Amp Primers | 2000271 | 4 |
Pipetting scheme for the Pre-Amplification Mix.
3m
Add sample (46 µL from step 15.13) to the Pre-Amplification Mix. Pipette mix and centrifuge briefly.
2m
Incubate in a Bio-Rad C1000 Touch Thermal cycler with a 96-Deep Well Reaction Module with the following protocol.
- Lid temperature: 105 °C
- Reaction volume: 100 µL
- Total run time: 00:30:00
| Step | Temperature | Run time | |
| 1 | 72°C | 00:05:00 | |
| 2 | 98°C | 00:03:00 | |
| 3 | 98°C | 00:00:20 | |
| 4 | 63°C | 00:00:30 | |
| 5 | 72°C | 00:01:00 | |
| 6 | Go to step 3, repeat 6x (7 cycles total) | ||
| 7 | 4°C | Hold | |
PCR program for pre-amplification PCR
30m
[STOP POINT] Store at 4 °C for up to 18 h or proceed to the next step.
Post Pre-Amplification SPRI cleanup
! Important: Differences to the 10x Genomics protocol !
(Different volumes will be used for library elution and splitting.)
Vortex to resuspend SPRIselect reagent. Add 160 µL SPRIselect reagent to each sample. Pipette mix 15 times. (pipette set to 200 ul).
2m
Incubate 00:05:00 at room temperature.
5m
Centrifuge briefly. Place on the magnet•High until the solution clears.
2m
Discard the supernatant.
30s
While sample on the magnet, add 300 µL freshly prepared 80 % (v/v) ethanol without disrupting the pellet.
30s
Wait 00:00:30 and then remove the ethanol.
30s
While on the magnet, add 200 µL 80 % (v/v) ethanol to the sample without disrupting the pellet.
1m
Wait 30 sec and then remove the ethanol.
1m
Centrifuge briefly. Place on the magnet•Low.
30s
Remove any remaining ethanol.
30s
! Difference to the 10x Genomics protocol !
Remove the tube strip from the magnet. Immediately add 100.5 µL Buffer EB.
30s
Pipette mix (pipette set to 90 µl) without introducing bubbles.
1m
Incubate 00:02:00 at Room temperature .
2m
Centrifuge briefly. Place on the magnet•High until the solution clears.
1m
! Difference to the 10x Genomics protocol !
Transfer 100 µL sample to a new tube strip and proceed to library construction as per protocol. The 100 µL pre-amplified, SPRI-cleaned sample includes barcoded transposed DNA fragments and barcoded cDNA.
30s
! Difference to the 10X protocol !
The sample is divided and used as input for three separate steps:
- 25 µL sample (25%) is used for ATAC Library Construction (step 6)
- 35 µL sample (35%) is used for cDNA Amplification (step 7)
- Store the remaining Pre-amplification product at -20 °C for long term.
[STOP POINT] Store at 4 °C for up to 72 h or -20 °C for long-term storage.
Step 6: ATAC Library Construction
1h 16m
As outlined in the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression user guide, CG000338 Rev E: Step 5.
! Important: Differences to the 10x Genomics protocol !
Material and pre-prep
Equilibrate the Single Index Plate N set A to room temperature. Upon thawing, centrifuge at 1000 x g, 4°C, 00:01:00 . Maintain On ice .
Thaw the SI-PCR primer B On ice . Vortex and centrifuge briefly.
Thaw the Amp Mix On ice . Pipette mix and centrifuge briefly.
ATAC-Sample Index (SI) PCR
! Difference to the 10x Genomics protocol !
Prepare the Sample Index PCR Mix On ice .
Note: due to the reduced resuspension volume in the previous step, water has to be added to the mix to fill the input for the ATAC Index PCR mix ad 40 µL total volume (25 µL library + 15 µL water).
| Component | 10X product number (PN) | 1 sample (µL) | |
| Amp Mix | 2000047 | 50 | |
| SI-PCR Primer B | 2000128 | 7.5 | |
| Nuclease-free water | - | 15 | |
| Total | 72.5 |
Pipetting scheme for the Sample Index PCR Mix
4m
! Difference to the 10x Genomics protocol !
Add 25 µL pre-amplified sample (from 5.2.16.a) to 72.5 μL the Sample Index PCR Mix. Pipette mix and centrifuge briefly.
2m
Add 2.5 µL of an individual Single Index N Set A to each well. Record the barcode/well that was used for each sample. Pipette mix and centrifuge briefly.
5m
Incubate in a Bio-Rad cycler with 96-Deep Well Reaction Module with the following protocol.
(For mtscATAC/mtscMultiome ATAC library construction, we typically add 1-2 PCR cycles compared to what is recommended by 10x Genomics for their standard single-cell ATAC-seq protocol.)
- Lid temperature: 105 °C
- Reaction volume: 100 µL
- Run time: ~
| Step | Temperature | Time | |
| 1 | 98°C | 00:00:45 | |
| 2 | 98°C | 00:00:20 | |
| 3 | 67°C | 00:00:30 | |
| 4 | 72°C | 00:00:20 | |
| 5 | Go to step 2, see table below for number of cycles | ||
| 6 | 72°C | 00:01:00 | |
| 7 | 4°C | Hold | |
PCR protocol for the ATAC sample index PCR
| Number of loaded cells | Recommended total PCR cycles | |
| <2,000 | 10 | |
| 2,001 - 6,000 | 9 | |
| 6,001 - 10,000 | 8 |
Cycle number optimization table. The number of PCR cycles needs to be adapted based on the number of cells per µL loaded on the Chip J as determined in step 7.6 of this protocol (step 1.0 in the 10x Genomics protocol).
40m
[STOP POINT] Store at 4 °C for up to 72 h or proceed to the next step.
Post Sample Index Double Sided Size Selection – SPRIselect
Vortex to resuspend the SPRIselect reagent. Add 60 µL (0.6X) SPRIselect reagent to each sample. Pipette mix 15 times. (pipette set to 150 µl).
1m 30s
Incubate 00:05:00 at Room temperature .
5m
Place on the magnet•High until the solution clears.
1m
Transfer 150 µL supernatant to a new strip tube. DO NOT discard the supernatant.
1m
Vortex to resuspend the SPRIselect reagent. Add 95 µL SPRIselect reagent (1.55X) to each sample. Pipette mix 15 times. (Pipette set to 180 µl).
1m
Incubate 00:05:00 at Room temperature .
5m
Place on the magnet•High until the solution clears.
1m
Discard the supernatant.
30s
While the sample is on the magnet, add 300 µL freshly prepared 80 % (v/v) ethanol without disrupting the pellet.
30s
Wait 30 sec and then remove the ethanol.
1m
While the sample is on the magnet, add 200 µL 80 % (v/v) ethanol without disrupting the pellet.
1m
Wait 30 sec and then remove the ethanol.
1m
Centrifuge briefly and place on the magnet•Low.
30s
Remove remaining ethanol.
30s
Remove from the magnet and immediately add 20.5 µL Buffer EB. Pipette mix.
1m
Incubate 00:02:00 at Room temperature .
2m
Centrifuge briefly. Place on the magnet•Low until the solution clears.
1m
Transfer 20 µL sample to a new tube strip.
30s
[STOP POINT] Store at 4 °C for up to 72 h or -20 °C for long-term storage.
ATAC Post Library QC & Quantification
Follow the instruction as described in the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression user guide, CG000338 Rev F: Step 5.3.
Expected result
For the scATAC libraries from a mtscMultiome ATAC + GEX run, we typically expect concentrations between 40-100 ng/µL and molarities of 100-400 nanomolar (nM)
This is how the BioA profile of a typical scATAC library from a mtscMultiome run should look like:
Example of an expected DNA fragment size profile of an ATAC library from a mtscMultiome ATAC + GEX run recorded with a BioAnalyzer (Agilent)
Step 7: cDNA Amplification
3h 15m 30s
Follow the steps as outlined in Chromium Next GEM Single Cell Multiome ATAC + Gene Expression user guide, CG000338 Rev E: Step 6.
cDNA Amplification
For the cDNA amplification PCR, use the following PCR program and PCR cycles.
| Step | Temperature | Time | |
| 1 | 98°C | 00:03:00 | |
| 2 | 98°C | 00:00:15 | |
| 3 | 63°C | 00:00:20 | |
| 4 | 72°C | 00:01:00 | |
| 5 | Go to step 2, see table below for the number of total cycles | ||
| 6 | 72°C | 00:01:00 | |
| 7 | 4°C | Hold | |
PCR program for cDNA amplification.
| Nr. of loaded cells per µL | Total PCR cycles | |
| >2,0001 | 9 | |
| 2,001-6,000 | 7 | |
| 6,001-10,000 | 6 |
Cycle number optimization table. The number of PCR cycles needs to be adapted based on the number of cells per µL loaded on the Chip J as determined in step 7.6 of this protocol (step 1.0 of the 10x Genomics protocol). This table is identical to the table "Recommended starting point for cycle number optimization" in step 6.1 of the 10x Genomics protocol.
[STOP POINT]
After the PCR, amplified cDNAs can be stored at 4 °C for up to 72 h or at -20 °C for up to 7 days or proceed to the next step.
50m
cDNA Cleanup – SPRIselect
[STOP POINT] After the clean-up, the libraries can be stored at 4 °C for up to 72 h or at -20 °C for up to a month.
cDNA QC & Quantification
Follow the instructions as described in Chromium Next GEM Single Cell Multiome ATAC + Gene Expression user guide, CG000338 Rev F: Step 6.3. DNA concentration quantification via Qubit is not required, 1 µL undiluted amplified cDNA can directly be used for the determination of peak sizes & distribution via BioAnalyzer. However, quantification via Qubit can is recommended to obtain a quick estimation of yield and to ensure that no dilution is required before loading the sample onto the BioA chip.
2h
Step 8: GEX Library Construction
6h
Follow the instruction as described in Chromium Next GEM Single Cell Multiome ATAC + Gene Expression user guide, CG000338 Rev F: Step 7.
6h
Acknowledgements
We thank Paul Mendez and Moritz Kochanek for critically reviewing the draft document of this protocol.

