Mar 26, 2026
Most Probable Number (MPN) Method for Quantitatively Estimating the Colony Forming Units in the Fastidious Pathogen Piscirickettsia salmonis
- Pamela Aravena1,2,
- Javiera Ortiz Severín1,
- Daniel E. Palma1,2,
- Verónica Cambiazo1,2
- 1Universidad de Chile;
- 2Millennium Science Initiative Program – ICN2021_044

Protocol Citation: Pamela Aravena, Javiera Ortiz Severín, Daniel E. Palma, Verónica Cambiazo 2026. Most Probable Number (MPN) Method for Quantitatively Estimating the Colony Forming Units in the Fastidious Pathogen Piscirickettsia salmonis. protocols.io https://dx.doi.org/10.17504/protocols.io.261ge8zoog47/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: June 03, 2025
Last Modified: March 26, 2026
Protocol Integer ID: 219506
Keywords: Most probable Number, P. salmonis, viability analysis, numbers of bacteria, bacterial analytical manual, colony forming units in the fastidious pathogen piscirickettsia, probable number technique, classic microbiology technique, most probable number, probable number, bacteria, growth in bacteriological medium, fastidious fish pathogen piscirickettsia salmoni, culture in nutrient broth, bacterial suspension, number of replicate, fastidious pathogen piscirickettsia, bacteriological medium, probability statistics, number of viable cell, frequency of occurrence, using probability statistics, mpn number, growth in each dilution, colony forming unit, nutrient broth, mpn table, number
Funders Acknowledgements:
Fondecyt
Grant ID: 1211893
Millennium Science Initiative Program
Grant ID: ICN2021_044
Abstract
The Most Probable Number (MPN) method is a useful and classic microbiology technique that is based on the direct observation of the binary pattern of 'growth vs. no-growth' of a culture in nutrient broth serially diluted to extinction. After recording the number of replicates with growth or no-growth in each dilution, the number of viable cells in the original culture is obtained using probability statistics and relates to 'the frequency of occurrence of a series of positive results most likely to occur when given numbers of bacteria are present in a sample' (J. L. Oblinger and J. A. Koburger. 1975. Understanding and Teaching the Most Probable Number Technique, J. Milk Food Technol. Vol39, No. 9, p. 540-545). The MPN number can be obtained by referring to MPN tables for three- or five-replicate designs (such as the ones in the FDA’s Bacterial Analytical Manual), or by using the R Programming Platform (V. 0.3.0 MPN R package). Here, we have adapted this technique to estimate the number of viable cells in a bacterial suspension of the fastidious fish pathogen Piscirickettsia salmonis, obtained by growth in bacteriological medium (nutrient broth) or obtained after intracellular replication in a cell culture (infection).
Image Attribution
Pamela Aravena
Guidelines
We have observed a discrepancy between the number of P. salmonis cells in a culture (quantified using a counting chamber) and the number of colonies the same culture can yield in an agar medium. In addition, the colonies appear between 7 and 14 days after seeding. Considering this, and the fact that P. salmonis growth in nutrient broth is faster and more consistent over time, we adapted this classic technique to the requirements of P. salmonis growth to estimate the number of viable cells in a culture.
Materials
Tryptic Soy BrothMerck MilliporeCatalog #1.05459
NaClMerck MilliporeSigma (Sigma-Aldrich)Catalog #S5886
L-CysteineMerck MilliporeSigma (Sigma-Aldrich)Catalog #C7352
Iron(III) chloride hexahydrateMerck MilliporeSigma (Sigma-Aldrich)Catalog #44944
Leibovitz's L-15 Medium, powderThermo FisherCatalog #41300039
Fetal Bovine Serum qualified heat inactivated BrazilGibco - Thermo Fisher ScientificCatalog #10500064
Gentamicin SolutionMerck MilliporeSigma (Sigma-Aldrich)Catalog #G1397
Triton X-100Merck MilliporeSigma (Sigma-Aldrich)Catalog #X100-1L
Gibco™ DPBS (10X), calcium, magnesiumFisher ScientificCatalog #14080055
Troubleshooting
Safety warnings
It is recommended to evaluate the most probable number protocol when the infection occurs within the established time frame or with bacteria that grow as expected. If a P. salmonis culture do not reach OD600 = 1.2-1.5, subculturing and infecting with that culture may not work.
P. salmonis cultures in nutrient broth as inoculum for MPN assay
4d
Grow a culture of P. salmonis in an appropriate medium, such as NPB (NPB; 30 gr/L Tryptic Soy Broth, 256.6 mM NaCl, 8.25 mM L-cysteine, 37 µM FeCl3), by inoculating a 50 mL tube containing 5 mL NPB with bacteria from a solid NPA plate (NPB + 1.5% agar), or with bacteria from a grown NPB culture (OD~1.2-1.5).180 rpm, 18°C, 96:00:00
4d
Take an aliquot to perform serial dilutions (about 100 µL of each culture to be tested). And store at 4ºC or in ice until use.
P. salmonis recuperated from infections in SHK-1 cultures as inoculum for MPN assay
1w 4d
Infect SHK-1 cell cultures at 80% confluence in a T75 flask with stationary state bacterial cultures using MOI 100.
1w
After 3 days post-infection, wash the cultures with PBS and incubate 40 mins with Leibovitz's L15 medium with gentamicin (100 µg/mL) at 18 ºC.
After incubation, wash the cells with PBS and incubate at 18ºC with 15 mL of fresh Leibovitz's L15 medium supplemented with 5% of FBS, without antibiotics.
4d
Incubate for the required amount of time, here we show the results for 7 days after infection (dpi) at 18ºC.
Incubate the infected cell cultures with 1X cold PBS and lyse the monolayer by adding 4 mL of 1.5% Triton X-100 into the flask.
Incubate for 15 min at 4 °C.
Transfer the entire supernatant to a 50 mL sterile centrifuge tube, wash the flask twice with PBS, which also recovered in the same 50 mL tube.
Centrifuge the samples for 3 min at 1,000 x g, and recover the supernatant and centrifuge again at 8,000 x g for 15 min at 4°C.
Discard the supernatant and resuspend the pellet in NPB.
Place on ice or at 4ºC until use.
Plate design and inoculation for the MPN assay
1w 3d
Fill a 96-well round-bottom plate with 180 µl of NPB or the culture medium you wish to use to test P. salmonis replication.
Inoculate the first well (A1) with 20 µL of the grown P. salmonis culture or the recovered P. salmonis cells from infected SHK-1 cultures.
Homogenize the inoculated well by pipetting and serially dilute the culture 1/10 in each subsequent well, until the 11th well (A11), thus obtaining dilutions 10-1 to 10-11.
Use the same aliquot to make two or four more replicates (rows B-C or B-E) thus obtaining a triplicate or a quintuplicate for each bacterial culture.
Leave the 12th column of each row without inoculum as a negative control.
Incubate the plate at 18ºC without agitation for 7 - 10 days.
1w 3d
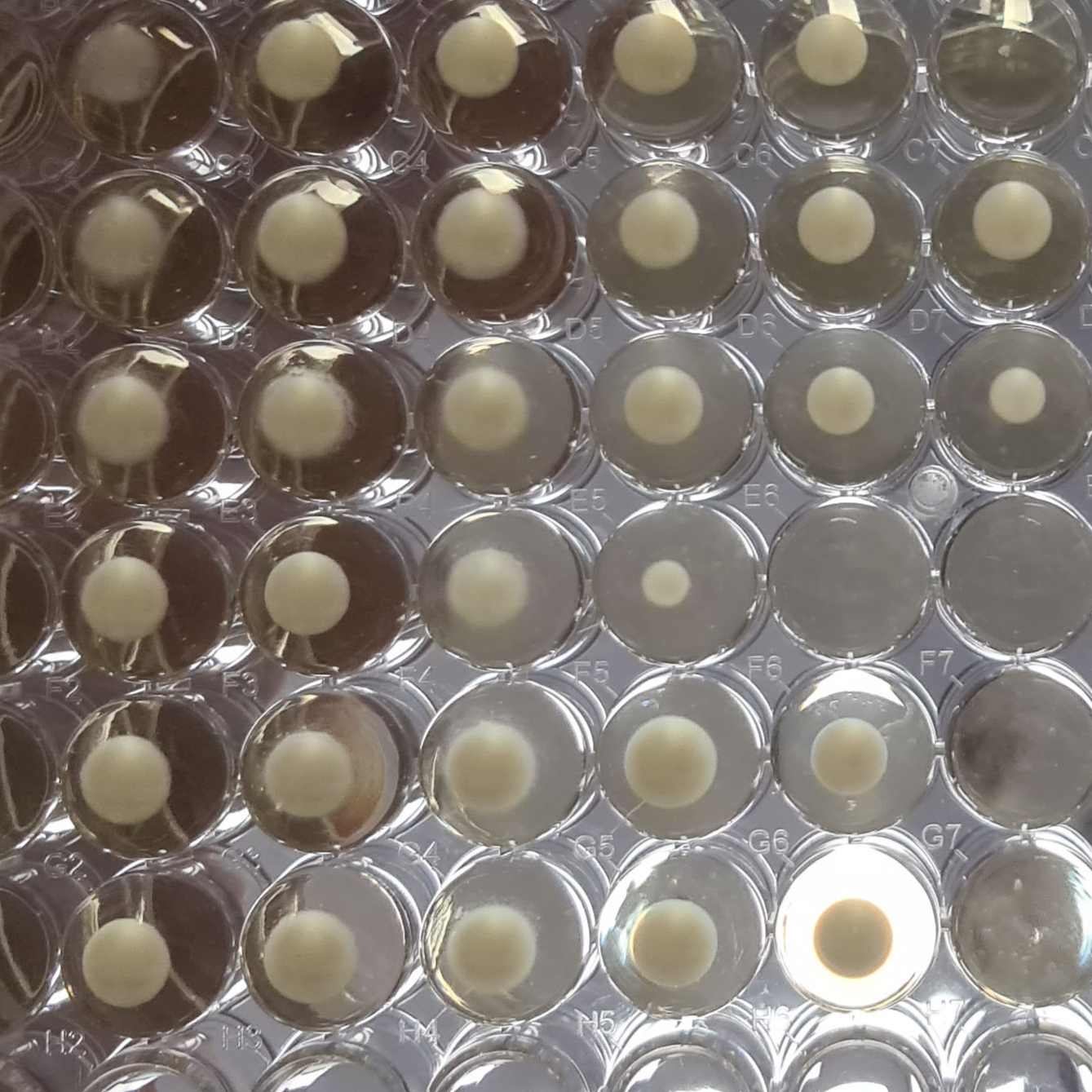
Record the number of wells with visible growth in every dilution for each replicate.
Calculate the MPN using https://mostprobablenumbercalculator.epa.gov/mpnForm or the MPN R package v0.3.0:
Install the required R packages:
Software
R Studio Desktop
NAME
The R Studio, Inc.
DEVELOPER
SOURCE LINK
install.packages(c("MPN", "data.table")).
Prepare your input table with the following format (as in Table 1): one column for dilution levels, one column for the number of replicates, and one column per sample, indicating the number of positive results at each dilution.
| A | B | C | D | E | |
| Dilution | Replicates | Sample 1 | Sample 2 | Sample 3 | |
| 1.00E-01 | 3 | 3 | 3 | 3 | |
| 1.00E-02 | 3 | 3 | 3 | 3 | |
| 1.00E-03 | 3 | 3 | 3 | 3 | |
| 1.00E-04 | 3 | 3 | 3 | 3 | |
| 1.00E-05 | 3 | 3 | 3 | 3 | |
| 1.00E-06 | 3 | 3 | 3 | 3 | |
| 1.00E-07 | 3 | 3 | 3 | 3 | |
| 1.00E-08 | 3 | 3 | 2 | 3 | |
| 1.00E-09 | 3 | 3 | 2 | 3 | |
| 1.00E-01-10 | 3 | 0 | 0 | 0 |
Table 1. Example of MPN input data.
Save file with the name 'MPN'. Export the table in txt format. Change the file extension to TSV (tab-separated values) format.
Use the following code to process the data and calculate the MPN.
library(data.table)
library(MPN)
# options ----
# sample names must
be unique
decimal_sep <- "."
input_file <- "MPN.tsv"
output_file <- "MPN_out.tsv"
conf_level <- 0.95
CI_method <- "Jarvis"
# data ----
data <- fread(input_file, dec=decimal_sep)
mdata <- melt(data, c("dilution", "replicates"), variable.name = "sample", value.name = "positive")
results <- mdata[, mpn(positive, replicates, dilution, conf_level =
conf_level, CI_method = CI_method), by=sample]
results[, std_dev := sqrt(variance)]
write.table(results, output_file, sep="\t", quote=FALSE, row.names=FALSE, dec=decimal_sep)
