Jun 20, 2025
High-throughput quantitative analysis of isoprenol (IP) by using an Agilent RapidFire-QqQ system

- Yan Chen1,
- David Carruthers1,
- Blake Simmons1,
- Taek Soon Lee1,
- Christopher Petzold1
- 1Lawrence Berkeley National Laboratory
- LBNL omics

Protocol Citation: Yan Chen, David Carruthers, Blake Simmons, Taek Soon Lee, Christopher Petzold 2025. High-throughput quantitative analysis of isoprenol (IP) by using an Agilent RapidFire-QqQ system. protocols.io https://dx.doi.org/10.17504/protocols.io.5jyl8d6w6g2w/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: January 08, 2025
Last Modified: June 20, 2025
Protocol Integer ID: 117828
Keywords: Metabolites, RapidFire, High-throughput, Quantitative, Isoprenol, throughput quantitative analysis of isoprenol, throughput measurement of isoprenol, isoprenol, analytical assay, such as microbial fermentation culture, microbial fermentation culture, agilent quantitative analysis software, throughput screening of thousands sample, qqq system this protocol details step, agilent 6460 qqq system, throughput quantitation of ip
Funders Acknowledgements:
Department of Defense (DOD) Tri-Service Biotechnology for a Resilient Supply Chain (T-BRSC)
Grant ID: AWD00007196
Department of Energy (DOE) Office of Science
Grant ID: DE-AC02-05CH11231
Disclaimer
This protocol is for research purposes only.
Abstract
This protocol details steps in high-throughput measurement of isoprenol (IP) with the Agilent RapidFire 400 coupled with Agilent 6460 QqQ system, and subsequent data processing using the Agilent Quantitative analysis software. This analytical assay provides accurate and high-throughput quantitation of IP from a broad variety of sample types, such as microbial fermentation cultures, chemical overlays, intracellular cell lysate, etc. The protocol provides flexible options for measurements of small number of samples or ultra high-throughput screening of thousands samples per day.
Materials
Acetonitrile LCMS qualityJT BakerCatalog #9829-02
Water LC-MS grade B&J BrandVWR International (Avantor)Catalog #BJLC365-2.5
Cis-3-Hexenyl AcetateMerck MilliporeSigma (Sigma-Aldrich)Catalog #W317101
3-METHYL-3-BUTEN-1-YL ACETATEMerck MilliporeSigma (Sigma-Aldrich)Catalog #S881171
- Bio-Rad hard shell 96-well PCR plate (Bio-Rad part number HSP9601) or other equivalent RapidFire compatible 96-Well plate.
- LC compatible non adhesive alumni seal (Agilent part number: 06644-001)
- PlateLoc Thermal Microplate Sealer or equivalent type
- BioMek FX or any equivalent liquid transfer automation platform
Equipment
RapidFire 400
NAME
High-throughput liquid chromatography
TYPE
Agilent
BRAND
G9532AA
SKU
LINK

Troubleshooting
Safety warnings
- Wear proper PPE (gloves, safety goggles, and lab coat), and prepare solvents in a chemical fume hood.
- Store organic solvents in a flammable solvent storage cabinet when not in use.
- Always prepare necessary control samples to validate sample preparation procedures.
- Dispose of used solvents and samples in accordance with your institute's Environmental, Health, and Safety requirements.
Before start
Refill RapidFire pumps with sufficient volumes of the following solvents to complete the analysis:
- LCMS grade water for pump 1.
- LCMS grade water for pump 2.
- LCMS grade methanol for pump 3.
Verify MS status:
- Perform Check-tune of QqQ system to ensure it passes calibration criteria. Service the QqQ and perform autotune if check-tune fails.
- Perform a quality control (QC) run to ensure QC compound peak full width at half maximum (FWHW), shape, and intensity are within quality standard range.
Isoprenol (IP) standard preparation
Prepare internal standard (IS) stock of Cis-3-Hexenyl Acetate (CAS: 3681-71-8) in methanol at 10 millimolar (mM) .
Prepare general diluent solution of 50 micromolar (µM) IS in 5 % volume Methanol (MeOH). This general diluent is used for diluting samples and preparing calibration standard diluent.
Prepare sample matrix that contains everything that is present in the typical sample except IP.
Note
A typical sample suitable for this protocol could be but not limited to microbial fermentation broth, IP trapping overlay such as dodecane, or extracted intracellular lysate of a microbial culture.
Prepare the calibration standard diluent (diluent used for generating calibration standards at a serial of concentrations) by spiking the same volume ratio of sample matrix into general diluent solution used to prepare test samples.
Note
Proper determination of sample dilution factor is important to make sure IP measurement is in calibration linear range.
Prepare IP standard stock at 100 millimolar (mM) in MeOH. Prepare the highest concentration IP calibration standard at 5 millimolar (mM) (430 mg/L) by diluting IP standard stock in calibration standard diluent.
Note
IP standard calibration range can be adjusted to fit estimated sample IP amount and chosen sample dilution factor.
Perform serial dilution of the highest concentration standard in calibration standard diluent by a dilution factor of 2 .
Leave one well containing Calibration standard diluent as Blank. At least 50 µL of blank and each standard are needed.
Sample preparation
Calculate the volumes of general diluent solution and test samples that are needed to prepare 100 µL Per sample with predetermined sample dilution factors. The following steps use dilution factor of 2 as an example.
Transfer 50 uL of general diluent solution (5% Methanol with the internal standard) to wells of Bio-Rad 96-well PCR plate that were designated for samples. Leave wells designated to Blank, QC, Calibration standards, and controls empty.
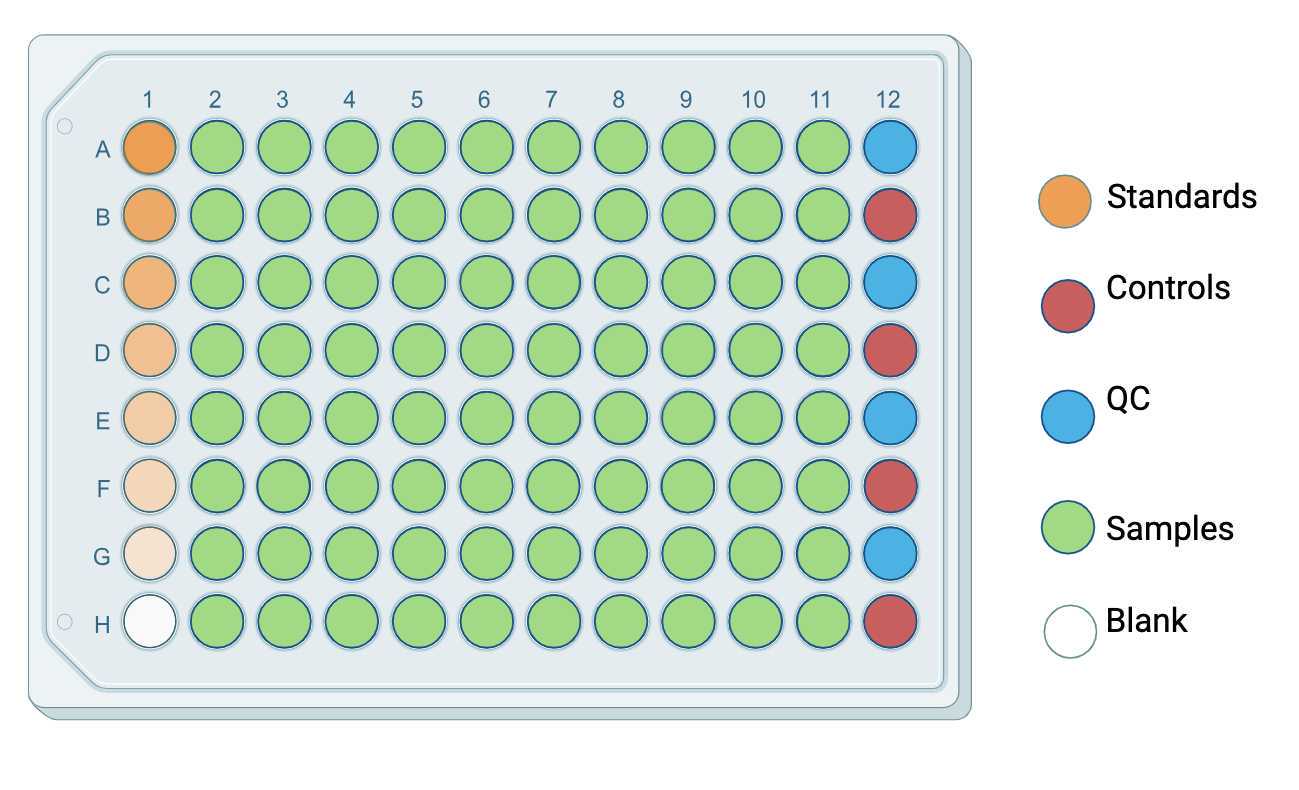
Example analytical plate layout. Calibration standards could be arranged together with samples in one plate, or be prepared in a separate plate. For every plate that contains test samples, controls and QC are required, but their well locations could be random.
Note
Using automated liquid handler reduces human error and increases accuracy and throughput, especially when multiple dilution factors are used and sample locations are randomized.
Transfer the test samples (at least 300 uL per well) to the sample well locations of 96 deep-well plates, then centrifuge at 3,200 rcf at4 °C for 30 minutes.
Note
The centrifugation time could be reduced when spinning at a higher centrifugation force. Alternatively, samples matrix could be removed through filtration process using commercially available filter plates, such as Agilent Captiva EMR-Lipid plate (Part Number:
5190-1001).
Transfer 50 uL of test samples from 96 deep-well sample plate, into the Bio-Rad 96-well PCR plate containing the general diluent solution (see Step 9).
Note
Using automated liquid handler reduces human error and increases accuracy and throughput, especially when multiple dilution factors are used and sample locations are randomized.
Finish preparing analysis plates by manually filling Blank, QC, Calibration standards and controls, then sealing the sample plate with LC compatible non adhesive alumni seal. Store the plates in -20 °C if they are not processed immediately.
Prepare RapidFire-MS system
Condition RapidFire pumps by purging pumps 1, 2, and 3 at 5 mL/min flow rate, and pump 4 at 10 RPM for 5 minutes.
Change the flow rates of pump 1, 2, and 3 to that of the running method, and change pump 4 speed to 3 RPM. Close the purge valves of pump 1, 2, and 3 afterwards.
| A | B | |
| RapidFire pump | Flow rate (mL/min) | |
| Pump 1 | 1.25 | |
| Pump 2 | 0.5 | |
| Pump 3 | 0.5 |
RapidFire pump flow rates for IP and internal standard quantitation
Check the back pressure of RapidFire high-throughput mass spectrometry system. Monitor all pump's pressure until they are stabilized at expected range.
| A | B | C | D | |
| RapidFire state | Pump 1 pressure (Mpa) | Pump 2 pressure (Mpa) | Pump 3 pressure (Mpa) | |
| Aspirate | 4.06 ~ 4.15 | 0.67 ~ 0.72 | 1.55 ~ 1.70 | |
| Load | 4.08 ~ 4.15 | 0.67 ~ 0.72 | 1.55 ~ 1.70 | |
| Extra Wash | 0.72 ~ 0.74 | 1.46 ~ 1.47 | 1.55 ~ 1.70 | |
| Elute | 0.72 ~ 0.74 | 0.72 ~ 0.79 | 2.20 ~ 2.23 |
RapidFire-QqQ system pumps back pressures at the method flow rates in each cycle state
Note
Pay close attention to the pressure of all the pumps. Make sure they are stable at the expected range before starting your run. Diagnose and service RapidFire if pump pressure exceeds normal range.
Load all sample plates in RapidFire stacker.
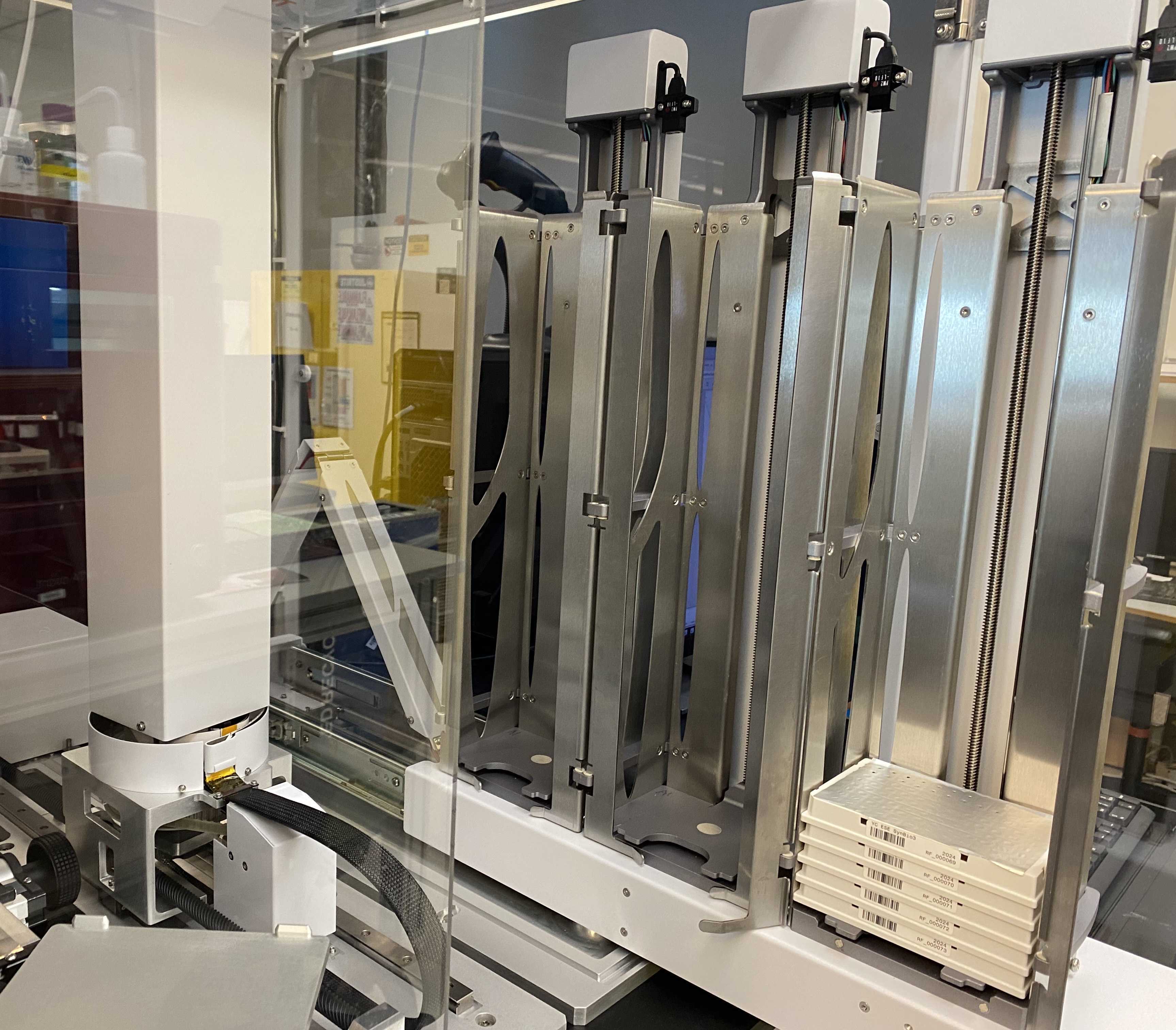

RapidFire plate stacker with loaded sample plates
RapidFire and QQQ method parameters
Create and save a RapidFire method using the following RapidFire Cycle parameters and pump flow rates:
| A | B | C | |
| State | Time (ms) | Flow rate (mL/min) | |
| Aspirate | 600 | ||
| Load | 3000 | 1.25 | |
| Extra Wash | 0 | 0.5 | |
| Elute | 5000 | 0.5 | |
| Reequilibrate | 500 | 1.25 |
RapidFire Cycle duration and flow rates for IP and internal standard quantitation
Install and condition RapidFire C18 4 µL Type C cartridge (Agilent Part Number: G9205A).
Note
Tracking sample counts on the working C18 cartridge and Install multiple ones if the number of total injection samples is greater than the maximum allowed injections per cartridge. Use the same cartridge type name if multiple C18 cartridges are installed.
Create and save a Agilent 6460 QqQ acquisition method that uses the following source parameters:
| A | B | |
| Gas Flow (L/min) | 10 | |
| Gas Temperature | 300 | |
| Vaporizer Temperature | 300 | |
| Nebulizer pressure | 30 | |
| Capillary Voltage | 3500 | |
| Corona Current (uA) | 4 |
Source parameters setting for IP and internal standard quantitation
Update the Agilent 6460 QqQ acquisition method with the following MRM acquisition parameters:
| A | B | C | D | E | F | |
| Compound name | Precursor (m/z) | Product (m/z) | Dwell time (ms) | CE | Polarity | |
| Cis-3-Hexenyl Acetate | 143.1 | 83.2 | 50 | 5 | Positive | |
| Isoprenol | 69.2 | 41.2 | 50 | 12 | Positive |
MRM acquisition parameters setting for IP and internal standard quantitation
Set up RapidFire-MS analysis
Create plate maps that contain the desired RapidFire injection sequences.
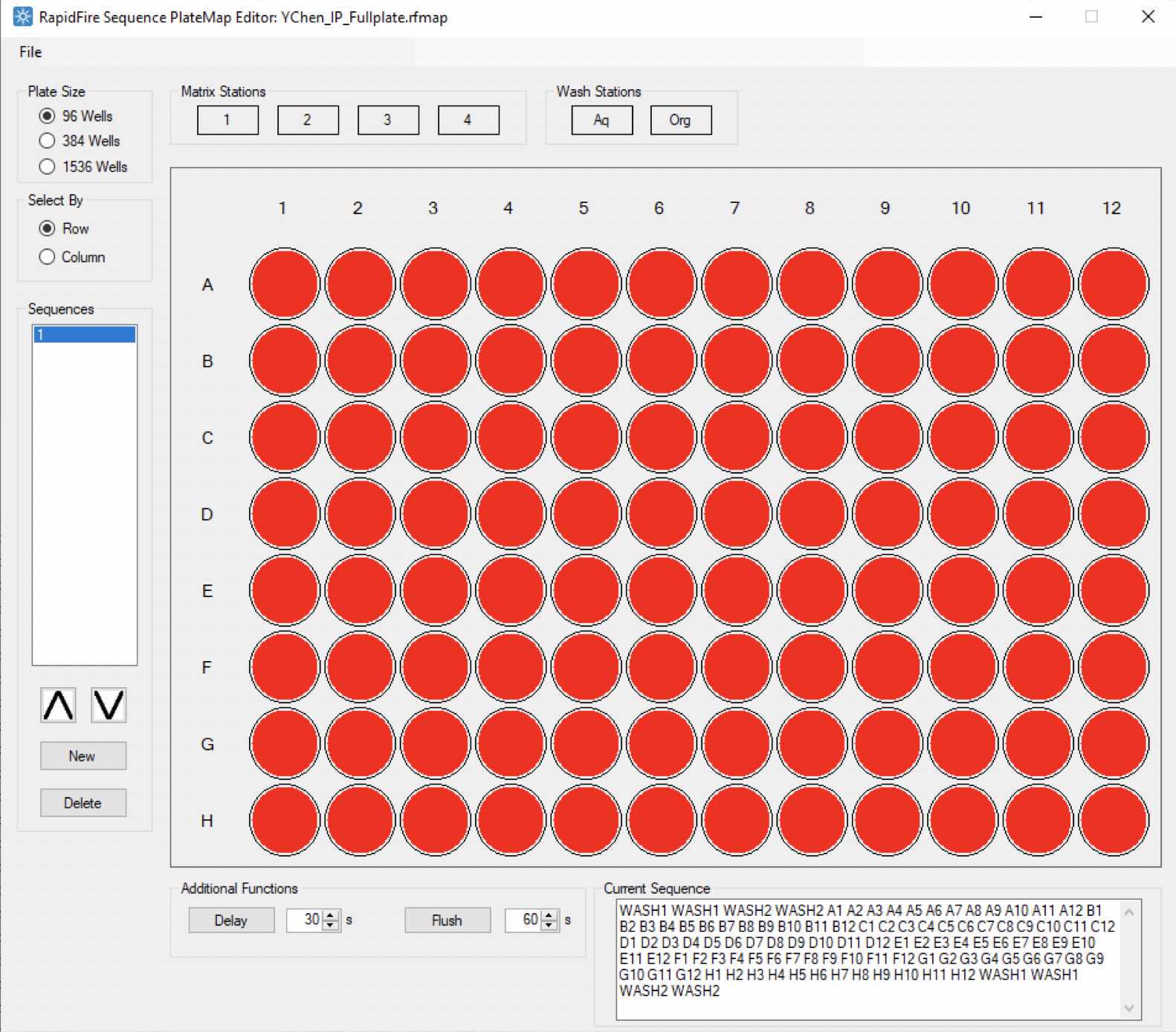
Example platemap that contains RapidFire injection sequence of a full 96-well sample plate
Create and save a batch file when the application runs single plate analysis or plates with multiple different injection sequences.

Example Batch file that contains plate info(Barcode etc) and their respective injection sequence, RapidFire method, Cartridge choice, and MS method.
Load the saved batch file under RapidFire UI File drop down menu. Click the green arrow button and follow the prompt to start the plateset.

RapidFire UI plateset run control panel
Create and save a Run Template file when the application runs high-throughput screen assay that analyze numerous full sample plates or plates with same injection sequence.

Example Run Template file that contains RapidFire injection sequence, RapidFire method, Cartridge choice, and MS method.
Create sample path file according to analytical sample plate layout. Fill in applied barcode information, well position, Sample name (File name), Sample type, assigned Calibration standard Levels and sample dilution factor(s).
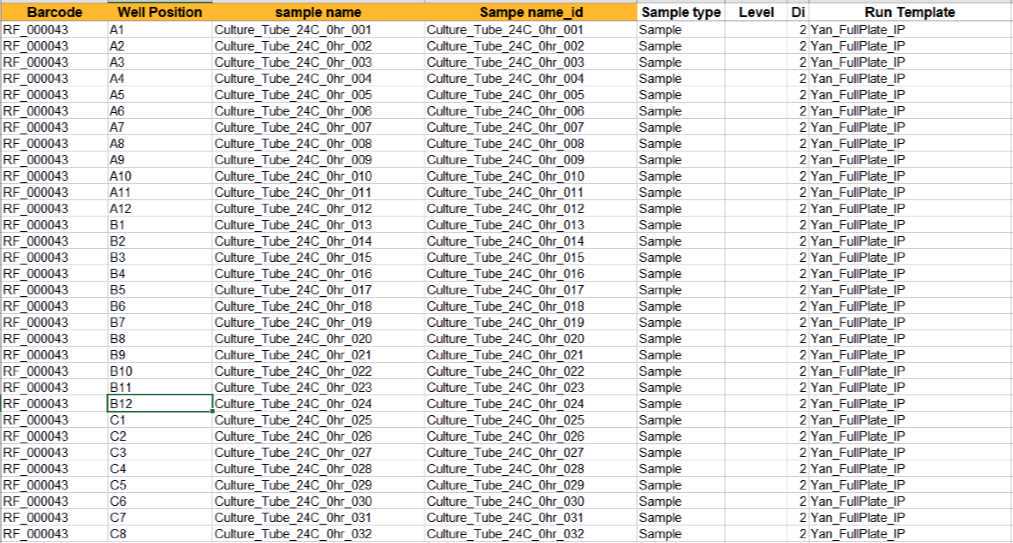
Example sample path file. Barcode column information needs to match applied barcode to corresponding plate. Well position, sample name, sample name_id and sample type are parsed from sample plate layout information. Calibration standard Levels are required for setting up quantitation method in Agilent QQQ quantitative analysis software. Dilution factor is a variable that could be modified accordingly. Run template file is generated and updated according to step 24.
Generate a Map File under RapidFire UI File drop down menu.
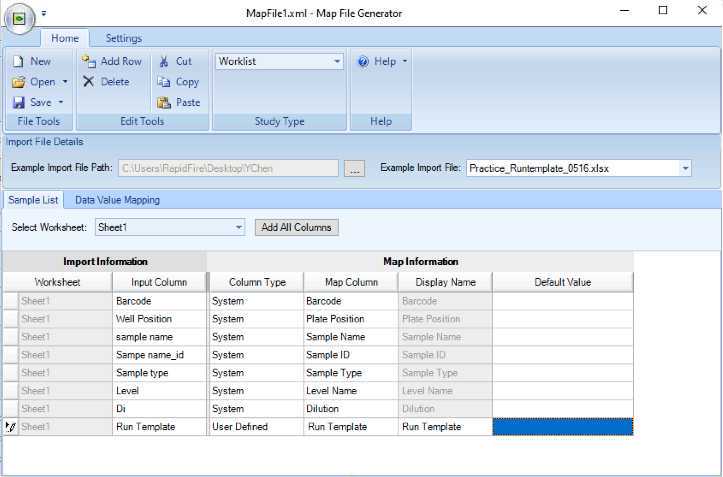
Example map file that parses information from sample path file to data file.
Import sample data file based on the map file generated above. Click import sample data under RapidFire UI file drop down menu. Select sample path file generated in step 25 and Map file generated in step 26 to fill in Sample Data File section.
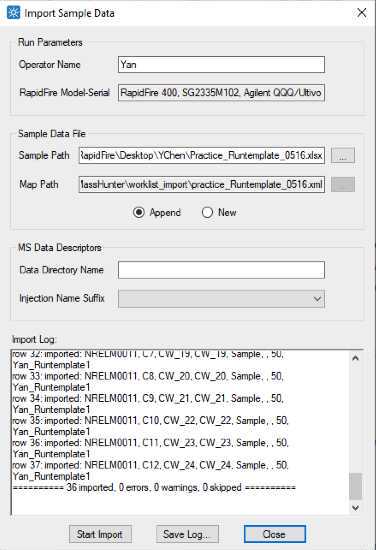
Example result after import sample data.
Note
Check the end of the import log to make sure that all samples are imported without errors, warnings and skipped samples.
Click the green arrow button and follow the prompt to start the plateset.
Data analysis
Convert RapidFire sequence data file(s) to individual injection data files.
Note
Sample data files could miss if 'Do not covert missed sips' option in the converting process is checked.
Create a new batch in Agilent MassHunter QQQ quantitative analysis software, and load all individual injection data files. If this is the first analysis of IP, create a new quantitative method from acquired IP standard data file.
Note
Other software such as the Agilent RapidFire Integrator could be also used for analyzing RapidFire MS data. Refer to each software user guide for details.
Validate and save the quantitative method after completing all tasks in the method setup sections. Apply the quantitative method and analyze the sample batch.
Note
The calibration standard levels and expected concentrations must match the levels defined in the sample batch table. Refer to MassHunter QQQ quantitative analysis software user manual for more details on setting up the quantitative method.

Example IP calibration curve that shows linear fitting ranging from 7.8 mg/L to 1000 mg/L.
Save sample batch and Export quantitative reports from batch table for further data processing and visualization.
Acknowledgements
This work was supported by the DOD Tri-Service Biotechnology for a Resilient Supply Chain (T-BRSC) program through Award AWD00007196 and by by the Joint BioEnergy Institute (http://www.jbei.org), U.S. Department of Energy, Office of Science, Biological and Environmental Research Program under Award DE-AC02-05CH1123 to Lawrence Berkeley National Laboratory.
Competing Interests:
BAS has a financial interest in Illium Technologies, Caribou Biofuels, and Erg Bio. None of the other authors have an outside financial interest to disclose.
