May 31, 2023
Helicase-like transcription factor (HLTF)-deleted CDX/TME model of colorectal cancer increased transcription of oxidative phosphorylation genes and diverted glycolysis to boost S-glutathionylation in intravascular metastatic niches
- Dalia Martinez-Marin1,2,
- Rebecca A. Helmer1,3,
- Gurvinder Kaur4,
- Rachel L. Washburn1,
- Raul Martinez-Zaguilan5,
- Souad R. Sennone1,
- Jannette M. Dufour1,6,
- Beverly S Chilton1,6,7
- 1Department of Cell Biology and Biochemistry, Texas Tech University Health Sciences Center, Lubbock, Texas, United States of America;
- 2Department of Immunology and Molecular Microbiology, Texas Tech University-Health Sciences Center, Lubbock, Texas, United States of America;
- 3Current address: Garrison Independent School District, Garrison, Texas, United States of America;
- 4Department of Medical Education, Texas Tech University Health Sciences Center, Lubbock, Texas, United States of America;
- 5Department of Cell Physiology and Molecular Biophysics, Texas Tech University Health Sciences Center, Lubbock, Texas, United States of America;
- 6Texas Center for Comparative Cancer Research, Texas Tech University School of Veterinary Medicine, Amarillo, Texas, United States of America;
- 7School of Medicine Cancer Center, Texas Tech University Health Sciences Center, Lubbock, Texas, United States of America

Protocol Citation: Dalia Martinez-Marin, Rebecca A. Helmer, Gurvinder Kaur, Rachel L. Washburn, Raul Martinez-Zaguilan, Souad R. Sennone, Jannette M. Dufour, Beverly S Chilton 2023. Helicase-like transcription factor (HLTF)-deleted CDX/TME model of colorectal cancer increased transcription of oxidative phosphorylation genes and diverted glycolysis to boost S-glutathionylation in intravascular metastatic niches. protocols.io https://dx.doi.org/10.17504/protocols.io.ewov1oe37lr2/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: May 31, 2023
Last Modified: May 31, 2023
Protocol Integer ID: 82645
Keywords: tme model of colorectal cancer, colorectal cancer, deletion in cancer cell, early stage colorectal cancer, transcription of oxidative phosphorylation gene, tumor microenvironment, hltf from tumor, cancer cell, oxidative phosphorylation gene, deleted cancer cell, metabolic pathway, prototypical example of cancer cell, hallmark of metastasis, oxidative phosphorylation, tumor cell, epigenetic silencing of hltf, deleted cancer, tumor suppressor, phenotype at the tumor border, putative role in altered metabolism, like transcription factor, reprogramming metabolism, glutathionylation in intravascular metastatic niche, metabolic requirements in response, altered metabolism, metastasis, redox homeostasis throughout the cdx, oxphos gene, master regulator of glycolysi, energy from glucose, negligible hltf expression in the tme, metabolic requirement, primary crc tumor
Funders Acknowledgements:
Harry Weitlauf Endowment for Cancer Research
TTUHSC Cancer Animal Facility Core, PI Dr. Scott L. Trasti This infrastructure grant provided funding for the ZEISS Axioscan 7 high-performance slide scanner
Grant ID: CPRIT Grant # RP190524
Abstract
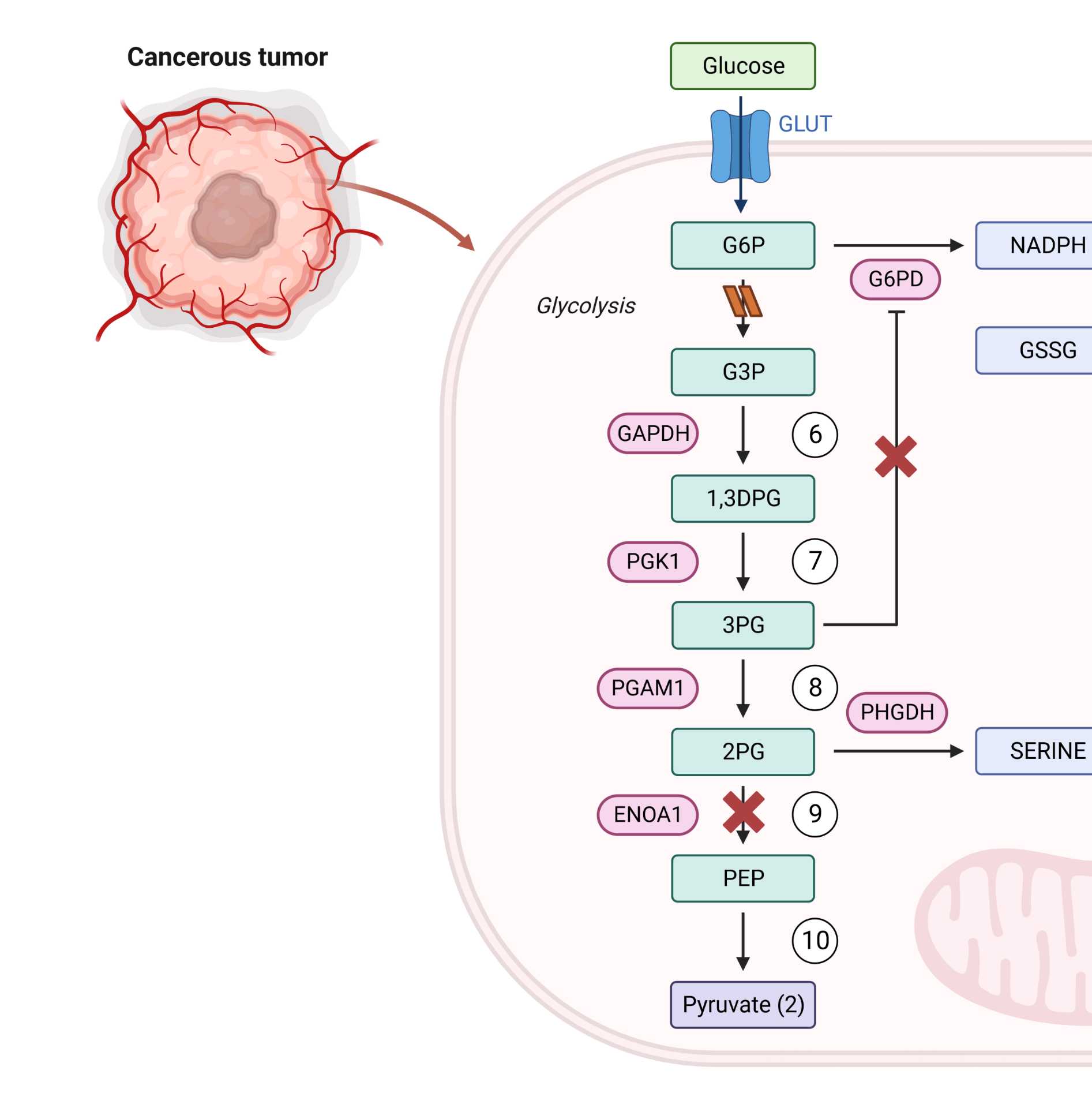
The tumor suppressor, helicase-like transcription factor (HLTF), is expressed in tumor cells but not in the tumor microenvironment (TME) in early stage colorectal cancer (CRC). With disease progression, epigenetic silencing of HLTF in primary CRC tumors coincides with negligible HLTF expression in the TME. Cell line-derived xenograft (CDX) models were developed to test the hypothesis that HLTF-deletion in cancer cells with resultant metabolic reprogramming—a hallmark of metastasis—is a prototypical example of cancer cells modifying their metabolic requirements in response to HLTF-deletion in the TME. The two metabolic pathways that derive energy from glucose—glycolysis and oxidative phosphorylation (OXPHOS)—are variously utilized by cancer cells depending upon the TME. HIF-1α, a master regulator of glycolysis, was eliminated from a role in reprogramming metabolism to satisfy CDX energetic requirements by RNAseq and spatial transcriptomics. Variability in the gut microbiome, with a putative role in altered metabolism, was also eliminated. In contrast, HLTF-deleted cancer cells recovered from DNA damage at a transcriptomic level induction of DNA repair and OXPHOS genes linked to an amoeboid-associated phenotype at the tumor border (confocal microscopy).HLTF-deleted cancer and endothelial cells of the intravascular niche in the TME share a site-specific protein S-glutathionylation signature (2D DIGE, MALDI-TOF/TOF mass spectrometry) for three glycolytic enzymes (PGK1 Cys379/380, PGAM1 Cys55, ENOA1 Cys119) that diverted glycolysis in support of continued glutathione biosynthesis. The collective absence of HLTF/Hltf from tumor and TME maintained redox homeostasis throughout the CDX and promoted metastasis.
Troubleshooting
Reagents and kits
Microbiome collection kits were obtained from TransnetYX (Cordova, TN). McCoy’s 5a Medium (30-2007) and fetal bovine serum (30-2020) were purchased from American Type Culture Collection (ATCC, Manassas, Virginia). Puromycin (ant-pr) was purchased from InvivoGen (San Diego, CA). XenoLight D-luciferin potassium salt (122799) was purchased from PerkinElmer (Waltham, MA). ThermoScientific (Waltham, MA) was the source of glutathione mouse monoclonal antibody (D8; MA1-7620) and the Pierce™ reagent HENS buffer (90106). PerkinElmer (Waltham, MA) was the source of Bioware® Brite Cell Line HCT116 Red-FLuc (BW124318). 10X Genomics (Pleasanton, CA) was the source of all Visium reagent kits for whole transcriptome profiling of intact formalin fixed paraffin embedded (FFPE) tissue sections including test slide (PN-1000347), slide kit (PN-1000188), reagent kit (PN-1000361), human transcriptome probe kit (PN-1000361), accessory kit (PN-1000194) and dual index kit TS Set A (PN-1000251). KAPA SYBR FAST qPCR master mix (KK4600) was purchased from Roche Diagnostic Corporation (Indianapolis, IN). SPRIselect (B23317) was purchased from Beckman Coulter Life Sciences (Indianapolis, IN). ProLong Gold antifade reagent with DAPI (P36935) and Streptavidin-FITC (19538-050) were from Invitrogen by ThermoFisher Scientific. 3-(N-Maleimidylpropionyl) biocytin (MPB) was from Santa Cruz Biotechnology (sc-216373). The following were purchased from Millipore Sigma: NADPH, tetrasodium salt (481973), Glutaredoxin-S2, E. coli (GRX1, 354406), Glutathione reductase (GSSG, G3664), and glutathione (GSH, Y0000517). Primary and secondary antibodies used with FFPE tissue sections in immunohistochemistry (IHC-P) and confocal immunocytochemistry (ICC/IF) are listed in Table 1.
Table 1. Source, application and concentration of antibodies
| Primary Antibodies | Secondary Antibodies | |
| Rabbit polyclonal HLTF antibody (NBP1-83256) Novus Biologicals (Centennial, CO) (1:100) Rabbit polyclonal CYTB (55090-1-AP) ThermoFisher Scientific (1:50) Rabbit polyclonal PGAM1 (16126-1-AP) ThermoFisher Scientific (1:50) Rabbit polyclonal ANXA1 (71-3400) ThermoFisher Scientific (1:20) Abcam (Cambridge, MA) rabbit monoclonal anti-γH2AX (phospho S139) ab81299 IHC-P (1:50) Abcam (Cambridge, MA) rabbit monoclonal ENO1 (ab227978) IHC-P (1:2,000) | IHC-P: Vector Laboratories RTU Biotinylated Goat Anti-rabbit IgG (H+L) (BP-9100-50) | |
| Mouse monoclonal PGAM1 (67470-1-1g) ThermoFisher Scientific (1:200) Rabbit polyclonal ANXA1 (71-3400) ThermoFisher Scientific (1 µg/mL) | ICC/IF: Goat anti-mouse, Alexa Fluor™ Plus 647 (A-32728) ThermoFisher Scientific (~4 µg/mL) ICC/IF: Donkey anti-rabbit, Alexa Fluor™ 555 (A-31572) ThermoFisher Scientific (~4 µg/mL) |
Gut Microbiome
Individual mice were allowed to defecate normally in autoclaved cages with no bedding. Using a sterile 26Gx1/2 needle (one per mouse), the first two fecal pellets per mouse were submerged in stabilization buffer in a barcoded sample collection tube. Collection tubes were shipped to TransnetYX for DNA extraction (Qiagen DNeasy 96 PowerSoil Pro QIAcube HT kit/protocol), library preparation (KAPA HyperPlus protocol), and sequencing. High molecular weight genomic DNA captured the true microbial diversity of stool samples from two cohorts: six- to eight-week old Hltf+/+ (n=9) and Hltf-/- (n=5) immune deficient (ID) male mice. Libraries were sequenced using Illumina NovaSeq at a dept of 2 million read pairs (2x150 configuration) sufficient for species and strain level taxonomic resolution. FASTQfiles were uploaded onto One Codex analysis software and analyzed against the One Codex database of ~127K whole microbial reference genomes. Classification results were filtered through several statistical post-processing steps to eliminate false positive results caused by contamination or sequencing artifacts. Mouse 12K metagenome-assembled genomes were included to screen out host reads. One Codex annotated metagenomes and generated a taxa plot of the top 10 species for all samples, and calculated alpha (within-sample) family diversity for comparison of the two cohorts.
HLTF-deleted isogenic HCT116 Red-FLuc cells
Bioware® Brite Cell Line HCT116 Red-FLuc is a luciferase-expressing derivative of a highly metastatic cell line (HCT116 parental line, CCL-247) that we used in the development of the CDX model. PerkinElmer’s recommendation that the Bioware® Brite Cell LineHCT116 Red-FLuc only be passaged 10 times, precluded the use of CRISPER/Cas9 to generate a stable knockout of HLTF. Thus, System Biosciences (Palo Alto, CA) produced a custom stable cell line with shRNA sequence targeting of HLTF. Briefly, synthesized HLTF shRNA sequence was cloned into pGreenPuro shRNA expression lenti vector (SI505A-1). HCT116 Red-FLuc HLTF-/- cells infected with 10 MOI of HLTF-shRNA-SI505 virus (lot#171009-008) yielded puromycin-resistant polyclonal HCT116 Red-FLuc HLTF-/- cells. For each CDX experiment, cell stocks in liquid nitrogen at passage 2 were thawed using T25 flasks, expanded in T150 flasks for 2 days, passaged overnight and harvested at 70-75% confluency. We established this protocol as a unified framework for comparative gene expression analysis.
Hltf-deleted mice
Immune competent (IC) global Hltf knockout (KO) mice were developed in collaboration with genOway (Lyon, France). Speed congenics accelerated the generation of IC Hltf KO C57BL/6J mice that were bred (IACUC# 02007) into the recombinase activating gene 2 (Rag2)/common gamma (IL2rg) double knockout background to generate ID Hltf KO mice >99% C57BL/6 background genome. ID Hltf KO mice were housed in a barrier facility with a 12:12 light/dark cycle, access to food and water ad libitum and bedding changes 2-3 times/week. Routine testing of sentinel mice ensured the colony was disease free. All studies and the anticipated mortality were conducted in accord with the NIH Guidelines for the Care and Use of Laboratory Animals, as reviewed and approved by the Institutional Animal Care and Use Committee at Texas Tech University Health Sciences Center (NIH Assurance of Compliance A3056-01; USDA Certification 74-R-0050, Customer 1481, S1 Checklist). TTUHSC's IACUC (# 02009) specifically approved this study.
The orthotopic HLTF-/-HCT116 xenograft model was established. For this study, briefly, six- to eight-week old ID Hltf KO mice (n=7) received direct orthotopic cell microinjections (OCMI) of HLTF-/-HCT116 Red-Fluc cells (2x106 cells/10 µl) between the mucosa and the muscularis layers of the cecal wall. Each surgery was performed with isoflurane (Isothesia) and the SomnoSuite® Low-Flow anesthesia system (Kent Scientific) with far infrared warming pads during surgery and recovery. Efforts to minimize suffering included an IP injection of Buprenorphine (Buprenex, 0.1 mg/kg) prior to surgery to manage incisional pain followed by a second dose 4-8 hours later, if needed. The cecum was exteriorized via small midline incision on the linea alba between the rectus abdominus muscles to eliminate bleeding. Non-invasive bioluminescence imaging (BLI) with an IVIS Spectrum In Vivo Imaging System was used to validate the quality and accuracy of the injection, and to track and quantify tumor growth and metastasis. Histopathology at necropsy confirmed placement of the inoculum. Mouse behavior and well-being were monitored daily. Tumor growth/metastasis was monitored weekly with BLI.
The humane endpoint of 28 days post CDX establishment in ID Hltf KO mice was determined in combinatorial experiments. Mice under continuous isothesia were imaged and killed immediately (<15 seconds) by cervical dislocation. Primary HLTF-/-CDX (hereafter HLTF-/-CDX) tumors were quickly removed, rinsed in physiological saline and either flash frozen for biochemical evaluation (RNAseq, Western blotting, MALDI-TOF/TOF MS, nanoLC-MS/MS) or formalin fixed paraffin embedded (FFPE) for routine histopathology, IHC-IP, laser scanning confocal ICC-IF and spatial transcriptomics.
Microscopy
FFPE HLTF-/-CDX tumors were serially sectioned (3-4 µm) for histopathology, IHC-IP and ICC-IF. Two sections were placed on each slide and deparaffinized prior to staining. Beginning with the first slide, sections on every fifth slide were stained with hematoxylin and eosin (H&E) for evaluation by light microscopy. Sections on alternate slides were processed for IHC-IP with heat-induced epitope retrieval (HIER). Two tissue sections per slide facilitated the use of one section for positive immunostaining, and the companion section for negative (minus primary antibody) control staining. Pairing of primary antibodies with secondary antibodies is shown in Table 1. Nuclei were counterstained (blue) with hematoxylin. ICC-IF (Table 1) was performed with serial sections from the above described FFPE tumors with HIER, aldehyde quench (50 mM NH4Cl in PBS), and ProLong Gold DAPI (DNA-specific blue-violet fluorescent dye).
Protein S-glutathionylation(Pr-SSG) was detected in situ in FFPE HLTF-/-CDX tumor sections (3-4 µm) with a biotin switch assay. Free thiol groups were blocked in HENS buffer (100mM HEPES, pH 7.8, 1 mM EDTA, 0.1 mM Neocuproine, 1% SDS) containing 20 mM MMTS (methyl methanoethiosulfonate) and 1% triton (vol/vol) for 10 minutes. After three rinses in PBS, S-glutathionylated cysteine groups were reduced by incubation with 27 µg/ml E coli GRX1, 4 U/ml GSSX, 1 mM GSH, 1 mM NADPH in 1 mm EDTA/10mM Tris pH 8 for 20 minutes. After three rinses in PBS, newly reduced cysteine residues were labeled with 1 mM MPB for 60 minutes.Excess MPB was removed with six rinses in PBS and tissue samples were incubated with 0.5 µg/ml streptavidin-FITC for 60 minutes. Excess streptavidin-FITC was removed with 6 rinses in PSB and cover slipped with ProLong Gold antifade reagent with DAPI to stain nuclei. Slides were allowed to cure overnight at room temperature and then stored at -20C until confocal imaging. Controls were treated the same except E. coli GRX1 was omitted from the deglutathionylation step.
Tumor transcriptome (RNAseq)
HLTF-/-CDX tumors (1 per ID Hltf KO mouse x 3 biological replicates) were flash frozen and sent to Genewiz, the next generation sequencing (NGS) division of Azenta Life Sciences (South Plainfield, NJ) RNA-sequencing. Briefly, total RNA was isolated, and evaluated for its integrity and purity with an Agilent Bioanalyzer (Table 2). RNA was polyA selected prior to library preparation followed by Illumina HiSeq, 2x150 configuration, single index per lane with 350 million raw paired-end (PE) reads per lane, i.e. 40-60 million PE reads per sample (>80% of bases >Q30). Sequence reads were trimmed to remove possible adapter sequences and nucleotides with poor quality using Trimmomatic v.0.36. The trimmed reads were mapped to either the Homo sapiens GRCh38 reference genome or the Mus musculus GRCm38 reference genome on ENSEMBL using the STAR aligner (v.2.5.2b) that detects splice junctions and incorporates them to help align the entire read sequences. BAM files were generated as a result of this step.
Unique gene hit counts were calculated by using featureCounts from the Subread package v.1.5.2. The hit counts were summarized and reported using the gene_id feature in the annotation file. Only unique reads that fell within exon regions were counted. After extraction, gene hit counts were used for downstream differential gene expression analysis (DESeq2) of HLTF-/-and HLTF+/+ CDX tumors from ID Hltf KOmice. Log2 fold changes (L2 FC) were calculated. P-values were obtained with the Wald test, and adjusted p-values (padj) were calculated with the Benjamini-Hochberg method. Genes with padj < 0.05 and absolute L2 FC >1 were called as differentially expressed genes for each comparison. New RNAseq data for Hltf-/-CDX tumors in ID Hltf KO are accessible through NCBI's Gene Expression Omnibus (GEO) Series. Our previously published RNAseq data for Hltf+/+CDX tumors in ID Hltf KO are accessible through NCBI’s Gene Expression Omnibus (GEO) Series accession number GSE161961.
Table 2. Sample quality control and RNAseq outcome.
| Sample ID | OD260/280 | RINa | DV200b | Yield (Mbases) | Total Reads | %bases >30c | |
| ID Hltf KO | 2.16 | 9.1 | 85.62 | 24,189 | 80,629,398 | 91.70 | |
| ID Hltf KO | 2.10 | 8.6 | 63.46 | 25,733 | 85,775,084 | 91.24 | |
| ID Hltf KO | 1.50 | 8.4 | 88.15 | 23,777 | 79,255,475 | 91.34 |
aRNA integrity number (RIN) from Agilent Bioanalyzer;
bPercentage RNA fragments >200 nucleotides (Tapestation)
cBase calling software assigns quality score (Q) to each base; Q30=accuracy of the base call is 99.9%
Spatial Transcriptomics
Work flow – five basic steps were necessary to implement spatial transcriptomics technology to quantify inter and intra tumor heterogeneity. Step 1, placement of FFPE CDX tumors on capture areas of a Visium gene expression (GEX) slide. Step 2, H&E staining followed by brightfield microscopic imaging with ZEISS Axioscan 7 high-performance slide scanner (White Plains, NY). Step 3, permeabilize tissue and construct barcoded libraries with a final sample index PCR according to the manufacturer’s instructions. Step 4, short-read sequencing (Illumina NovaSeq) of barcoded libraries by Genewiz. Step 5, data analysis of tissue images and sequencing files in FASTQ format with Space Ranger run on Ubuntu 22.04 LTS –Thelio Mira-b3 by System76, Inc. (Denver, CO). The space ranger aggr pipeline aggregated data from replicate samples and from replicate regions. Loupe browser visualization software was accessed in a desktop application via Windows (Dell Optiplex 990).
FFPE sections –CDX tumor sections (5 µm) were processed with the RNeasy FFPE kit for DV200 analysis. Replicate sections from either HLTF-/-or HLTF+/+ CDX tumors from ID Hltf KOmice were placed within fiducial frames of capture areas A,B and C,D respectively, on Visium GEX slide V11D13-089-A1.
GEX slide –each of 4 capture areas (6.5 x 6.5 mm each) inside fiducial frames (8 x 8 mm each) contain 5,000 gene expression spots (55 µm in diameter) with a distance of 100 µm between the centers of each spot (1-10 cells/spot). Visium for FFPE uses RNA-templated ligation (RTL) probes targeting the whole transcriptome. The assay captures probes via a capture sequence, e.g. poly-A for FFPE probes. Each gene expression spot has primers with a unique spatial barcode. Probes are designed against the entire human genome, each with primers that include Illumina TruSeq Read 1 (partial read 1 sequencing primer), 16 nt spatial barcode, 12 nt unique molecular identifier (UMI), and 30 nt poly(dT) sequence (captures ligation product). Spatially barcoded, ligated products were released from the slide, and harvested for qPCR with KAPA SYBR Fast qPCR master mix. The threshold for determining the Cq value for each sample was set along the exponential phase of the amplification plot at ~25% of the peak fluorescence value with QuantStudio 12 K Flex real-time PCR system (ThermoFisher Scientific). Sample index sets were selected to distinguish each of the 4 samples in a multiplexed sequencing run. Samples were amplified using Ilumina-compatible indexing primers, cleaned up with SPRIselect reagent, and bi-directionally sequenced.
Human Probe Set – Visium Human Transcriptome Probe Set v1.0 contains 19,144 gene ids targeted by 19,902 probes. Gene ids (1,201; 6.3%) targeted by 1,272 probes were excluded by default due to predicted off-target activity to a different gene. As a result, 17,943 gene_ids (targeted by 18,630 probes) were present in the final filtered output. During data analysis, read 2 sequences were mapped against the reference human genome (GRCh38/hg38) and read 1 sequences were used for UMI filtering to obtain spatial information.
Sequencing – Illumina NovaSeq NGS was performed at GenWiz. Unique dual indexing — unique identifiers on both ends of the sample — allows for an increase in the number of samples sequenced per run. Sequencing depth, a minimum of 50k read pairs per spot covered with tissue, was calculated by estimating the percent of capture area covered by the tissue section based upon the H&E brightfield image. Actual values are provided in Table 3.
Table 3. Statistics for spatial transcriptomics outcome for Visium_FFPE_HLTF-/-CDX in ID Hltf KOmice (A and B) and HLTF+/+CDX in ID Hltf KO (C and D).
| A | B | C | D | E | F | G | H | I | |
| ID | DV200 | # of spots under tissue | Mean reads/spot | Median genes/spot | # of reads | Validated barcodes | Sequencing saturation | Genes Detected | |
| A | 94.42 | 3,159 | 277,048 | 5,107 | 875,195,352 | 98.1% | 89.2% | 17,089 | |
| B | 94.42 | 2,757 | 208,484 | 2,629 | 574,790,269 | 98.0% | 93.3% | 15,883 | |
| C | 92.64 | 2,018 | 309,788 | 9,724 | 625,152,273 | 98.5% | 61.9% | 17,299 | |
| D | 92.64 | 2,206 | 341,062 | 9,362 | 752,382,625 | 98.4% | 73.5% | 16,386 |
Bioinformatics analysis
Visium Spatial Gene Expression Software Suite includes Space Ranger and Loupe Browser. Spaceranger mkfastq demultiplexed the Illumina sequencer’s base call files (BCLs) for each flow cell directory into FASTQ files. Spaceranger count combined a brightfield microscope slide image and FASTQ files from spaceranger mkfastq and performed alignment, tissue detection, fiducial detection, barcode/UMI counting, and prepared a full resolution slide image for visualization in Loupe Browser. The pipeline used the Visium spatial barcodes to generate feature-spot matrices, determine clusters, and perform gene expression analyses. The pipeline used a probe aligner algorithm for FFPE tissues. Spaceranger aggr used the output of multiple runs of spaceranger count from related samples and aggregated their input, normalizing those runs to the same sequencing depth, and then recomputed the feature-barcode matrices and the analysis on the combined data. The aggr pipeline combined data from multiple samples into an experiment-wide feature-barcode matrix and analysis. Loupe Browser was used to interrogate significant genes, characterize and refine gene clusters, and perform differential expression analyses.
Protein S-Glutathionylation (Pr-SSG)
Parallel analysis of Pr-SSG in HLTF-/-HCT116 Red-Fluc cells used for OCMI and resultantHLTF-/-CDX tumors in ID Hltf KOmice was performed by Applied Biomics, Inc (Hayward, CA). Briefly: cell pellet (control) and-CDX tumors (n=2 mice) were homogenized/sonicated (Polytron) in 4 volumes HENS buffer. Protein concentrations were determined (OD280) with NanoDrop One (ThermoFisher), adjusted to a final concentration of 2 µg/µl with HENS buffer and frozen for analysis.
Differential Pr-SSG between cell and tumor proteins was evaluated by large-format two-dimensional gel electrophoresis (Western blot) and two-dimensional difference gel electrophoresis (2D DIGE) in which cell (gel 1) and tumor proteins (gel 2) labeled with Cy2 (internal standard) were interrogated with mouse monoclonal anti-glutathione (0.5 µg/ml) and Cy5-labeled donkey anti-mouse IgG (1:2000 dilution). These two gels provided information on the difference in glutathionylation between cells and tumor (glutathionylation ratio). Difference in protein levels between the two samples (ratio) was determined by comparing Cy2 labeled cell proteins with Cy3 labeled tumor proteins by 2D DIGE (gel 3). The final glutathionylation ratio for each spot was obtained by adjusting the glutathionylation ratio by the protein ratio. 2D-DIGE guided the excision of 61 spots. Protein identification was based on peptide fingerprint mass mapping (using MS data) and peptide fragmentation mapping (using MS/MS data). MASCOT search engine was used to identify human and mouse proteins from primary sequence databases (SWISSProt). Posttranslational modification (glutathionylation) authentication was based on matrix-assisted laser-desorption ionization time-of-flight (MALDI-TOF/TOF).
