Oct 04, 2024
GoT-ChA: Genotyping of Targeted loci with single-cell Chromatin Accessibility
- Franco Izzo1,2,
- Robert M Myers1,2,
- Saravanan Ganesan1,2,
- Levan Mekerishvili1,2,
- Sanjay Kottapalli1,2,
- Tamara Prieto1,2,
- Elliot O Eton1,2,
- Theo Botella1,2,
- Ivan Raimondi1,2,
- Catherine Potenski1,2,
- Dan A Landau1,2
- 1Weill Cornell Medicine, New York, NY, USA.;
- 2New York Genome Center, New York, NY, USA.
- SMaHT

External link: https://github.com/landau-lab/Gotcha
Protocol Citation: Franco Izzo, Robert M Myers, Saravanan Ganesan, Levan Mekerishvili, Sanjay Kottapalli, Tamara Prieto, Elliot O Eton, Theo Botella, Ivan Raimondi, Catherine Potenski, Dan A Landau 2024. GoT-ChA: Genotyping of Targeted loci with single-cell Chromatin Accessibility. protocols.io https://dx.doi.org/10.17504/protocols.io.ewov19pn2lr2/v1
Manuscript citation:
Izzo, F., Myers, R.M., Ganesan, S. et al. Mapping genotypes to chromatin accessibility profiles in single cells. Nature 629, 1149–1157 (2024). https://doi.org/10.1038/s41586-024-07388-y
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: October 02, 2024
Last Modified: October 04, 2024
Protocol Integer ID: 108969
Keywords: cell chromatin accessibility somatic mutation, cell genotyping approach, cell sequencing approach, seq platform from 10x genomic, assaying chromatin accessibility, cell sequencing approaches in order, chromatin accessibility, impact of somatic mutation, somatic mutation, chromatin accessibility measurement, crucial for cancer initiation, 10x genomic, direct utilization of genomic dna, dependence on gene expression, mutation, genomic dna, genotyping information, end bias of most scrna, sequencing, aberrant clonal expansion, captured mrna transcript, mutation location, mrna transcripts as source, targeted loci, gene expression, cancer initiation, gene, mrna transcript, scrna, most scrna, genotyping
Abstract
Somatic mutations are crucial for cancer initiation and evolution, and have been identified across a number of healthy tissues in the human body. These mutations can disrupt normal cellular functions, leading to aberrant clonal expansions via acquired fitness advantages or skewed differentiation topologies. GoT-ChA and similar methods (e.g. GoT, TARGET-seq) aim to pair targeted genotyping with single-cell sequencing approaches in order to understand the impact of somatic mutations directly in human patient samples, in both malignant and non-malignant contexts.
GoT-ChA is unique in that it is a high-throughput single-cell method that pairs targeted genotyping with chromatin accessibility measurements based on the broadly utilized scATAC-seq platform from 10x Genomics. Previous single-cell genotyping approaches were largely based on scRNA-seq protocols, utilizing expressed and captured mRNA transcripts as sources for genotyping information. This results in a limiting dependence on gene expression and mutation location (due to 3' end bias of most scRNA-seq methods), precluding the usage of such technologies on lowly expressed or inconveniently located loci of interest. GoT-ChA surmounts these limitations via direct utilization of genomic DNA for genotyping, whilst simultaneously assaying chromatin accessibility.
Guidelines
How does GoT-ChA work?
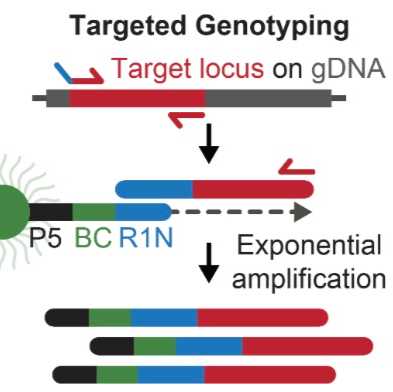
In order to capture genotypes within droplet-based scATAC-seq, two GoT-ChA primers are added into the cell barcoding PCR reaction that are designed to flank the locus of interest. One primer contains the partial Nextera Read 1N sequence in its handle, which allows for GoT-ChA amplicons to interact with the 10x Genomics gel bead oligonucleotide and obtain a unique cell barcode, just as tagmented chromatin fragments do. Further, the second GoT-ChA primer allows for exponential amplification of GoT-ChA amplicons while tagmented chromatin fragments are only linearly amplified.
After the single-cell emulsion is broken, a small portion of the sample is taken for GoT-ChA library construction, comprised of a hemi-nested PCR, biotin-streptavidin pull-down, and an on-bead sample indexing PCR. The final GoT-ChA library can be pooled with scATAC-seq libraries and sequenced together using standard ATAC sequencing parameters.
Troubleshooting
Before start
Designing a GoT-ChA experiment.
To utilize GoT-ChA, three primers need to be designed that flank the genomic region of interest. GoT-ChA_R1 (containing the partial Nextera Read1 sequence in its handle) and GoT-ChA_Rev flank the loci of interest, ideally forming an amplicon between 200-500 bps in size. GoT-ChA_nested is utilized in the hemi-nested PCR during GoT-ChA library construction, and crucially needs to bind within 50bp (inclusive of binding site) of the mutation to be covered with standard scATAC-seq sequencing parameters.
Buffer Preparations
Pre-chill swing-bucket centrifuge to 4˚C
Pre-heat block to 65˚C for later use dissolving digitonin
for GoT-ChA on Isolated Nuclei
Lysis Buffer
| A | B | C | D | E | |
| reagent | stock | final | µL for 1 mL (whole cell) | µL for 1 mL (Nuclei) | |
| Tris-HCl (pH 7.4) | 1 M | 10 mM | 10 | 10 | |
| NaCl | 5 M | 10 mM | 2 | 2 | |
| MgCl2 | 1 M | 3 mM | 3 | 3 | |
| NP-40 | 10% | 0.1% | 10 | 10 | |
| BSA | 10% | 1% | 100 | 100 | |
| Tween20 | 10% | 0.1% | - | 10ul | |
| Digitonin | 5% | 0.01% | - | 2ul | |
| Nucelase-free H2O | 875 | 863 |
prepare fresh, keep at 4˚C
Wash buffer
| A | B | C | D | E | |
| reagent | stock | final | µL for 1 mL | µL for 10 mL | |
| Tris-HCl (pH 7.4) | 1 M | 10 mM | 10 | 100 | |
| NaCl | 5 M | 10 mM | 2 | 20 | |
| MgCl2 | 1 M | 3 mM | 3 | 30 | |
| BSA | 10% | 1% | 100 | 1000 | |
| Tween 20 (for Nuclei prep only) | 10% | 0.1% | 10 | 100 | |
| Nuclease-free H20 | 875/865 | 8750/8650 |
prepare fresh, keep at 4˚C
PBS + 0.04% BSA
| A | B | C | D | |
| reagent | stock | final | µL for 1 mL | |
| BSA | 10% | 0.04% | 4 | |
| PBS | 996 |
GoT-ChA on Permeabilized Whole Cells
Obtain all single cell suspensions (filter if needed) and measure viability and density (if
viability is <90% proceed with live cell enrichment and/or use best judgement
depending on sample source / importance / rarity).
Resuspend cells in 450 μl PBS.
if low cell number, scale down accordingly.
Add 30 μl 16% formaldehyde (1% f.c) and incubate 10min at room temperature, swirl,
invert occasionally
Quench by adding glycine to 0.125M f.c.
Wash with 1x ice-cold PBS by filling up the tube, invert 5 times
Spin 5 minutes 400g at 4 ̊C.
Discard supernatant and repeat wash with 1ml 1x ice-cold PBS
Spin 5 minutes 400g at 4 ̊C, discard supernatant.
Resuspended cell pellet in 100 μl chilled lysis buffer, mix by pipetting.
Incubate on ice for 3min (primary cells), 5min (cell lines)
Add 1 ml chilled wash buffer to the lysed cells, mix by pipetting
Spin 5 minutes 500g at 4 ̊C.
Remove supernatant, resuspend in 150 μl 1x nuclei buffer (10x Genomics)
Filter through 40 μm strainers
Use 2 µL nuclei suspension mixed with 6 µL Diluted Nuclei Buffer and 2 µL trypan blue to determine nuclei concentration
Calculate amount of µL for target number of recovered cells
(max volume for loading is 5 µL for 10x transposition reaction)
Carry with Transposition for sc-experiment using 10x kit and follow the protocol. (remember to spike in 1µL of combined Forward and Reverse (each at 22.5µM conc) to the emulsion Master Mix)
corresponding to step 2.1 from 10X genomics ATAC user guide
GoT-ChA on Isolated Nuclei
Obtain all single cell suspensions (filter if needed) and measure viability and density (if
viability is <90% proceed with live cell enrichment and/or use best judgement
depending on sample source / importance / rarity).
Spin down at 300 x g for 5 minutes at 4˚C in PCR tubes if the number of cells are very low.
Remove supernatant and resuspend in 100 µL room temperature PBS + BSA
if low number of cells, scale done volume
Spin down at 300 x g for 5 minutes at 4˚C
Resuspend cell pellet in 100 µL chilled lysis buffer and mix via pipetting
Incubate on ice for 3 minutes (primary cells) or 5 minutes (cell lines)
Add 1 ml chilled wash buffer to the lysed cells and mix via pipetting
Spin 5 minutes at 700 x g at 4 ˚C
Remove supernatant in two steps to not touch pellet
Resuspend the pellet in X µL of chilled Diluted Nuclei Buffer for a final volume close to 7-8 µL
Use 2 µL nuclei suspension mixed with 6 µL Diluted Nuclei Buffer and 2 µL trypan blue to determine nuclei concentration
Calculate amount of µL for target number of recovered cells
(max volume for loading is 5 µL for 10x transposition reaction)
Carry with Transposition for sc-experiment using 10x kit and follow the protocol. (remember to spike in 1µL of combined Forward and Reverse (each at 22.5µM conc) to the emulsion Master Mix)
corresponding to step 2.1 from 10X genomics ATAC user guide
Emulsion generation
As per 10x protocol. (remember to spike in Gotcha primers (22.5uM - 1ul each) per reaction during emulsion generation step)
Step 2.5 from the 10X Genomics ATAC User guide
SPRI Clean up
Continue with SPRI clean up and elute DNA with 44.5ul of EB buffer. (40ul for ATAC and 4ul for GOTCHA)
Step 3.2 from the 10X Genomics ATAC User guide
Proceed to ATAC-library process as per 10x protocol with 40ul of eluted DNA.
Nested PCR
Perform 10-20 cycles of PCR with the following conditions (with nested primer)
GoTChA Nested PCR
| A | B | |
| KAPA reagents | 1X (in µL) | |
| DNA input | 20 ( 4 DNA+16 Water) | |
| p5 generic (10 µM) | 2.5 | |
| GoTChA nested primer (10 µM) Biotiylated | 2.5 | |
| KAPA (2X) | 25 | |
| total volume: | 50 |
GoTChA Nested PCR thermocycler program
| A | B | C | |
| 1 cycle | 3 min | 95˚C | |
| 10-20 cycles | 20 sec | 95˚C | |
| 30 sec | 65˚C | ||
| 1 min | 72˚C | ||
| 1 cycle | 2 min | 72˚C | |
| hold | 4˚C |
SPRI clean up with 1.2X. Elute in 40 µL.
Biotin- Pull Down
Allocate 15 µL of streptavidin M-280 beads per sample (+10%)
Wash beads three times with 100ul of 1x SSPE buffer
Resuspend in 5X SSPE buffer for 10 µL per sample (+10%)
Add 10 µL of beads to 40 µL of SPRI'd sample
Incubate at room temperature for 15 minutes
Wash two times with 100ul of 1x SSPE buffer
Wash one time with 100ul of 10 mM Tris HCl (pH 8.0)
Resuspend beads in 20 µL
On-bead Sample Indexing PCR
Perform 6-10 cycles of PCR with the following conditions:
GoTChA Sample Index PCR
| A | B | |
| KAPA reagents | 1X (in µL) | |
| DNA input | 20 | |
| p5 generic (10 µM) | 2.5 | |
| RPI-X (10 µM) | 2.5 | |
| KAPA (2X) | 25 | |
| amt to add: | 30 | |
| total volume: | 50 |
GoTChA Sample Index thermocycler program
| A | B | C | |
| 1 cycle | 3 min | 95˚C | |
| 10-20 cycles | 20 sec | 95˚C | |
| 30 sec | 65˚C | ||
| 1 min | 72˚C | ||
| 1 cycle | 2 min | 72˚C | |
| hold | 4˚C |
Run an e-gel to assess correct product formation. Use 5 µL (10%) of total sample
Purify libraries using AMPureXP beads at a 1.2X ratio
Elute in 20 µL of EB or Water
Assessing library quality
Run a BioA
Expected results:
Representative bioanalyzer traces of GoT-ChA genotyping (left) and GoT-ChA scATAC (rigth) libraries. FU, fluorescent units.
Measure samples via Qubit
Dilute samples for sequencing. GoT-ChA libraries can be sequenced together with ATAC libraries assuming i7 indexes do not clash. Sequencing parameters are as follows Read 1N - 50 cycles, i7 index -8 cycles, i5 Index - 16 cycles. Read 2N - 50 cycles.
Protocol references
Izzo F, Myers RM, Ganesan S, Mekerishvili L, Kottapalli S, Prieto T, Eton EO, Botella T, Dunbar AJ, Bowman RL, Sotelo J, Potenski C, Mimitou EP, Stahl M, El Ghaity-Beckley S, Arandela J, Raviram R, Choi DC, Hoffman R, Chaligné R, Abdel-Wahab O, Smibert P, Ghobrial IM, Scandura JM, Marcellino B, Levine RL, Landau DA. Mapping genotypes to chromatin accessibility profiles in single cells. Nature. 2024 May;629(8014):1149-1157. doi: 10.1038/s41586-024-07388-y. Epub 2024 May 8. PMID: 38720070; PMCID: PMC11139586.
