Sep 18, 2025
Genotyping Mice by PCR and Copy Number Standards
- Thom Saunders1,
- Honglai Zhang1,
- Jennifer Leo1,
- Sivakumar Jeyaraja1,
- Eden A. Dulka1,
- Zachary T. Freeman1
- 1University of Michigan
- Thom Saunders: Orcid ID: 0000-0003-2015-101X
- Sivakumar Jeyaraja: Orcid ID: 0000-0001-6614-5489
- Eden A. Dulka: Orcid ID: 0000-0002-7724-8923
- Zachary T. Freeman: Orcid ID: 0000-0003-1291-382X
- Transgenic Animal Model Core

External link: https://michmed.org/z45g9
Protocol Citation: Thom Saunders, Honglai Zhang, Jennifer Leo, Sivakumar Jeyaraja, Eden A. Dulka, Zachary T. Freeman 2025. Genotyping Mice by PCR and Copy Number Standards. protocols.io https://dx.doi.org/10.17504/protocols.io.8epv5krynv1b/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: August 29, 2025
Last Modified: September 18, 2025
Protocol Integer ID: 225753
Keywords: genomic mouse dna, sensitivity of the genotyping assay, experimental artifact pcr assay, genomic copy, genotyping mice, genotyping assay, mice by pcr, transgene dna, crispr founder, amount of transgene dna, transgenic founder, other genomic dna, inserted dna, copy number standards the purpose, pcr, important because transgenic founder, pcr screen, use of copy number standard, copy number standard, identification of transgene, insensitive assay, new primer pair, standardized copy, specific primer, transgene, copy standard, primer
Abstract
The purpose of this protocol is to describe the identification of transgenes in genomic mouse DNA. The use of copy number standards allows one to determine the sensitivity of the genotyping assay. This is important because transgenic founders or CRISPR founders are often mosaic and may carry less than one genomic copy of the inserted DNA when tail tip or other genomic DNA is assayed. PCR screens must be designed to detect transgene DNA at 0.1 copy level or lower. Insensitive assays are known to be the reason that transgenic founders were not detected on numerous occasions. To avoid this experimental artifact PCR assays need to be standardized copy standard template prepared by mixing non-transgenic tail DNA with a known amount of transgene DNA. When sensitive and specific primers are used then the occurrence of false negative results during genotyping can be eliminated. If primers do not meet sensitivity requirements or if they produce more than one band on a gel with copy standards then new primer pairs are ordered and tested until a suitable pair is identified.
Image Attribution
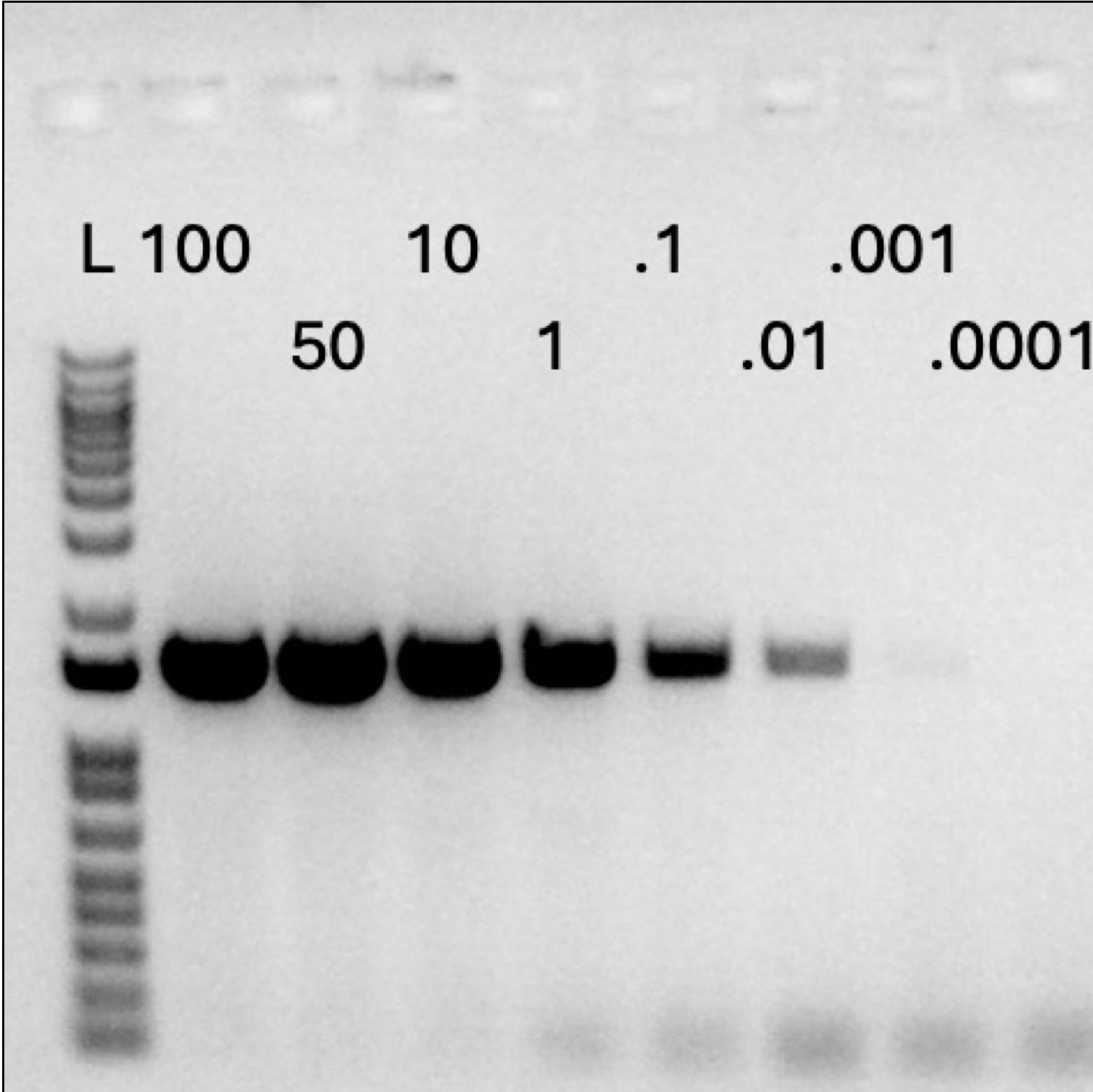
Copy sensitivity test with C468 KO spanning PCR primers on synthetic template spiked into wild type genomic DNA. Note that the C468 KO primers can detect as little as 0.01 copies of the transgene template when it is present in genomicDNA.
Lane 1: Invitrogen 1 Kb Plus Ladder.
Lane 2: 100 transgene copy standard.
Lane 3: 50 transgene copy standard.
Lane 4: 10 transgene copy standard.
Lane 5: 1.0 transgene copy standard.
Lane 6: 0,1 transgene copy standard.
Lane 7: 0.01 transgene copy standard.
Lane 8: 0.001 transgene copy standard.
Lane 9: 0.0001 transgene copy standard.
Lane 10: Wild type DNA (zero copy transgene).
Lane 11: Negative control. No template.
Lane 12: Negative control. No template.
Lane 13: Invitrogen 1 Kb Plus Ladder.
Image provided by Jennifer Leo.
Troubleshooting
Safety warnings
All activities involving animals require approval by an IACUC or other relevant regulatory body prior to executing this protocol.
Ethics statement
All animal activities described were reviewed and approved by the University of Michigan IACUC.
Procedure: Prepare Copy Number Standards
Assumption: the length of the inserted transgene DNA is 5480 bp.
Assumption: the Haploid content of a mammalian genome is 3 X 10E9 bp.
Assumption: 2 µg of mouse tail DNA are available.
Since the transgenic founder mice are hemizygous or mosaic calculate the amount of transgene DNA that would occur in 1 microgram of genomic DNA in a one copy mouse:
Divide the number of base pairs in the transgene by 3 X 10E9 bp or
5,480 bp ÷ 3 X 10E9 bp = 1.83 X 10E-7
For a mass of 1 microgram of DNA you need to add
1.83 X 10E-7 micrograms of transgene DNA to 2 micrograms of DNA to make a copy standard of one transgene copy per hemizygous genome.
Thus, for a 10 copy standard 18.3 pg transgene DNA is mixed with 2 µg tail DNA
Thus, for a 1 copy standard 1.83 pg transgene DNA is mixed with 2 µg tail DNA
Thus, for a 0.1 copy standard 0.183 pg transgene DNA is mixed with 2 µg tail DNA
Thus, for a 0.01 copy standard 0.0183 pg transgene DNA is mixed with 2 µg tail DNA
Suggestions for Primer Design
Primer design suggestions are taken from the Mouse Blastocyst DNA Extraction and PCR Amplification protocol.
DOI dx.doi.org/10.17504/protocols.io.5qpvodr5bg4o/v1
Procedure: Primer Design for Specific and Sensitive PCR Assays.
Use Primer-Blast to pick primers http://www.ncbi.nlm.nih.gov/tools/primer-blast
Adjust Primer Parameter default settings.
Minimum primer melting temperature: change to 60°C.
Optimal primer melting temperature: change to 63°C.
Maximum primer melting temperature: change to 66°C.
Minimum primer melting temperature difference: change to 1°C.
Adjust Primer Pair Specificity Checking Parameters
Click box to turn on “Enable search for primer pairs specific to the intended PCR template”
Set Search Mode to “Automatic”
Set Database to “Genomes for selected eukaryotic organisms (primary assembly only)”
Set Organism to “Mus musculus (taxid:10090)” for mouse.
Open Advanced Parameters
Change Primer Minimum size to 14
Change Primer Optimum size to 29
Change Primer Maximum size to 31.
nota bene. Stratman et al. reported primers of 27-30 nucleotides made up of 50-60% GC content will produce a 100-500 bp PCR product that uniformly detects genomic DNA with single copy sensitivity.
Stratman JL, Barnes WM, Simon TC. 2003. Universal PCR genotyping assay that achieves single copy sensitivity with any primer pair. Transgenic Res. 12:521-522.
Isolate Mouse Tail Tip DNA (or other DNA source)Untitled section
Prepare Reagent: TNES Buffer
| A | B | C | |
| Constituent | Quantity | Final Concentration | |
| 1 M Tris-HCl, pH 7.5 | 1.0 ml | 10 mM Tris, pH 7.5 | |
| 5 M NaCl | 12.5 ml | 400 mM NaCl | |
| 500 mM EDTA | 20.0 ml | 100 mM EDTA | |
| 10% SDS | 6.0 ml | 0.6% SDS | |
| Mol. Bio. Water | 60.5 ml | ||
| Total Volume | 100 ml |
Produces 100 ml of Buffer. Store at Room Temperature.
Prepare Reagent: Proteinase K Stock Solution.
| A | B | C | |
| Constituent | Quantity | Final Concentration | |
| Proteinase K | 10 mg | ||
| Mol. Bio. Water | 1 ml | 10 mg/ml | |
Produces 1 ml of 10 mg/ml Proteinase K. Store at -20°C.
Prepare Reagent: Saturated NaCl.
| A | B | C | |
| Constituent | Quantity | Final Concentration | |
| NaCl | 350.6 g | ||
| Mol. Biol. Water | 1000 ml | 6 molar | |
Store at 37°C to aid NaCl solubility, salt crystals will be apparent.
Obtain tail biopsies from 2- to 3-week-old mice in 1.5 ml microtubes per your approved animal use protocol. Hold mouse firmly at base of tail with one hand, with the other cut off 0.5 of the tail tip with a scalpel or single edge razor blade. Ear punches from older mice can be used in place of tail tips. Keep tissues frozen at -80°C if they will not be immediately processed.
Add 600 microliters of TNES and 35 microliters Proteinase K.
Incubate overnight (8-24 hr.) at 55°C.
Add 166.7 microliters 6M NaCl. Shake vigorously for 15 sec.
Remove supernatant to new 1.5 ml microtube and add 1 – 2 volumes cold 95% ethanol.
Insert closed end glass capillary into tube and twirl to spool DNA onto the glass capillary. If DNA does not spool then spin down in a microcentrifuge at top speed for 5 minutes and work with DNA pellet.
Rinse spooled DNA by dipping glass capillary in a tube filled with 70% EtOH and allow to air dry 5-10 min.
Place capillary with DNA into a microtube containing 100-500 µl of TE buffer (10mM Tris, pH 8.0, 1mM EDTA). TE volume depends empirically on pellet size.
Heat at 65°C for 10 minutes to aid dissolution of DNA.
Quantitate DNA and store at 4°C until needed.
Method adapted from: Miller SA. Dykes DD. Polesky HF. 1988. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Research. 16:1215.
PCR Tests of Primers for Sensitivity and Specificity.Untitled section
Use 200 ng of the transgene DNA copy standards DNA described above as a template in 25µl PCR reactions as described below.
Prepare Reagent: Prepare PCR Cocktail
| A | B | C | |
| Constituent | Quantity For 1 Reaction | Quantity For 20 Reactions | |
| Mol. Biol. Water | 14.0 µl | 280.0 µl | |
| 10X PCR Buffer | 2.5 µl | �50.0 µl | |
| 10 millimolar Nucleotides | 4.0 µl | 80.0 µl | |
| Primer A (10pmol/microliter) | 1.25 µl | 25.0 µl | |
| Primer B (10pmol/microliter) | 1.25 µl | 25.0 µl | |
| Taq Polymerase (0.3125 U/µl) | 1.0 µl | 20.0 µl | |
| Total Volume | 24.0 µl | 480.0 µl |
Aliquot 24.0 µl of cocktail into PCR tubes.
Add 1.0 microliter of DNA to each PCR tube.
nota bene: if tail DNA concentration is less than 200 ng/µl then add 200 ng of DNA in a volume up to 3 µl.
Final volume of PCR reaction is 25.0 µl.
Overlay with mineral oil and place into thermal cycler.
Amplify 30 to 35 cycles.
The first step in the cycle is denaturation for 30 seconds at 98°C.
Analyze on a 1 to 1.5% agarose gel in TBE, add 0.5 µg/ml of ethidium bromide to the gel prior to casting the gel. Visualize DNA on a UV transilluminator.
Remember to include a negative control of water only with PCR reagents.
Remember to include a DNA 0.1 copy standard as a positive control.
The first time primers are used they should be tested on a range of copy standards to demonstrate their sensitivity. Sensitive primers will detect as little as 0.001 transgene copies per genome.
If more than one band is produced discard the primers and order new primers.
If the primers can not detect 0.1 copies of the transgene in the copy standard discard the primers and order new.
For every sample run a parallel PCR with internal positive control primers that amplify genomic DNA from any mouse on the face of the earth. This will ensure that the DNA sample is "amplifiable" and free of PCR inhibiting contaminants.
For example, amplification of beta globin indicates that the tail DNA is free of PCR inhibitors. If it is positive for beta global and negative for the expected PCR product expected from sensitive and specific primers this indicates the animal is not transgenic.
Paralle PCR reactions or an internal positive control prevents false negative results, and positive animals will not be accidentally discarded as a consequence of PCR inhibitors in the DNA preparation.
Beta globin primers are used to amplify the endogenous mouse beta globin gene.
Amplification will thus demonstrate that the DNA sample is an “amplifiable” substrate for PCR.
5' CCA ATC TGC TCA CAC AGG ATA GAG AGG GCA GG 3' 32 mer
5' CCT TGA GGC TGT CCA AGT GAT TCA GGC CAT CG 3' 32 mer
Size of expected amplification product is 494 base pairs.
Thermal cycler profile:
Heat at 98°C for 60 seconds.
Cycles 1 through 35:
94°C 30 seconds
60 °C 90 seconds
72° C 120 seconds
Final treatment:
72°C 10 min
4° C Hold
Results of Copy Standard PCR
The PCR products are analyzed by agarose gel electrophoresis.
Copy sensitivity test with C467 5’ transgene specific primers on synthetic template spiked into wild type (wt) genomic DNA. Note that the C467 5’ transgene primers can detect as little as 0.001 copies of the transgene template when it is present in genomic DNA.
Lane 1: Invitrogen 1 Kb Plus Ladder.
Lane 2: 100 transgene copy standard.
Lane 3: 50 transgene copy standard.
Lane 4: 10 transgene copy standard.
Lane 5: 1.0 transgene copy standard.
Lane 6: 0,1 transgene copy standard.
Lane 7: 0.01 transgene copy standard.
Lane 8: 0.001 transgene copy standard.
Lane 9: 0.0001 transgene copy standard.
Lane 10: Wild type DNA (zero copy transgene).
Lane 11: Negative control. No template.
Lane 12: Negative control. No template.
Protocol references
Konkel DA, Tilghman SM, Leder P. 1978. The Sequence of the Chromosomal Mouse beta-globin Major Gene: Homologies in Capping, Splicing and Poly(A) Sites. Cell 15:1125-1132.
Miller SA. Dykes DD. Polesky HF. 1988. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Research. 16(3):1215.
Ralser M, Querfurth R, Warnatz HJ, Lehrach H, Yaspo ML, Krobitsch S. 2006. An efficient and economic enhancer mix for PCR.
Biochem Biophys Res Commun. 347:747-751.
Stratman JL, Barnes WM, Simon TC. 2003. Universal PCR genotyping assay that achieves single copy sensitivity with any primer pair. Transgenic Res. 12(4):521-2.
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden T. 2012. Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics. 13:134.
