Jan 29, 2026
Generation of Stickleback intestinal organoid
- Arshad A Padhiar1,
- Xihao Huang1,
- Daniel Bolnick1
- 1Department of Ecology and Evolutionary Biology, University of Connecticut, 75 North Eagleville Rd, Storrs CT 06269 USA
- University of Connecticut

Protocol Citation: Arshad A Padhiar, Xihao Huang, Daniel Bolnick 2026. Generation of Stickleback intestinal organoid. protocols.io https://dx.doi.org/10.17504/protocols.io.bp2l6e3pzgqe/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: January 28, 2026
Last Modified: January 29, 2026
Protocol Integer ID: 241696
Keywords: Threespine Stickleback, gut organoid, 3d Culture, Primary cell culture, Teleost fish, Intestinal cell culture, maintaining intestinal organoid, intestinal organoid, intestinal organoid intestinal organoid, term cell cultures from stickleback, resulting organoid, generation of stickleback, threespine stickleback, adaptable to other teleost species, term cell culture, epithelial in origin, gasterosteus aculeatus, other teleost species, stickleback, pathogen study, evolutionary biology, juvenile stickleback, cell, adaptive immunity
Funders Acknowledgements:
Gordon and Betty Moore Foundation
Grant ID: GBMF9323
Abstract
Intestinal organoids offer significant advantages over conventional 2D cultures, providing a physiologically relevant 3D architecture that enables more realistic disease modeling, host-microbe interactions, and host-pathogen studies. The threespine stickleback (Gasterosteus aculeatus) is an established teleost model widely used in evolutionary biology, adaptive immunity, and developmental genetics. However, robust long-term cell cultures from stickleback (whether fibroblast or epithelial in origin) have been challenging to establish, largely due to its cold water physiology and species-specific media requirements. Here, we present a first method for generating and maintaining intestinal organoids derived from juvenile stickleback. This protocol provides a novel experimental platform that is potentially adaptable to other teleost species. The resulting organoids are capable of expansion over multiple passages and are suitable for diverse downstream applications, including genetic, transcriptional, proteomic, and functional assays.
Guidelines
- Perform tissue dissection on a clean bench using clean, disinfected instruments.
- Use sterile consumables and reagents for all post digestion, Matrigel embedding, and culture steps.
- Minimize environmental exposure time during transfer to culture conditions.
Materials
Equipment and Consumables
- Vannas Spring Scissors (Fine Science Tools, Item No. 15000-08)
- Micro Knife – Side Cut (Fine Science Tools, Item No. 10315-12)
- Dumont #5SF Forceps (Fine Science Tools, Item No. 11252-00)
- Dumont #5 Fine Forceps (Fine Science Tools, Item No. 11254-20)
- Sterile Petri dishes
- 15 mL Falcon tubes
- 2 mL sterile Eppendorf tubes
- 6-, 12-, 24- or 48-well plates
- 70 μm cell strainers
- Sterile pipette tips
- Centrifuge
- Cryovial tubes
- Controlled rate freezing container
- Incubator (without CO₂)
Reagents
- PBS without Ca²⁺ and Mg²⁺ (Gibco)
- MS-222 (Syndel)
- DTT (Roche)
- Liberase DL (Roche)
- DNase I (Thermo Fisher Scientific)
- Fetal bovine serum (FBS) (Gibco)
- DMEM/F-12 (Thermo Fisher Scientific)
- Antibiotic-Antimycotic (Gibco)
- Corning Matrigel (356255)
- IntestiCult Organoid Growth Medium (Mouse) (STEMCELL Technologies)
- Gentle Cell Dissociation Reagent (GCDR)
- CryoStor CS10 (STEMCELL Technologies)
Troubleshooting
Problem
Low or no organoid formation
Solution
Ensure larvae are 30–60 dpf. Younger larvae often yield mixed or poorly defined epithelial tissue. Confirm complete removal of DTT before enzymatic digestion
Problem
Excessive debris or clumping after digestion
Solution
Add DNase I after initial digestion and gently pipette at defined intervals. Avoid over digestion beyond 60 minutes.
Problem
Matrigel domes spreading or collapsing
Solution
Keep Matrigel and tips ice cold during handling. Allow domes to partially solidify before transferring plates to the incubator.
Problem
Poor growth after first passage
Solution
Do not allow organoids to exceed ~80% confluency before passaging. Use gentle dissociation and avoid excessive mechanical force.
Safety warnings
- MS 222 is hazardous and must be handled according to institutional safety guidelines.
- DTT and Liberase are irritants; use gloves and eye protection.
- Matrigel solidifies rapidly at room temperature; prolonged handling can compromise embedding.
Ethics statement
All animal procedures must be approved by the Institutional Animal Care and Use Committee (IACUC) and performed in accordance with institutional and national guidelines.
Before start
- Prepare all buffers and required reagents in advance.
- Pre cool centrifuge to 4 °C.
- Thaw Matrigel on ice or at 4 °C overnight
Buffers and Media
Washing buffer: 1X PBS + 4X Antibiotic-Antimycotic
Mucus removal solution: 19 mL 1X PBS + 1 mL 0.1 M DTT + 3X Antibiotic-Antimycotic
Enzymatic digestion buffer: 1.4 Wünsch units/mL Liberase TM + 24 U/mL DNase I in 7 mL 1X PBS (without Ca²⁺/Mg²⁺) +2X antibiotic-antimycotic
Neutralization medium: DMEM/F-12 + 10% FBS + 2X antibiotic-antimycotic
Cryopreservation medium: 50% cryostar CS10 + 50% InstesiCult organoid growth medium
Organoid culture medium: IntestiCult Organoid Growth Medium (Mouse) + 1X antibiotic-antimycotic
Agar Plate for Dissection
Add 4 g agar to 200 mL sterile water. Microwave until the solution is clear.
Let the solution cool for approximately 15 minutes (ensure it remains above 50°C), then pour it into 60 mm Petri dishes, filling each about one-third full.
Allow plates to solidify at room temperature and store at 4 °C until use.
Euthanasia and Gut Isolation from Juvenile
Use 30–60 days post-fertilization (dpf) stickleback larvae. At earlier stages (<30 dpf), the gut tissue is often not fully separated from surrounding epithelium and muscle, increasing the risk of contamination from other cell types.
Euthanize one larva fish at a time in 0.5 g/L MS-222 (pH 7.4), following the approved IACUC guidelines.
Place the euthanized fish on a 2% agar plate under a dissection microscope (Figure 1).
Clamp the fish trunk with tweezers for stabilization.
Rip the abdomen longitudinally from the anus to the heart using the micro-knife.
Using a micro-knife, make a longitudinal incision from the anus towards the heart to open the abdomen.
Carefully pull the intestinal tract out by holding the distal segment and pulling gently towards the head of the fish (Figure 2).
Place the isolated gut into ice-cold washing buffer.
Mucus Removal
Squeeze out the gut content by pressing the gut with closed tweezers. Larvae fed with brine shrimp have orange-colored gut contents that can be distinguished and removed easily (Figure 3)
Transfer the gut to fresh PBS with 3X Antibiotic-Antimycotic and place the Petri dish over crushed ice.
Repeat the dissection procedure for the remaining larvae.
Pool guts from approximately 10 larvae into a new dish containing ~20 mL of mucus removal solution.
Chop tissue into small pieces (∼1–2 mm) using the micro-knife directly in the chopping buffer.
Transfer tissue fragments to a 15 mL tube and centrifuge at 2000 RPM for 3 min at 4 °C.
Carefully aspirate the supernatant, resuspend pellet in 10 mL ice-cold washing buffer, and centrifuge again to ensure complete removal of DTT, which can inhibit subsequent enzymatic digestion.
Enzymatic Digestion
Resuspend the washed tissue pellet from 10 larvae in 5 mL of enzymatic digestion buffer.
Incubate at room temperature for 30 minutes.
Every 15 min, gently pipette tissue fragments up and down 10–15 times using a P1000 tip to mechanically disrupt stroma and release crypts.
Add 20 U/mL DNase I after 30 min to prevent clumping from released DNA.
Continue incubation for an additional 20–30 min (total digestion time 45–60 min).
Neutralization and crypt filtration
Add an equal volume of neutralization medium to the digestion mixture to terminate Liberase/DNase activity.
Pass the suspension through a 70 μm cell strainer into a new 50 mL or 15 mL tube to remove undigested fragments. A 40 μm mesh filter can also be used and may reduce contaminating debris, but could lower the crypts yield.
Centrifuge the filtrate at 800 × g for 5 min at 4 °C or room temperature.
Carefully discard the supernatant and place the tube on ice to keep the pellet cold.
Matrigel Embedding and Plating
Resuspend the pellet in cold matrigel. For a preparation from 10 juvenile fish, approximately 400 μL total Matrigel is sufficient.
Mix gently with pre-chilled tips to avoid introducing bubbles.
Plate Matrigel domes in tissue culture plates (35 μL for 24-well, 60 μL for 12-well, or 90 μL for 6-well plates, per well).
Let the plate sit in the culture hood for 5 min to initiate solidification, then incubate at 25 °C for 10–15 min to allow Matrigel domes to fully solidify without disturbance.
Carefully overlay each dome with organoid culture medium, ensuring the domes remain intact.
Fill neighboring unused wells with PBS or sterile water to maintain humidity during culture.
Initial Culture and Medium Changes
Change the medium after 24 h and again after 48 h to reduce debris and minimize contamination risk.
After the first 48 h, maintain the cultures by adding fresh medium on top (for example, ~1 mL per well of a 6-well plate, ~500 μL per well of a 12-well plate), rather than fully replacing the medium, to reduce mechanical disturbance of early organoids.
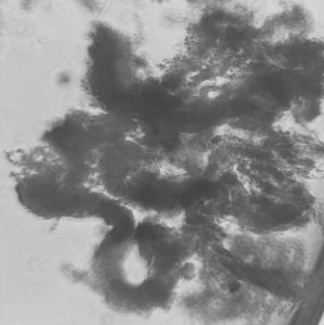
Continue culture at 25 °C and monitor for organoid colony formation. Colonies typically become evident within 2–3 weeks (Figure 4).
Organoid Expansion and Passaging
Allow organoids to reach an appropriate size before passaging. Then aspirate the medium and gently wash the Matrigel domes once with 1X PBS.
Add Gentle Cell Dissociation Reagent (GCDR) to cover each dome and incubate for 1–2 min at room temperature in the culture hood.
Pipette up and down 15–20 times with a P1000 tip to dislodge organoids from the Matrigel. Ensure the pipette tip is pre-wetted to minimize adherence of organoids to the plastic.
Optionally, to obtain clonal organoid lines, individual colonies can be isolated under an inverted microscope using a sterile 18–20 G needle, aspirated into a 200 μL pipette tip, and transferred into a separate Eppendorf tube. Each colony can then be mechanically broken into smaller fragments for clonal expansion.
Transfer the organoid suspension to a 15 mL tube and add DMEM/F-12 containing 2% FBS to a final volume of ~10 mL to neutralize the dissociation reagent completely.
Centrifuge at 2,000 rpm for 5 min. Discard the supernatant and resuspend the pellet in cold Matrigel.
Typically, single well containing a mature organoid dome can be split into 4–5 new wells. After the first passage, cells usually proliferate faster, with new organoids emerging within 2–3 days.
Seed new Matrigel domes and overlay with IntestiCult medium as above.
Do not allow organoids to exceed more than ~80% confluency within the Matrigel dome, as excessive density can promote differentiation and loss of stemness.
Cryopreservation of Stickleback Intestinal Organoids
Aspirate culture medium and gently wash the Matrigel domes once with chilled 1X PBS (without Ca²⁺/Mg²⁺).
Add sufficient Gentle Cell Dissociation Reagent (GCDR) or chilled PBS to fully cover each dome and incubate for 1–2 min at room temperature under the culture hood.
Using a pre-wetted P1000 tip (pre-rinsed with cold PBS or GCDR), pipette up and down multiple times to dislodge organoids from the Matrigel and mechanically fragment large domes into smaller clusters.
Transfer the suspension to a pre-chilled 15 mL tube and bring the volume up to ~10 mL with ice-cold PBS.
Keep the tube on ice for several minutes to facilitate further solubilization of residual Matrigel and separation of organoids from matrix components.
If GCDR is used instead of PBS then neutralize by adding an equal volume of chilled DMEM/F-12 containing 5% FBS.
Centrifuge at 2,000 rpm for 5 min at 2–4 °C and carefully discard the supernatant.
Optional: Wash the pellet again with chilled PBS if matrigel is visible from the naked eye. Centrifuge again under the same conditions.
Prepare cell freezing medium consisting of 50% CryoStor CS10 (or equivalent 10% DMSO-based cryopreservation medium) and 50% IntestiCult organoid growth medium (Mouse).
Gently resuspend the organoid pellet in 2–3 mL of pre-cooled freezing medium (50% CryoStor CS10 + 50% IntestiCult), depending on organoid density; for example, organoids from one 12 or 6-well dome can be resuspended in 2–3 mL and then aliquoted into 3 labeled cryovials with 0.5–1.0 mL per vial, ensuring an even distribution of organoid fragments in each tube.
Place cryovials into a controlled-rate freezing container (isopropanol-free with metal coil/foam box or isopropanol-based) and transfer immediately to a -80 °C freezer.
Keep vials at -80 °C for at least 24–48 h before transferring them to liquid nitrogen for long-term storage.
Note: Organoids generally maintain acceptable morphology and viability for at least several weeks at -80 °C, but long-term storage is recommended in liquid nitrogen.
Thawing and Organoids recovery
Remove cryovials from -80 °C or liquid nitrogen and immediately place them at room temperature.
Thaw each vial manually by gently rotating it between the palms; perform this action slowly to avoid uneven heating and thermal shock.
Once a small ice crystal remains, disinfect the outside of the vial with 70% ethanol and transfer it into the culture hood.
Gently transfer the thawed cell suspension to a 15 mL tube. In the culture hood, gently add 1 mL room temperature DMEM/F-12 directly to the cryovial containing the thawed organoid suspension, dropwise while swirling to dilute the cryoprotectant.
Centrifuge the 2 mL cryovial directly at 2,000 rpm for 5 min at room temperature to pellet the organoids.
Carefully aspirate the supernatant and place the tube on ice for 1–2 min.
Resuspend the pellet in an appropriate volume of ice-cold Matrigel and mix gently to obtain a homogeneous suspension.
Seed Matrigel domes into 6-, 12-, 24- or 48-well plates according to the experimental needs.
Allow the plate to sit under the hood (room temperature) for ~5 min to let the domes settle and begin to gel, then transfer to the 25 °C incubator for 10 to 15 min for complete solidification.
Once the domes have solidified, carefully overlay them with pre-warmed IntestiCult organoid growth medium, taking care not to disturb the Matrigel domes.
Return the plate to the incubator and monitor organoid recovery over the following days.


