Jun 18, 2025
Generation of iPSC-derived Microglia
- Yasmine Nonose1,
- Niraj Sawarkar2,
- Gist Croft1
- 1New York Stem Cell Foundation;
- 2NYSCF

Protocol Citation: Yasmine Nonose, Niraj Sawarkar, Gist Croft 2025. Generation of iPSC-derived Microglia. protocols.io https://dx.doi.org/10.17504/protocols.io.eq2lyqo4wvx9/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: June 17, 2025
Last Modified: June 18, 2025
Protocol Integer ID: 220423
Keywords: ASAPCRN, iPSC-derived microglia, differentiation, microglia, brain macrophages, embryoid bodies, ipsc differentiation to microglia, derived microglia, mature microglia, microglial identity, microglia, relevant microglia, redundant for microglial identity, neuroinflammatory mechanisms in neurodegenerative disease, neuroinflammatory mechanism, physiology in neuroinflammation, neurodegenerative disease, generation of ipsc, ipsc differentiation, neuroinflammation, ipsc
Funders Acknowledgements:
Aligning Sciences Across Parkinson's (ASAP)
Grant ID: ASAP-000472
Disclaimer
This is a working protocol based on the work of Washer and colleagues (Washer et al., 2022), and was used to validate differentiation conditions for obtaining iPSC-derived microglia.
Abstract
This is a working protocol based on the publication of Washer et al., 2022 (Nat Sci Reports). This protocol can be used to obtain fully mature microglia to investigate neuroinflammatory mechanisms in neurodegenerative diseases. One of the advantages of this protocol is that it assessed systematically medium components, coatings, and growth factors required for iPSC differentiation to microglia, optimizing several components that were redundant for microglial identity. They use defined medium which produces both transcriptionally and functionally relevant microglia for modeling these cells’ physiology in neuroinflammation and for drug discovery.
Attachments
Image Attribution
Images proprietary of authors; schematics done with Biorender
Guidelines
For this protocol internal validation, we used H9 iPS cell lines, especially H9 WT.
Materials
- Base Medium
| A | B | C | |
| MEDIUM | SUPPLIER | CAT. NO | |
| Stemflex | Thermo Fisher | A3349401 | |
| X VIVO 15 | Lonza | BE02-060F | |
| Advanced RPMI | Thermo Fisher | 12633020 |
- Solutions
| A | B | C | |
| REAGENT | SUPPLIER | CAT. NO | |
| Anti-adherence rising solution | STEMCELL Technologies | 7010 | |
| PBS | Thermo Fisher | 10010049 | |
| GCDR | STEMCELL Technologies | 100-0485 | |
| Accutase | Thermo Fisher | A1110501 | |
| GlutaMAX‱ | Life Technologies | 35050-061 |
- Substrate
| A | B | C | |
| REAGENT | SUPPLIER | CAT. NO | |
| Fibronectin | Thermo Fisher | CB-40008A |
- Small molecules
| A | B | C | D | |
| SMALL MOLECULE | SUPPLIER | CAT. NO | QUANTITY (ug) | |
| BMP-4 | R&D | 314-BP-500 | 500 | |
| VEGF | R&D | 293-VE-500 | 500 | |
| SCF | R&D | 255-SC-01M | 1000 | |
| M-CSF | R&D | 216-MC-100 | 100 | |
| IL-3 | R&D | 203-IL-500 | 500 | |
| IL-34 | R&D | 5265-IL-010 (ORDER BULK) | ||
| GM-CSF | R&D | 215-GM-500 | 500 | |
| TGF-B1 | R&D | 216-MC-100 | 100 |
Quantity as a suggestion, please evaluate how much will suit your experiments.
- Culture Vessels
| A | B | C | D | |
| PLATE | # OF WELLS | SUPPLIER | CAT. NO | |
| AggreWell 400 | 24 | STEMCELL Technologies | 34415 | |
| AggreWell 800 | 24 | STEMCELL Technologies | 34815 | |
| 6-well plates | 6 | Fisher Scientific | 3516 | |
| 6-well plates | 6 | Fisher Scientific | 140675 |
AggreWell400 and AggreWell800 both work, with the AggreWell800 helping avoid EBs to fuse.
- Others
| A | B | C | |
| REAGENT | SUPPLIER | CAT. NO | |
| Y-27632 | Tocris | 1254 |
Troubleshooting
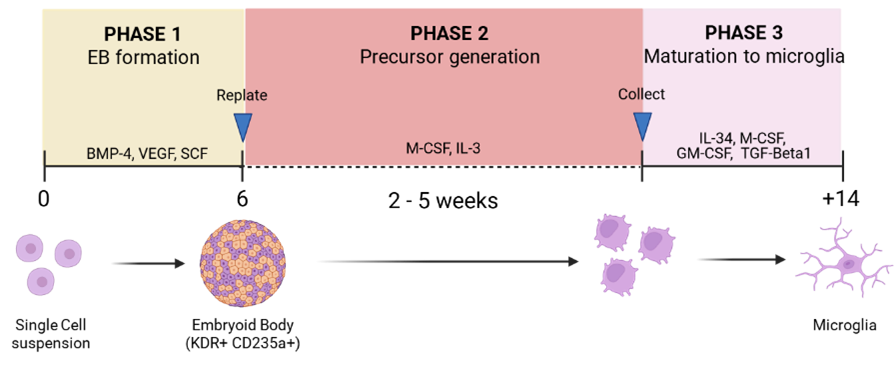
PHASE 1 - Generation of Embryoid bodies (EBs)
We maintain iPSCs in Laminin 521 (1:20 in PBS +/+) or in Cultrex (1:100 in DMEM/F12) and feed them with StemFlex supplemented every other day until they reach 70-80% confluency. After thawing, it usually takes 4 days.
Day 0: Prepare the Aggrewell plate with 0.5 mL of anti-adherence rising solution.
Spin at 1300 – 3000 x g for 5-3 min to remove bubbles from the microwells.
Remove the anti-adherence rising solution, and wash the wells with PBS/water 3
times before the addition of 1 mL of 2x concentrated EB medium supplemented
with Rhok inhibitor (Y-27632).
EB Media: Add small molecules fresh.
| A | B | C | D | |
| Component | Stock concentration | Final concentration | Final Volume for 10 mL of EB Media (uL) | |
| Stem Flex supplemented | N/A | N/A | 10 mL | |
| BMP-4 | 100 μg/mL | 50 ng/mL | 5 | |
| VEGF | 100 μg/mL | 50 ng/mL | 5 | |
| SCF | 100 μg/mL | 20 ng/mL | 2 | |
| Y-27632 * | 10 mM | 20 μM | 20 |
* Only on day 0. No need to add on the feeds at d1-6.
Dissociate iPSCs (70-80% confluency) by washing with PBS 1x and then incubate with GCDR for 30 min at 37 °C, 5% CO2. Transfer cells to a tube containing DMEM/F12 and pellet by centrifugation at 300 x g for 5 min.
Aspirate supernatant and resuspend cells in StemFlex for counting.
Cell viability needs to be >75% to ensure good iPSC quality and EB formation.
Prepare cell seeding stock: Seed 4 x 10^6 cells/mL (AggreWell 800) or 2 x 10^6 cells (AggreWell 400) per well; add 1 mL of cells to one well of the Aggrewell 800 containing 1 mL of 2x concentrated EB media.
After seeding, spin plate at 100 x g for 3 min to capture cells in the microwells.
Leave the EBs on the plate for 4-6 days, doing daily feeds of 50-75% media change with 1x EB media. Be careful to not dislocate the EBs and cause them to fuse. Aspirate from the meniscus to preserve the EBs on the bottom. Handle the plate gently.
[CHECKPOINT - OPTIONAL] On day 7, you can run a FACS as a checkpoint for hemangioblasts (KDR+/CD235a/b+) (Guttikonda et al., 2021).
| A | |
| Separate 100 uL of EB suspension. Spin down at 300 x g for 5 min, aspirate media and start dissociation with Accutase for 30 min. It may need mechanical dissociation as well to break the EBs. Verify if dissociation was successfull before next step. ALWAYS have a non-stained control for cell autofluorescence. | |
| Add FACS Buffer to tubes containing dissociated EBs and Accutase. BSA fraction V may help stop Accutase reaction. Spin down at 300 x g for 5 min and aspirate media. | |
| Add FACS Buffer containing the KDR-PE (1:20) and CD235a (1:20). Incubate on ice or 4dg for 30 min. | |
| Prepare Sytox solution (1 to 1000 in FACS buffer). Make enough solution so all the samples can be resuspended in 300 uL | |
| Spin down at 300 x g for 5 min and add 300 uL of FACS Buffer. | |
| Read on Attune. At least 10-15% of double-positive is great to proceed with the differentiation. 20% and above is exceptional and will have a high yield. |
PHASE 2 - Microglia precursor generation (monolayer)
Day 5: Thaw enough aliquots of fibronectin in 4 °C overnight.
Day 6: Coat the culture vessels you will need with Fibronectin 10 μg/mL in PBS +/+ (containing calcium and magnesium). Let it coat in 4 °C overnight, or you can do it fresh on d7 (coat for 3 h at room temperature).
Day 7: Pipette up and down to collect all the EBs from the Aggrewell. Collect them in a 15ml/50mL conical.
Spin at 120 - 300 x g for 5 min or just let the EBs sediment in the tube.
Prepare “FACTORY” / Precursor Medium (see below)
FACTORY / Precursor Medium: Add small molecules fresh.
| A | B | C | D | |
| Component | Stock concentration | Final concentration | Final Volume for 10 mL of EB Media (uL) | |
| X VIVO 15 | N/A | N/A | 10 mL | |
| GlutaMAX | 100X | 1X | 100 | |
| M-CSF | 100 μg/mL | 100 ng/mL | 10 | |
| IL-3 | 100 μg/mL | 25 ng/mL | 2.5 |
Aspirate the old supernatant and resuspend the EBs in Precursor Medium.
Seed aproximately 100-200 uL of EB suspension in one well of a 6-well plate coated with fibronectin (for pooling 5 wells from a AggreWell400; reference value), or alternatively use a T-75 flask.
Distribute the EBs fairly in the well and leave the plate at 37 °C. Check the attachment on the next day.
OBS: It can take 48 - 72 h before they recover and form a reasonable monolayer - leave the plate alone for this period, then change half the medium if necessary. Otherwise, half media changes can be done once a week using enough volume (minimum of 3 mL per well in a 6-well plate).
PHASE 3 - Maturation to microglia
Day 14-35: When changing the Precursor Media, collect precursors (floating cells), and filter through a 40 μm strainer. Spin down at 300 x g for 5 min.
Resuspend in Maturation media 1. COUNT CELLS.
Maturation Media 1:
| Component | Stock concentration | Final concentration | Final Volume for 10 mL of EB Media (uL) | |
| Advanced RPMI | N/A | N/A | 10 mL | |
| GlutaMAX | 100X | 1X | 100 | |
| IL-34 | 100 μg/mL | 100 ng/mL | 10 | |
| M-CSF | 100 μg/mL | 25 ng/mL | 2.5 | |
| GM-CSF | 100 μg/mL | 10 ng/mL | 1 | |
| TGF-Beta1 | 50 μg/mL | 50 ng/mL | 10 |
[OPTIONAL] Take an aliquot to run Attune on CD43/CD45. Cells are expected to be >90% double-positive. Alternatively, use CD11c-PE or CD11c-647 as a dendritic cell marker to observe the precursors under the microscope.
FOR MATURATION: Plate in a ULA or TC-treated / fibronectin-coated vessel at 50,000 cells/cm2 with enough Maturation Media volume.
OBS: If your plating density is higher, you will have to feed the cells more often, as the IL-34 will be limited, and you will start seeing cell death. Plating at 50,000 cells/cm2 allows you to feed twice a week with enough media volume. Half media changes are encouraged.
Mature for 12-14 days with Maturation medium 1.
After, maintain cells in Maturation medium 2 until experiments.
Maintenance: Change media every 3 days, being consistent in the time of the feeds (e.g., around 2pm every time).
Maturation
Medium 2 (Maintenance medium):
| Component | Stock concentration | Final concentration | Final Volume for 10 mL of EB Media (uL) | |
| Advanced RPMI | N/A | N/A | 10 mL | |
| GlutaMAX | 100X | 1X | 100 | |
| IL-34 | 100 μg/mL | 100 ng/mL | 10 | |
| M-CSF | 100 μg/mL | 25 ng/mL | 2.5 |
FOR CRYOPRESERVATION - Resuspend the counted cells in 500 μL of Maturation medium (base media) per 0.5M - 2.0 M cells. Add 500 μL of 2xFM per cryovial.
The next day, remove cryovials from -80 and store them in a liquid nitrogen tank.
Protocol references
Washer SJ, Perez-Alcantara M, Chen Y, Steer J, James WS, Trynka G, Bassett AR, Cowley SA. Single-cell transcriptomics defines an improved, validated monoculture protocol for differentiation of human iPSC to microglia. Sci Rep. 2022 Nov 14;12(1):19454. doi: 10.1038/s41598-022-23477-2. PMID: 36376339; PMCID: PMC9663826.
Guttikonda SR, Sikkema L, Tchieu J, Saurat N, Walsh RM, Harschnitz O, Ciceri G, Sneeboer M, Mazutis L, Setty M, Zumbo P, Betel D, de Witte LD, Pe'er D, Studer L. Fully defined human pluripotent stem cell-derived microglia and tri-culture system model C3 production in Alzheimer's disease. Nat Neurosci. 2021 Mar;24(3):343-354. doi: 10.1038/s41593-020-00796-z. Epub 2021 Feb 8. PMID: 33558694; PMCID: PMC8382543.
Acknowledgements
We thank Sigi Benjamin for helping run FACS/Attune during the optimization of this protocol, Matt Zimmer for setting the panel in Attune, and for consulting on the fluorophores that would work best for our experiments.