Aug 02, 2024
Generating Nuclei Segmentations for vEM datasets using Cellpose
- Emma Avetissian1,
- Alyson Petruncio1,
- Aubrey Weigel1,
- CellMap Project Team1,
- Jeff Rhoades1
- 1HHMI Janelia Research Campus

Protocol Citation: Emma Avetissian, Alyson Petruncio, Aubrey Weigel, CellMap Project Team, Jeff Rhoades 2024. Generating Nuclei Segmentations for vEM datasets using Cellpose . protocols.io https://dx.doi.org/10.17504/protocols.io.n2bvjnq8pgk5/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: August 01, 2024
Last Modified: August 02, 2024
Protocol Integer ID: 104387
Keywords: Image segmentation, nucleus, volume electron microscopy, cellpose, nuclei segmentations for vem dataset, generating nucleus segmentation, deep learning models on cell, generating nuclei segmentation, nuclei segmentation, organelle segmentation, image stacks of cell, nucleus segmentations in fib, vem dataset, 3d predictions for the entire dataset, segmentation, application of cellpose, tuning 2d model, cell, training deep learning model, using cellpose, 3d prediction, focused ion beam scanning electron microscopy, training other machine learning model, other machine learning model, image stack, 3d, nuclei
Abstract
This protocol details the process of generating nucleus segmentations in FIB-SEM (Focused Ion Beam Scanning Electron Microscopy) image stacks of cells and tissues using Cellpose. These segmentations are crucial for gaining biological insights and training deep learning models on cell and organelle segmentation. The protocol covers the application of Cellpose for fine-tuning 2D models specifically for nuclei segmentation. Once fine-tuned, the model is used to generate 3D predictions for the entire dataset. These segmentations can be further refined through manual approaches, utilized for visualization, or employed in training other machine learning models.
Materials
Software
Cellpose
NAME
Marius Pachitariu & Carsen Stringer
DEVELOPER
REPOSITORY
SOURCE LINK
Introduction:
The purpose of this protocol is to provide comprehensive instructions on how to create nuclei segmentations in FIB-SEM datasets on OpenOrganelle.org using Cellpose.
Cellpose is a generalist algorithm developed by Stringer et al., (2020) to segment cellular structures. Cellpose also provides software for manual segmentation and model training. In this protocol we discuss the use of Cellpose v3.0.9.
Citation
LINK
OpenOrganelle.org is an online platform that provides access to high-resolution 3D electron microscopy images of cellular structures for scientific research and education.
In this protocol you can expect to find:
- OpenOrganelle Overview
- Using Fiji to save files in the cloud locally as 2D tiffs
- Cellpose in 2D and Model Training
- Cellpose in 3D and Predictions
- Final Touches
OpenOrganelle Overview
OpenOrganelle (Heinrich et al, 2021) is a collection of focused ion beam scanning electron microscopy (FIB-SEM) datasets curated by the Janelia CellMap Project Team. Each dataset on OpenOrganelle includes important background information about the sample, protocol, contributors, and more. The “VIEW” button opens a new tab to view the chosen dataset in Neuroglancer. Each dataset is also accompanied by a Fiji link which copies the location of the dataset in the S3 bucket (see Figure 1). This link was used to read data into Fiji and save as 2D slices for Cellpose.
Figure 1: OpenOrganelle page. The Fiji link is located on the right-hand side.
Citation
LINK
Citation
LINK
Using Fiji to save files in the cloud locally as 2D tiffs
Figure 2: How to open a dataset from OpenOrganelle in Fiji
Head to: File → Import → HDF5/N5/Zarr/...
Paste the link into the search bar.
Click Detect Datasets
Navigate through the file container by clicking the dropdown, e.g. “em”.
Choose resolution, e.g., s0 (original), s1 (2x downsample), s3 (4x downsample), etc.
Note
This step involves reading the data from the S3 bucket into your local computer memory. The datasets can be very large; care should be taken when choosing which resolution to read. The approximate size of the image volume is listed e.g., "s0 (12400x500x9382, int16 → 16-bit) (116.3 GB)"
Click OK. See Figure 2.
Once the dataset is open in Fiji, it can be saved as a sequence of 2D slices for reading into Cellpose.
From the open dataset window go to:
File → Save As → Image Sequence → Save into a new folder
See Figure 3.
Figure 3: Process of saving 2D slices in Fiji.
Before closing the image in Fiji, save a second copy of the dataset as a 3D tiff. This will be used for generating 3D predictions in a later step in Cellpose. If the image is closed before creating the 3D tiff, you can simply drag the folder containing the image sequence into the main Fiji bar to open it. To save it as a 3D tiff:
File → Save As → Tiff
See Figure 4.
Figure 4: How to save 2D slices as a 3D tiff.
Note
While creating annotations in Cellpose for model finetuning, it is helpful to keep the image volume open in Fiji as a reference. It is not always evident what an organelle is from just the 2D slices presented in Cellpose. To open the 3D tiff, drag the tiff file into the main Fiji menu. A scroll bar will appear at the bottom for viewing the entire stack.
Cellpose in 2D and Model Training
To begin using Cellpose on the dataset, open the program (see Appendix). More detailed instructions for general Cellpose use can be found on https://cellpose.readthedocs.io/en/latest/gui.html. Within Cellpose go to:
File → Load Image → Select a slice for segmentation
See Figure 5.
Figure 5: How to load a single slice for annotation in Cellpose.
| A | B | |
| Zoom | Hold right click and drag | |
| Zoom | Scroll wheel | |
| Pan | Hold left click and drag | |
| Create mask | Tap right click and drag | |
| Remove mask | Ctrl left click | |
| Undo remove mask | Ctrl Y | |
| Toggle mask visibility | X | |
| Toggle mask outlines | Z |
Table 1: Basic key binds for annotation in Cellpose
Annotate the nuclei as accurately as possible.
At first, it may be beneficial to use an existing model to make annotating easier. Out of the box, the cyto2_cp3 model tends to give promising results for nuclei in FIB-SEM volumes of tissue. If the model does not catch anything, try annotating a couple slices manually and training a model. Continue to use and improve the model by periodically adding more training slices.
Note
Make sure to clean up missing spots, mistaken predictions, and tidy up the edges.
Refer to Figure 6 for what a completed slice might look like.
Figure 6: Example of a fully annotated slice from the dataset jrc_mus-kidney-2.
Once the annotation is complete, save the slice along with the masks.
File → Save masks and image (as *_seg.npy)
This file will automatically be saved into the folder where the rest of the 2D slices can be found.
Note
Do not change the name as this is how Cellpose will recognize that the file should be used for training.
To train a model on annotated slices, go to:
Models → Train new model with image+masks in folder
Set the initial model to cyto2_cp3
Adjust n_epochs to change the number of epochs (we generally went with 1000)
Name the model for organizational purposes. Click “OK” and wait for the training to finish.
Note
We found that training took at most 15 minutes and the largest amount of training slices required for an adequate model was 38 slices. Progress can be tracked in the terminal (See Figure 7). The model’s file will automatically be created in a separate “models” folder that is placed inside the folder that contains the 2D slices.
Figure 7. Screenshot of the terminal while training is in progress. The number after “[INFO]” displays the current epoch.
To run this model on a new 2D slice
Open the blank 2D slice and locate the section labeled “Other models” on the left-hand side of the program.
Open the dropdown for custom models, select the new model, and click run.
Repeat this process, making use of previous models, annotating additional slices until the model performs satisfactorily on 2D slices.
Cellpose in 3D and Predictions
Cellpose can extend models that were trained exclusively on 2D data to perform predictions in 3D. Once the model demonstrates a satisfactory performance in 2D, we attempt to run a prediction in 3D.
Note
In Cellpose v3.0.9, to create a 3D prediction, the program requires explicitly instructing the program to handle 3D data via a specific flag. This adjustment ensures that Cellpose interprets and processes the volumetric data correctly.
Close Cellpose
Into the terminal enter:
python -m cellpose --Zstack
This will alert Cellpose to load data as 3D
Now load in the previously saved 3D volume by:
File → Load Image → Select the tiff file
Once the tiff has loaded, run the custom model as described for 2D in Step 15.
This may take a while (on the order of tens of minutes) and progress can be tracked in the terminal.
The process will be complete when the masks are visible (see Figure 8).
Figure 8: A finetuned Cellpose model has been applied to the dataset jrc_mus-kidney-2.
The terminal will also contain how many masks were found at the end of the run (see Figure 9).
Figure 9: Screenshot of the terminal after Cellpose ran a 3D prediction. The arrow points to where the terminal lists out how many masks were found.
To save the masks go to:
File → Save masks as PNG/tiff
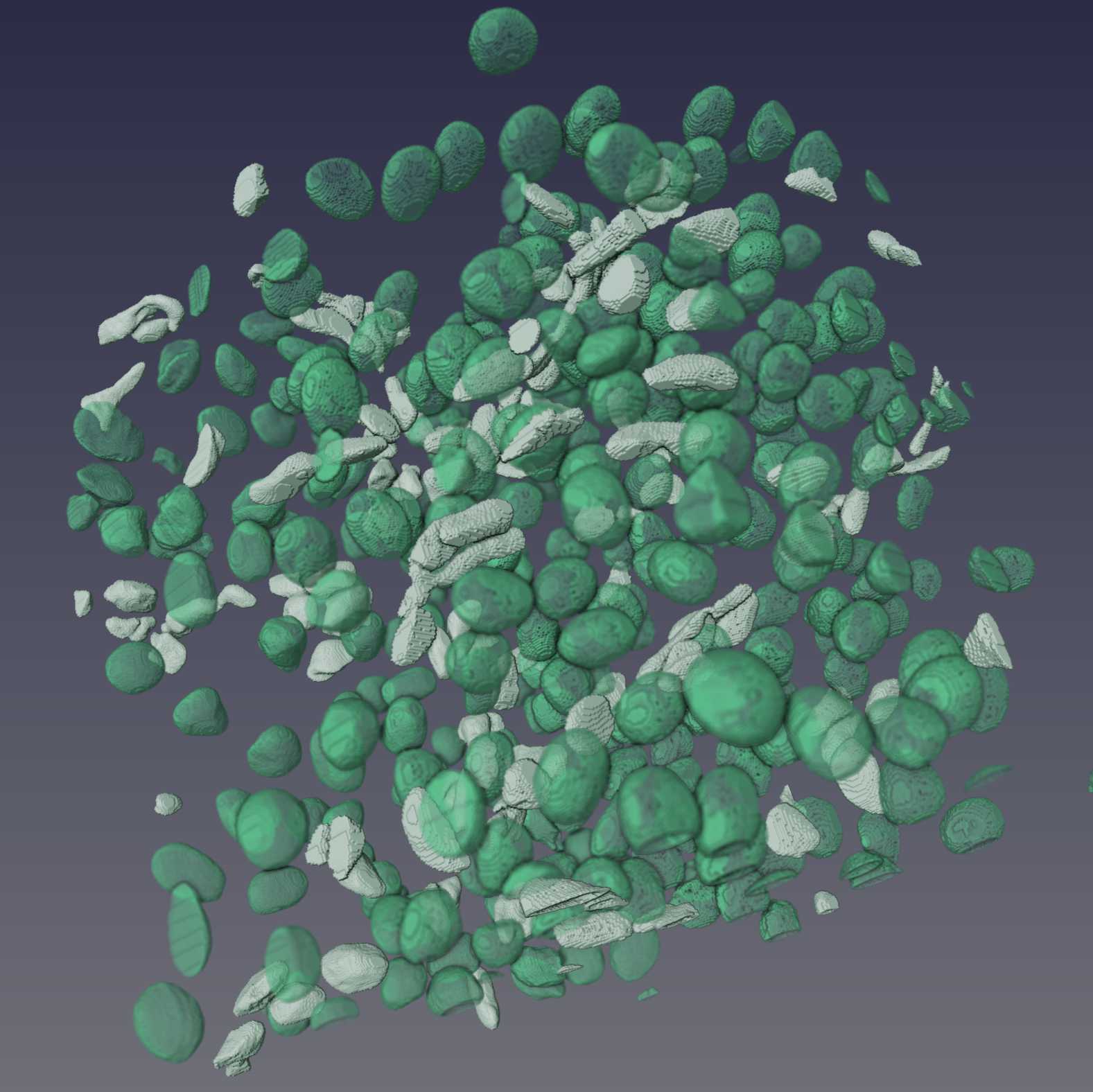
Figure 10: Example results of Cellpose segmentations from three datasets. Shown are 3D renderings of the segmentations that have been manually proofread.
Protocol references
Citation
LINK
Citation
LINK
Citation
LINK
Citation
LINK
Citations
Heinrich L, Bennett D, Ackerman D, Park W, Bogovic J, Eckstein N, Petruncio A, Clements J, Pang S, Xu CS, Funke J, Korff W, Hess HF, Lippincott-Schwartz J, Saalfeld S, Weigel AV, COSEM Project Team. Whole-cell organelle segmentation in volume electron microscopy.
https://doi.org/10.1038/s41586-021-03977-3Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A. Fiji: an open-source platform for biological-image analysis.
https://doi.org/10.1038/nmeth.2019Stringer C, Wang T, Michaelos M, Pachitariu M. Cellpose: a generalist algorithm for cellular segmentation.
https://doi.org/10.1038/s41592-020-01018-xPachitariu M, Stringer C. Cellpose 2.0: how to train your own model.
https://doi.org/10.1038/s41592-022-01663-4Step 1
Stringer C, Wang T, Michaelos M, Pachitariu M. Cellpose: a generalist algorithm for cellular segmentation.
https://doi.org/10.1038/s41592-020-01018-xStep 2
Heinrich L, Bennett D, Ackerman D, Park W, Bogovic J, Eckstein N, Petruncio A, Clements J, Pang S, Xu CS, Funke J, Korff W, Hess HF, Lippincott-Schwartz J, Saalfeld S, Weigel AV, COSEM Project Team. Whole-cell organelle segmentation in volume electron microscopy.
https://doi.org/10.1038/s41586-021-03977-3Step 2
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A. Fiji: an open-source platform for biological-image analysis.
https://doi.org/10.1038/nmeth.2019