Jul 07, 2025
Version 1
Formulation and Characterization of Primary Tissue-Derived Microfluidic Droplet-Engineered Organoids V.1
- Wanlong Wang1,2,
- Yongde Cai1,2,
- Xiaoyong Dai1,2,3,
- Haowei Yang1,2,
- Davit Khutsishvili1,
- Jiawei Li1,
- Yu Zhu1,
- Jin Wang2,
- Xiu Yan2,
- Zitian Wang1,
- Shaohua Ma1,2,4
- 1Tsinghua Shenzhen International Graduate School (SIGS), Tsinghua University, Shenzhen, China;
- 2Synorg Biotechnology (Shenzhen) Co. Ltd., Shenzhen, China;
- 3Department of Physiology, School of Medicine, State Key Laboratory of Bioactive Molecules and Druggability Assessment, and Guangdong Province Key Laboratory of Pharmacodynamic Constituents of TCM and New Drugs Research, Jinan University, Guangzhou 510632, China;
- 4Key Laboratory of Industrial Biocatalysis, Ministry of Education, Tsinghua University, Beijing, China
- Wanlong Wang: These authors contributed equally to this work;
- Yongde Cai: These authors contributed equally to this work;
- Shaohua Ma: Corresponding email: [email protected];

Protocol Citation: Wanlong Wang, Yongde Cai, Xiaoyong Dai, Haowei Yang, Davit Khutsishvili, Jiawei Li, Yu Zhu, Jin Wang, Xiu Yan, Zitian Wang, Shaohua Ma 2025. Formulation and Characterization of Primary Tissue-Derived Microfluidic Droplet-Engineered Organoids. protocols.io https://dx.doi.org/10.17504/protocols.io.yxmvmb77og3p/v1
Manuscript citation:
Jiang S, Zhao H, Zhang W, et al. An automated organoid platform with inter-organoid homogeneity and inter-patient heterogeneity[J]. Cell Reports Medicine, 2020, 1(9).
Wang W, Cai Y, Davit K, et al. Automated Device for Small-Tissue Extraction and Primary Organoid Modeling[J]. Bio-Design and Manufacturing, 2025 (in press)
Yang H, Li J, Zheng Y, et al. Ultra-small tissue-compatible organoid printer for rapid and controllable modeling of respiratory organoids[J]. Device, 2024, 2(8).
Ma S, Wang W, Zhou J, et al. Lamination-based organoid spatially resolved transcriptomics technique for primary lung and liver organoid characterization[J]. Proceedings of the National Academy of Sciences, 2024, 121(46): e2408939121.
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: July 04, 2025
Last Modified: July 07, 2025
Protocol Integer ID: 221784
Keywords: droplet-engineered organoid, microfluidics, automatic device, cell extraction, organoid lamination, high-throughput, organoid formulation, derived microfluidic droplet, microfluidic droplet, characterization of microfluidic droplet, microfluidic bioprinter, engineered organoids organoid technology, integrated microfluidic bioprinter, engineered organoid, rapid organoid characterization, organoids organoid technology, native tissue microenvironment, existing organoid protocol, use of organoid, organoid protocol, primary tissue sample, tissue extraction device, derived organoid, organoid, precise organoid, throughput fabrication of uniform cell, purification of viable primary cell, characterization of primary tissue, modeling human tissue, cell analysis, primary tissue, laden microsphere, viable primary cell, including diverse cell type, human tissue, throughput applications such as drug screening, diverse cell type, clinical biopsy, single cell, tissue sample
Funders Acknowledgements:
National Key-Area Research and Development Program of China
Grant ID: 2024YFA0919800
National Natural Science Foundation of China
Grant ID: 32371470 and 82341019
Merck Research Grant, and the Cross-disciplinary Research and Innovation Fund of Tsinghua SIGS
Grant ID: No. JC2022007
Guangdong Basic and Applied Basic Research Foundation
Grant ID: 2023B1515120025
Shenzhen Fundamental Research Program
Grant ID: No. JCYJ20240813112004006
Disclaimer
The authors declare no competing interests.
Abstract
Organoid technology offers a powerful platform for modeling human tissues, studying disease mechanisms, and developing personalized therapies. However, widespread clinical application is hindered by challenges in scalability, reproducibility, and the handling of ultra-small tissue samples typical of clinical biopsies. Here, we present a comprehensive and automated protocol for the formulation and characterization of microfluidic droplet-engineered organoids (DEOs) derived from primary tissue samples. This protocol integrates 3 key stages: (1) Extraction and purification of viable primary cells from ultra-small tissue specimens using the small-Tissue Extraction Device (sTED); (2) High-throughput fabrication of uniform cell-laden microspheres using an integrated microfluidic bioprinter (OrgFab), capable of generating over 100 organoids from just 10 μL of bioink; and (3) Rapid organoid characterization using lamination-based processing for single-cell analysis while preserving spatial context. The automated workflow minimizes manual intervention, reducing variability and enhancing reproducibility, making it suitable for high-throughput applications such as drug screening and disease modeling. Our method allows for the generation of patient-derived organoids that closely mimic the native tissue microenvironment, including diverse cell types and structural features, within a significantly reduced timeframe. This 3-stage protocol enhances the use of organoids in personalized medicine by enabling the rapid assessment of drug efficacy in patient-specific models. Here, the integration of advanced techniques supports existing organoid protocols, providing a valuable resource for researchers and clinicians seeking to improve patient outcomes through more efficient and precise organoid-based applications.
Guidelines
Cancerous tumors remain a leading global health threat, with persistently high incidence and mortality rates. Their intrinsic characteristics, such as tumor heterogeneity and dynamic evolution, pose significant challenges for traditional treatments and even genetic sequencing-based precision medicine. Rapid and extensive screening of tumors using patient-derived genetic and epigenetic living models, particularly those generated from ultra-small biopsy samples, is crucial for identifying the most effective treatments and improving patient outcomes. Three-dimensional (3D) organoid models have emerged as powerful tools for replicating the architecture and function of original tissues, providing invaluable platforms for disease modeling, drug screening, and personalized medicine. Organoids derived from patient tumors, known as patient-derived organoids (PDOs), retain the genetic and histological features of the original tumor, enabling more accurate predictions of therapeutic responses. However, the widespread clinical application of organoid technology faces significant challenges. Traditional methods for organoid culture are labor-intensive, time-consuming, and suffer from issues related to scalability and reproducibility. Moreover, handling ultra-small tissue samples, which are common in clinical biopsies, remains particularly problematic due to limited cell numbers and difficulties in maintaining viability. Recent advancements in automated cell extraction, microfluidic organoid formulation, and spatial transcriptomics offer promising solutions to these challenges. Automated tissue dissociation devices enable efficient and reproducible extraction of viable cells from small tissue samples. Microfluidic technologies facilitate the high-throughput formation of uniform organoid microspheres, enhancing scalability and consistency. While single-cell sequencing analysis is rapidly advancing, revealing more about cell-cell communication, the integration of spatial transcriptomics enables the comprehensive characterization of organoids at the single-cell level while preserving spatial context, providing insights into cellular heterogeneity and tumor microenvironment interactions. This protocol presents a comprehensive, high-throughput system for treating, generating, and characterizing patient-derived organoids from ultra-small tissue samples. It integrates automated cell extraction, microfluidic droplet-based organoid printing, and lamination-based spatial transcriptomics. Streamlining organoid workflow presents an opportunity to accelerate the clinical translation of organoid technology, enabling applications such as personalized therapy selection in a shorter timeframe.
Materials
Biological Materials: Human samples were collected after informed written consent was obtained from all donors in accordance with study protocols conforming to the provisions of the Declaration of Helsinki. For the pediatric participants, written informed consent was obtained from adult legal guardians of the children before enrollment. The study was approved by the Ethics Committees of Tsinghua Shenzhen International Graduate School (Refs. 202171 and 202295). Mouse: Male BALB/c mice (4-6 weeks old) were purchased from the Medical Laboratory Animal Center of Guangdong, China, and housed in the animal facility of Tsinghua University Shenzhen International Graduate School (SIGS). Environmental conditions were maintained at a temperature of 23±2°C, with a relative humidity of 50-65%, in a pathogen-free environment, under a 12-hour light/12-hour dark cycle, with ad libitum access to food and water. Human: Patient-derived colorectal cancer and liver cancer samples were obtained from Shenzhen People's Hospital, Shenzhen, China (Ethical Development No. 2024-08401). Reagents: Tissue digestion solution: Collagenase Ⅰ (Yuanye, cat. no.S10053-1g), Collagenase Ⅳ (Yuanye, cat. no. S10056-1g), Dispase Ⅱ (Yuanye, S25046-1g), DNase (Solarbio, D8071), DPBS (Biosharp, cat. no.BL310A), FBS (QmSuero, mu001SR), P/S (Biosharp, BL505A), AO/PI (Countstar, cat. no. RE010212). Organoid formulation and culture: 7000 Engineered Fluid (3M Novec, cat. no.25556), MasterAim® organoid complete medium for colorectal cancer (MasterAim, cat. no.10-100-018), MasterAim® organoid complete medium for liver cancer (MasterAim, cat. no.10-100-296). Organoid lamination and characterization: Cell Recovery Solution(Corning, cat. no.354253), Methanol (Rhawn, cat. no.R007536), Trypsin (Gibco, cat. no.15050057), PTFE (Millipore, cat. no.IPVH00010), BSA (for non-sterile use) (Biosharp , cat. no. BS114-100g ), DAPI (Biosharp, cat. no.BL739A), AntiFade Mounting Medium (Beyotime, cat. no.P0131), 4% Paraformaldehyde (PFA) (Biosharp, cat. no. BL539A), PBS (nonsterile) (Biosharp, cat. no. BL601A), Sucrose (Solarbio, cat. no. S8271), Tissue-Tek O.C.T. compound (Biosharp, cat. no. BL557A), TritonX-100 (Solarbio, cat. no. T8200), Tween20 (Leagene, cat. no. PW0026), CEA (Huabio, cat. no.ER1906-09), Lysozyme (Huabio, cat. no.ET1609-35), MUC1 (Huabio, cat. no. ET1611-14), FAPB1 (Huabio, cat. no.ET1704-23), RDX (Huabio, cat. no. ET1610-41), AFP (Huabio, cat. no. EM1701-31), OPN (Huabio, cat. no. EM1701-31). Drug screening: CellTiter-Glo®Luminescent Cell Viability Assay (Promega, cat. no.G7570), Calcein-AM/PI (Solarbio, cat. no.CA1630), CTG reagent (Promega, cat. no. G7572), 5-FU (Aladdin, cat. no. F100149-1g), Epirubicin (Abmole, cat. no. M5617), Oxaliplatin (MCE, cat. no. HY-17371), Gefitinib (Aladdin, cat. no. G125799), Gemcitabine (Aladdin, cat. no. G127944-5g), Cisplatin (Aladdin, cat. no. HY-17394), Paclitaxel (Aladdin, cat. no. P106869), Anti-PD-1 (Aladdin, cat. no. C412007), Lenvatinib (Aladdin, cat. no. L125518).
Troubleshooting
Development of the Protocol
All three steps are performed using automated devices (Synorg, model. no.P01). Manual sample transport is required between stages and devices.
This protocol builds upon previous advancements in primary tissue cell extraction, organoid microsphere formulation and printing, and organoid profiling. Our focus is on primary tissue or patient-derived organoids, emphasizing high-throughput formation and automated processing to enhance scalabi lity and reproducibility. The protocol includes three key stages (Figure 1):
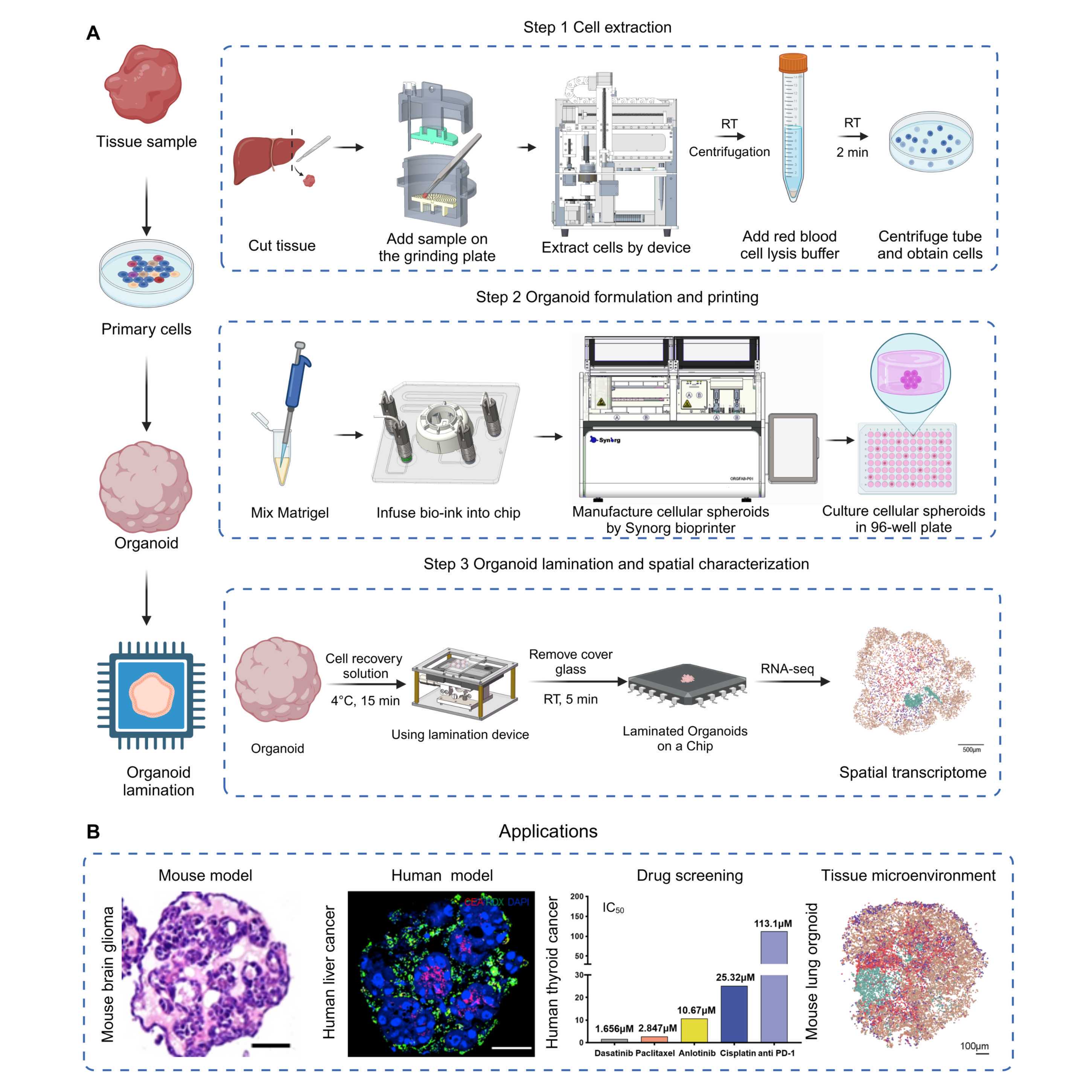
Figure 1. Workflow to formulate and characterize microfluidic droplet-engineered
organoids (DEOs)
A) Step 1. Extract cells from tissue, including using forceps and scalpels to cut tissue into small pieces, grinding tissue cuts using the automated small-Tissue Extraction Device (sTED), red blood cell lysis, and centrifuging cell suspension to obtain cell pellets.
Step 2. Formulate and print DEOs using extracted cells from Step 1, including suspending cells in Matrigel to create bioink, infusing bioink to sampling chip, and installing chip to OrgFab printer to formulate and print DEOs into 96-well plates. Details are described in Box 2.
Step 3. DEO profiling by laminating 3D organoid cells into 2D mono-layer distribution. It includes Matrigel removal by matrix digestion, laminating organoids on a spatial transcriptome chip using a lamination device, and spatial sequencing profiling.
B) Application examples of DEOs.
From left to right: HE staining of mouse brain glioma organoids, fluorescent staining of human liver cancer organoids (red: CEA, green: RDX, blue: DAPI), anticancer drug screening of human thyroid cancer organoids (IC50, the drug concentration at which anticancer drugs inhibit tumor cell activity by 50%), visualization of cell annotation in spatial transcriptomics of a primary lung organoid, scale bar, 100μm.
Extraction and Purification of Primary Cells from Tissue Samples (Figure 2)
Formulation and Bioprinting of Cell-Laden Microspheres (Figure 3)
Lamination of Organoids for Rapid Characterization (Figure 4)
Stage 1: Obtaining Primary Cells from Tissues
30m
Extract cells from primary tissues:
The acquisition of viable primary cells is critical for constructing accurate patient tissue-derived organoid models. Traditional methods often involve manual grinding of tissue followed by enzymatic digestion. However, manual techniques are inherently variable, as the degree of tissue fragmentation and operator proficiency can significantly affect cell viability and yield. For instance, inconsistencies in tissue preparation can impact the establishment and growth of organoids. Moreover, specialized tissues like the liver require perfusion-based extraction methods, which are complex and difficult to standardize. To address these challenges, we employ the automated small-Tissue Extraction Device that standardizes tissue processing (Figure 2D), minimizing operator-dependent variability and enhancing the consistency of primary cell isolation.
▲CRITICAL The cell extraction process must be performed within a sterile laminar flow hood. Prior to
commencing the procedure, the equipment should be sterilized with ultraviolet light.
Figure 2. Workflow to extract cells from primary tissues
A) Select liver tissue from BALB/c mouse
B) Use forceps and sterilized scalpels to cut the primary tissue sample on ice
into small fragments (approximately < 1mm in size)
C) Open the grinding chamber, and add the tissue fragments with forceps
D) Run the grinding device, which includes 6 steps (a-f):
a) Select the grinding mode, and lower the upper grinding plate until the grinding surfaces fuse together
b) Add the grinding buffer, set the lower grinding plate rotating at 60r/min, and start the motor to run for one minute
c) Select the stirring mode and lift the upper grinding plate for 10 mm to separate the grinding plates
d) Add 10× tissue digestion reagent, and rotate the lower grinding plate for 5 min at 60r/min and 37 °C
e) Pass the cell suspension through a 100 μm cell filter
f) Add an equal volume of digestion termination reagent, which contains 10% serum, to the cell suspension
E) Centrifuge the cell suspension, remove the supernatant, add 2 mL of red blood cell lysis solution to the cell pellet, pipette the mixture, and incubate it at room temperature for 2 min
F) Re-centrifuge the cell suspension, remove the supernatant, add 2 mL culture medium to the cell pellet, and pipette the mixture to obtain primary cell suspension
30m
Take out one 60 mm diameter Petri dish, add 2 mL of DPBS (add 2% P/S), and place the tissue block to be processed into the dish.
Wash the tissue 3 times with DPBS (add 2% P/S).
Use forceps and sterilized scalpels to cut the primary tissue sample on ice into small fragments (approximately < 10cm in size).
!CAUTION Avoid excessive cutting during the cutting process.
Open the grinding chamber, and add the tissue fragments with forceps
Run the grinding device, which includes 6 automated steps (a-f):
a) Lower the upper grinding plate until the grinding surfaces fuse together.
b) Add the grinding buffer, set the lower grinding plate rotating at 60 r/min, and start the motor to run for 1 min.
c) Lift the upper grinding plate for 10 mm to separate the grinding plates.
d) Add 10× tissue digestion reagent, and rotate the lower grinding plate for 5min at 60 r/min and 37 °C.
e) Pass the cell suspension through a 100 μm cell filter.
f) Add an equal volume of digestion termination reagent, which contains 10% serum, to the cell suspension.
!CAUTION Close the cover during operation and be cautious of mechanical hazards.
Centrifuge the cell suspension and remove the supernatant.
!CAUTION When aspirating the supernatant using a pipette, avoid disturbing the cell pellet.
Add 2 mL of red blood cell lysis solution to the cell pellet, pipette the mixture, and incubate it at room temperature for 2 min.
Re-centrifuge the cell suspension, remove the supernatant, add 2 mL culture medium to the cell pellet, and pipette the mixture to obtain primary cell suspension.
Thaw the AO/PI reagent and prepare a premix solution containing 50 µg/mL PI and 10 µg/mL AO in DPBS.
!CAUTION Avoid contact of AO/PI dye with skin.
Resuspend the cells in 1 mL of basal culture medium and mix thoroughly by pipetting up and down.
Aspirate 10 µL of the cell suspension and 10 µL of the AO/PI working solution, mix them together, and then add the mixture to a hemocytometer for cell counting. Calculate the cell number.
Stage 2: Constructing Primary Cell-Based In Vitro 3D Cell Microspheres
1w 6d 1h
Formulate and print organoid precursor, using the OrgFab printer:
The streamlined, automated process not only improves the spatial precision and reproducibility of organoid precursor fabrication but also enhances scalability for high-throughput applications in drug screening and disease modeling. At this stage, we focus on the automated fabrication of primary cell-based in vitro 3D microspheres using an integrated microfluidic bioprinting system. This approach addresses the challenges of scalability, consistency, and handling ultra-small sample sizes in organoid production. Building upon advancements reported in our previous work on the OrgFab printer, primary cells are combined with Matrigel to form a bioink, which is then processed using a microfluidic technique employing oil-phase shearing to generate uniform, multi-cell-laden droplets. These droplets undergo online Matrigel solidification, ensuring immediate stabilization of the organoid precursors. Subsequently, a 3D bioprinting nozzle precisely deposits the solidified microspheres onto multi-well plates, our target substrates for further culture (Figure 3D). The integration of microfluidics and bioprinting technologies allows for high spatial precision in droplet formation and placement, significantly improving the reproducibility of organoid fabrication. The streamlined process not only enhances spatial precision but also improves scalability for high-throughput applications in drug screening and disease modeling. The OrgFab system is capable of handling ultra-small sample volumes down to 5 μL, producing more than 100 organoid precursors from just 10 μL of bioink. For example, this is valuable for needle biopsy samples. The fully automated workflow minimizes manual intervention, reducing variability and enhancing reproducibility. The OrgFab printer integrates a computer vision-based bioprinting module, which enables accurate detection and deposition of organoid precursors into multi-well plates. This precision facilitates downstream high-throughput assays and ensures consistency across experiments without requiring an operator. Additionally, the high-density microfluidic droplet encapsulation used in this method has been shown to accelerate organoid maturation due to increased paracrine signaling among densely packed cells. This results in more rapid organoid development and the potential for earlier experimental readouts, which is advantageous in time-sensitive research and clinical contexts. By integrating these techniques, our protocol tries to address the key limitations of existing methods, such as low throughput and limited automation. It provides an integrated platform for the controllable modeling of organoids, with promising developments for personalized medicine, drug discovery, and disease modeling.
▲CRITICAL The entire process should be placed and operated within a biosafety cabinet to prevent bacterial contamination during use.
Figure 3. Workflow to formulate and print organoid precursor, i.e. cellular spheroids, using the OrgFab printer
A) Centrifuge the cell suspension and remove the supernatant to obtain a cell pellet
B) Use a pre-cooled pipette tip to aspirate Matrigel on ice, pipette it to the EP tube, and mix with cells by gently pipetting
C) Manual preparation steps before using the OrgFab printer (a-e):
a) Turn on UV sterilization for 20 min
b) Install the microfluidic chip to the semiconductor cooling platform, and connect the inlet/outlet ports to corresponding fluids, including two oil inlet ports, one bioink inlet port, and one droplet outlet port
c) Insert bioink inlet tubing into the bioink EP tube, aspirate oil out of the channel from the front right (Oil inlet 1) to port aspirate bioink with flow rates of 250uL/min from the tube, and infuse the channel
d) Install the print head, and connect one end of the print head to the collection/incubation tubing and the other end to the high-pressure gas. The third port is placed facing straight to the recipient substrate, e.g. 96-well plate
e) Place the EP tube with bioink in the sample tank and insert the bioink tube
D) OrgFab printer operation, which includes 8 automated steps (a-h):
a) Move down the mechanical turntable to fix the chip on the semiconductor cooling platform
b) Insert the chip bioink inlet tubing into the EP tube for waste collection, and pump oil into the chip channels with flow rates of 312.5uL/min via the two oil inlets to repel air in the channel. Afterward, the chip channels are filled with oil. All fluids are conducted via PTFE tubing
c) Insert bioink inlet tubing into the bioink EP tube, aspirate bioink from the tube, and infuse the channel, meanwhile, perfuse oil out of the channel segment connecting to the front right (inlet) port
d) Rotate the mechanical turntable (knob) to change the chip channel tandem, which connects the bioink channel to the front left port for the bioink outlet
e) Pump oil via the two oil inlets, with flow rates of 312.5uL/min in Oil inlet 2 and 25uL/min in Oil inlet 1
f) Shear bioink flow into uniform droplets by oil co-flow, forming structural templates of organoid precursors (or cellular spheroids)
g) Stop pumping oil toward the two inlet channels, store the cell-laden droplets in the collection tubing, and solidify them at 37°C for 30 min
h) Pump oil from Oil inlet 1 at 12.5uL/min and Oil inlet 2 at 25uL/min, use a camera and visual detection algorithm to control the instantaneous opening of the high-pressure gas toward the print head. Each air beam firing enables one shot of gelled droplet, i.e. organoid precursor/cellular spheroid, into the 96-well plate, with each well containing one or a certain number of spheroid
E) Culture organoid precursors/cellular spheroids in a 96-well plate
F) Picture of the consumable microfluidic chip module
G) Picture of organoid precursors after printing in a 96-well plate
H) Violin plot of area and roundness of printed organoids.
1h
Centrifuge the cell suspension and remove the supernatant to obtain a cell pellet.
Place the cell pellet on ice to pre-chill for 1 min.
Use a 100μl pre-cooled pipette tip to aspirate Matrigel on ice, pipette it to the EP tube, and mix with cells by gently pipetting.
▲CRITICAL STEP The uniformity of the mixture of cells with Matrigel determines the consistency of the organoids.
!CAUTION During the mixing process, maintain a low temperature at all times and avoid touching the EP tube with hands.
!CAUTION The Matrigel should not be subjected to repeated freeze-thaw cycles.
!CAUTION If the matrix gel that has been mixed with cells cannot be used immediately, it should be stored in a refrigerator at 4℃ and used within 2-4 hours.
Turn on the sterilization switch of the equipment and activate the UV sterilization for 20 min.
Install the microfluidic chip to the semiconductor cooling platform, and connect the inlet/outlet ports to
corresponding fluids, including two oil inlet ports, one bioink inlet port, and one droplet outlet port.
!CAUTION Ensure that the chip alignment platform is properly aligned to avoid gaps.
Install the collection tubing in the heating chamber, connect the collection port to the droplet formulation outlet port, and connect the other end to the port on the print head.
▲CRITICAL STEP When connecting the PTFE tubing to the chip component, after securing the tube, confirm whether the tube is in contact with the bottom of the chip.
!CAUTION During the connection process, do not bend the PTFE tubing.
Install the print head, and connect one end of the print head to the collection/incubation tubing and the other end to the high-pressure gas. The third port is placed facing straight to the recipient
substrate, e.g. 96-well plate.
!CAUTION Before installation, confirm the orientation of the print head to ensure proper alignment and maintain airtight connections.
Install the cell culture plate and the waste liquid collection trough on the printing platform on the right side of the device.
!CAUTION Place the culture plate on the plate tray and secure the plate to the tray by aligning and locking it into position.
Place the EP tube with bioink in the sample tank and insert the bioink tubing.
!CAUTION During the installation process, handle the cap of the EP tube and avoid contact with the part of the tube containing the bioink.
Initiating OrgFab printer, which includes 8 automated steps (a-h):
!CAUTION Before starting, ensure that the temperature displayed on the software is set to 4°C.
!CAUTION Close the chamber lid when starting the printing process and pay attention to mechanical safety.
!CAUTION During the operation of the equipment, ensure that the ventilation function of the biosafety cabinet is activated. The temperature should be maintained between 20-25℃.
a) Move down the mechanical turntable to fix the chip on the semiconductor cooling platform.
b) Insert the chip bioink inlet tubing into the EP tube for waste collection, and pump oil into the chip channels with flow rates of 312.5uL/min via the two oil inlets to repel air in the channel. Afterward, the chip channels are filled with oil. All fluids are conducted via PTFE tubing.
!CAUTION Nove7000 is volatile. After adding it, the cap should be closed promptly.
c) Insert bioink inlet tubing into the bioink EP tube, aspirate oil out of the channel from the front right (Oil inlet 1) to port aspirate bioink with flow rates of 250uL/min from the tube, and infuse the channel.
d) Rotate the mechanical turntable (knob) to change the chip channel tandem, which connects the bioink channel to the front left port for the Bioink outlet.
e) Pump oil via the two oil inlets, with flow rates of 312.5uL/min in Oil inlet 2 and 25uL/min in Oil inlet 1.
f) Shear bioink flow into uniform droplets by oil co-flow, forming structural templates of organoid precursors (or cellular spheroids).
!CAUTION During the initial microsphere preparation, observe the morphology and uniformity of the microspheres. If any abnormalities are detected, click the interrupt button to stop the process and
adjust the ratio of the oil-pushing speed.
g) Stop pumping oil toward the two inlet channels, store the cell-laden droplets in the collection tubing, and solidify them at 37°C for 30 min.
h) Pump oil from Oil inlet 1 at 12.5uL/min and Oil inlet 2 at 25uL/min, use a camera and visual detection algorithm to control the instantaneous opening of the high-pressure gas toward the print head. Each air beam firing enables one shot of gelled droplet, i.e. organoid precursor/cellular spheroid, into the 96-well
plate, with each well containing one or a certain number of spheroid.
Culture organoid precursors/cellular spheroids in a 96-well plate.
Organoid culture:
▲CRITICAL The entire process should be placed and operated within a biosafety cabinet to prevent bacterial contamination during use.
1w
Open the organoid printer cover and take out the organoid printed on the 96-well plate.
Place the plate under a microscope and observe the morphology of the organoid to check the morphology of the organoid and make sure each well of the plate has 1 organoid.
Place the culture plate in a 37°C CO2 incubator for culture.
!CAUTION Pay attention to the humidity and CO2 concentration of the incubator, and avoid vibrating the plate.
Change the medium every other day, until organoids fully grow for 7-14 d.
Remove 50 ml of culture medium each time, and then add new culture medium to 100 ml.
!CAUTION Be careful not to suck up the organoids when changing the culture medium. Tilting the 96-well plate will make it easier to observe.
Drug screening:
▲CRITICAL Choose a light-protected plate when culturing the organoids.
6d
For the drug to be tested, prepare a 50 mM drug stock solution and filter it through a membrane. Aliquot 5 μL per tube.
After culturing for 7-14 days (depending on the organoid type), place the culture plate under the microscope to observe the organoid structure.
Dilute the drug to the highest concentration and then perform a 5-fold gradient dilution. Calculate the volume of drug and culture medium required for each gradient.
Use a 1000 μL pipette to add the culture medium in a 15 ml centrifuge tube, and then add the
drug stock solution.
Use a 100 μL pipette to aspirate and remove the culture plate medium completely.
Add the drug-containing culture medium to the plate wells in ascending order of drug concentration.
!CAUTION For each concentration, set up three replicate wells, and also reserve one additional well for live/dead staining at each concentration.
Replace the culture medium with a drug-containing medium to prevent drug degradation every other day.
After culture 5 days, thaw the CTG reagent at room temperature and equilibrate the CTG reagent.
!CAUTION Do not vortex the reagents.
After culture 6 days, Thaw the Calcein-AM/PI reagent. In a light-protected EP tube, sequentially add 900 μL ddH₂O, 100 μL 10× Assay Buffer, 1 μL Calcein-AM Solution (2 mM), and 3 μL PI Solution (1.5 mM). Mix them and set aside for use.
Using a 100 μL pipette, aspirate the culture medium from the live/dead staining wells. Wash
with DPBS, then add the Calcein-AM/PI working solution.
Stain at room temperature in the dark for 5 minutes. Then, using a 100 μL pipette, aspirate the
staining solution, wash with DPBS, and observe under a fluorescence microscope for imaging.
Add 100 μL of CTG reagent to each well (maintaining consistency with the culture medium volume).
Place the culture plate on a shaker and shake for 2 minutes. Use a microscope to observe whether the organoid spheroids have dispersed. If not dispersed, use a multichannel pipette to pipette up and down 10-15 times, and then recheck under the microscope to ensure the spheroids are dispersed.
!CAUTION When using a multichannel pipette to blow and mix, be careful to avoid creating bubbles.
Incubate at room temperature for 10 minutes. Record the luminescence (LUM) signal intensity using a microplate reader and construct an IC50 curve.
Stage 3: Laminating Organoids for Rapid Characterization
30m
Laminate organoids on the chip:
Traditional methods for organoid characterization, such as sectioning, immunostaining, and even more recent single-cell sequencing, are labor-intensive and can result in cell loss and incomplete spatial information. Recent advancements in spatial transcriptomics have enabled high-resolution mapping of gene expression within tissues and organoids. However, applying these techniques to primary tissue-derived organoids remains challenging due to their small size and low cell count, which limits the feasibility of tissue sectioning as a pre-processing step before profiling. Based on these limitations and advancements, we developed a lamination device that compresses organoids into a single-cell-layer morphology without disrupting cellular architecture (Figure 4, Figure 7). This lamination technique allows projecting 3D organoid spheres onto a 2D plane, preserving spatial relationships between cells. By integrating lamination, which enables spatial transcriptomics profiling, our protocol facilitates the comprehensive characterization of organoids at the single-cell level, followed by immunofluorescence staining. This approach is particularly valuable for understanding cellular heterogeneity and interactions within tumor organoids.
Figure 4. Workflow to laminate organoids on the chip
A) Select and pick out organoids using a Pasteur pipette under the guidance of a microscope
B) Clean the culture medium by repeated washing with 1× SSC buffer, add cell recovery solution for the Organoid, and incubate it at 4°C for 15min
C) Wash the cell recovery solution for Organoid with 1× SSC buffer, add cell dissociation reagent, and incubate at room temperature for 2min
D) Use the lamination device to create 2D cell distribution, which includes 8 manual steps (a-h):
a) Use a Pasteur pipette to load the organoid on the chip
b) Use a gas cylinder to blow and dry the liquid around the organoid
c) Use tweezers to cover the chip with a piece of PTFE film
d) Install the chip on the lamination device
e) Place a glass slide on the laminating device and place a weight on top of the glass slide
f) Switch the knob to lift the chip until it touches the glass slide installed on the top
g) Gently and vertically remove the weight and then the glass slide
h) Uninstall the chip from the lamination device, heat the chip at 37°C for 5 min, and use tweezers to remove the PTFE film
E) Immunofluorescence staining of a laminated organoid, showing DAPI staining of the laminated cellular spheroid. Scale bar=100μm
F) Laminated organoids on a spatial transcriptomic chip, enabling sequencing and resolution of cell types and cellular interactions
30m
Place the cell culture plate under the microscope.
Select and pick out organoids using a Pasteur pipette under the guidance of a microscope.
Aspirate the culture medium from the wells using a pipette and wash with 1× SSC buffer.
!CAUTION When using the pipette, take care not to touch the organoids.
Add cell recovery solution for the organoid and incubate it at 4°C for 15min.
Wash the cell recovery solution for organoid with 1× SSC buffer, add cell dissociation
reagent, and incubate at room temperature for 2 min.
Place a drop of ddH₂O on the microscope slide and adhere the chip to the slide.
!CAUTION If lamination is performed on the microscope slide, this step is not required.
Use a Pasteur pipette to load the organoid on the chip/microscope slide.
!CAUTION When using a Pasteur pipette, take care not to touch the surface of the chip.
Use a gas cylinder to blow and dry the liquid around the organoid.
▲CRITICAL STEP Remove the liquid to ensure that there is no slipping during the lamination process.
!CAUTION When using a gas cylinder, pay attention to the gas flow rate to avoid blowing away the organoids.
!CAUTION The surface of the organoids needs to be dried as much as possible, but complete desiccation should be avoided. A thin layer of water film should be retained on the surface of the organoids.
Use scissors to cut the PTFE film into a size of 3 cm × 3 cm.
!CAUTION During the cutting, avoid the creased sections of the PTFE film.
Use tweezers to cover the chip with a piece of PTFE film.
!CAUTION Avoid the formation of wrinkles during the covering process. If the cover is misaligned, do not lift it to reposition.
!CAUTION This step should be completed as quickly as possible to avoid the disappearance of the water film on the organoids.
Install the microscope slide on the lamination device.
Place another glass slide on the lamination device and place a weight on top of the glass slide
!CAUTION When installing another microscope slide, align the edge of the slide with the raised platform on the device.
!CAUTION Before installing the glass slide, lower the manual height adjustment stage to its lowest position to avoid contact between the glass slide and the PTFE film.
Switch the knob to lift the chip until it touches the glass slide installed on the top.
!CAUTION The upper microscope slide should be detached from the boss and maintained in this position for 10s.
Remove the weight and the glass slide gently and vertically in sequence.
▲CRITICAL STEP When removing the weights and the microscope slide, avoid any movement or sliding of the PTFE film.
Uninstall the chip from the lamination device, heat the chip at 37°C for 5min, and use
tweezers to remove the PTFE film.
▲CRITICAL STEP Use tweezers to lift the PTFE film from one side, ensuring it does not slip.
Immunofluorescence staining of a laminated organoid, showing DAPI staining of the laminated cellular spheroid.
Laminated organoids on a spatial transcriptomic chip, enabling sequencing and resolution of cell types and cellular interactions.
Protocol references
REFERENCES
1 Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a cancer journal for clinicians 74, 229-263 (2024).
2 Tuveson, D. & Clevers, H. Cancer modeling meets human organoid technology. Science 364, 952-955 (2019).
3 Thorel, L. et al. Patient-derived tumor organoids: a new avenue for preclinical research and precision medicine in oncology. Experimental & Molecular Medicine 56, 1531-1551 (2024).
4 Kim, J., Koo, B.-K. & Knoblich, J. A. Human organoids: model systems for human biology and medicine. Nature Reviews Molecular Cell Biology 21, 571-584 (2020).
5 Zhao, Z. et al. Organoids. Nature Reviews Methods Primers 2, 94 (2022).
6 Lancaster, M. A. & Knoblich, J. A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345, 1247125 (2014).
7 Clevers, H. Modeling development and disease with organoids. Cell 165, 1586-1597 (2016).
8 Ma, S. et al. Lamination-based organoid spatially resolved transcriptomics technique for primary lung and liver organoid characterization. Proceedings of the National Academy of Sciences 121, e2408939121 (2024).
9 Jiang, S. et al. An automated organoid platform with inter-organoid homogeneity and inter-patient heterogeneity. Cell Reports Medicine 1 (2020).
10 Bose, S., Clevers, H. & Shen, X. Promises and challenges of organoid-guided precision medicine. Med 2, 1011-1026 (2021).
11 Hofer, M. & Lutolf, M. P. Engineering organoids. Nature Reviews Materials 6, 402-420 (2021).
12 Garreta, E. et al. Rethinking organoid technology through bioengineering. Nature materials 20, 145-155 (2021).
13 Wang, X., Luo, Y., Ma, Y., Wang, P. & Yao, R. Converging bioprinting and organoids to better recapitulate the tumor microenvironment. Trends in biotechnology 42, 648-663 (2024).
14 Cho, S., Discher, D. E., Leong, K. W., Vunjak-Novakovic, G. & Wu, J. C. Challenges and opportunities for the next generation of cardiovascular tissue engineering. Nature Methods 19, 1064-1071 (2022).
15 Mateo, J. et al. Delivering precision oncology to patients with cancer. Nature medicine 28, 658-665 (2022).
16 Hofman, P. The challenges of evaluating predictive biomarkers using small biopsy tissue samples and liquid biopsies from non-small cell lung cancer patients. Journal of thoracic disease 11, S57 (2019).
17 Zeng, G. et al. Advancing cancer research through organoid technology. Journal of Translational Medicine 22, 1007 (2024).
18 Zuo, J., Fang, Y., Wang, R. & Liang, S. High-throughput solutions in tumor organoids: from culture to drug screening. Stem Cells, sxae070 (2024).
19 Liu, X. et al. Tumor-on-a-chip: from bioinspired design to biomedical application. Microsystems & Nanoengineering 7, 50 (2021).
20 Wahle, P. et al. Multimodal spatiotemporal phenotyping of human retinal organoid development. Nature Biotechnology 41, 1765-1775 (2023).
21 Kishi, J. Y. et al. Light-Seq: light-directed in situ barcoding of biomolecules in fixed cells and tissues for spatially indexed sequencing. Nature Methods 19, 1393-1402 (2022).
22 Ooft, S. N. et al. Patient-derived organoids can predict response to chemotherapy in metastatic colorectal cancer patients. Science translational medicine 11, eaay2574 (2019).
23 Millen, R. et al. Patient-derived head and neck cancer organoids allow treatment stratification and serve as a tool for biomarker validation and identification. Med 4, 290-310. e212 (2023).
24 Lombardo, J. A., Aliaghaei, M., Nguyen, Q. H., Kessenbrock, K. & Haun, J. B. Microfluidic platform accelerates tissue processing into single cells for molecular analysis and primary culture models. Nature communications 12, 2858 (2021).
25 Van de Wetering, M. et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 161, 933-945 (2015).
26 Yang, H. et al. Ultra-small tissue-compatible organoid printer for rapid and controllable modeling of respiratory organoids. Device (2024).
27 Heumos, L. et al. Best practices for single-cell analysis across modalities. Nature Reviews Genetics 24, 550-572 (2023).
28 Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell–cell communication from single-cell transcriptomics. Nature Protocols 20, 180-219 (2025).
29 Wälchli, T. et al. Shaping the brain vasculature in development and disease in the single-cell era. Nature Reviews Neuroscience 24, 271-298 (2023).
30 Moncada, R. et al. Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nature biotechnology 38, 333-342 (2020).
31 Meran, L., Tullie, L., Eaton, S., De Coppi, P. & Li, V. S. Bioengineering human intestinal mucosal grafts using patient-derived organoids, fibroblasts and scaffolds. Nature Protocols 18, 108-135 (2023).
32 Liu, L., Yu, L., Li, Z., Li, W. & Huang, W. Patient-derived organoid (PDO) platforms to facilitate clinical decision making. Journal of Translational Medicine 19, 1-9 (2021).
33 Cordts, S. C. et al. Microdissection tools to generate organoids for modeling the tumor immune microenvironment. Microsystems & Nanoengineering 10, 126 (2024).
34 Zhou, C. et al. Standardization of organoid culture in cancer research. Cancer Medicine 12, 14375-14386 (2023).
35 Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nature protocols 11, 1724-1743 (2016).
36 Driehuis, E., Kretzschmar, K. & Clevers, H. Establishment of patient-derived cancer organoids for drug-screening applications. Nature protocols 15, 3380-3409 (2020).
37 Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299-312 (2015).
38 Ko, J. et al. Microfluidic high-throughput 3D cell culture. Nature Reviews Bioengineering 2, 453-469 (2024).
39 Zhang, Y. S. et al. 3D extrusion bioprinting. Nature Reviews Methods Primers 1, 75 (2021).
40 Brassard, J. A., Nikolaev, M., Hübscher, T., Hofer, M. & Lutolf, M. P. Recapitulating macro-scale tissue self-organization through organoid bioprinting. Nature Materials 20, 22-29 (2021).
41 Schuster, B. et al. Automated microfluidic platform for dynamic and combinatorial drug screening of tumor organoids. Nature communications 11, 5271 (2020).
42 Mekhileri, N. et al. Automated 3D bioassembly of micro-tissues for biofabrication of hybrid tissue engineered constructs. Biofabrication 10, 024103 (2018).
43 Rumker, L. Modeling Inter-Individual Variation in Single-Cell Datasets to Detect Cell State Abundance Associations to Clinical Features and Genetic Variants, Harvard University, (2023).
44 Dekkers, J. F. et al. High-resolution 3D imaging of fixed and cleared organoids. Nature protocols 14, 1756-1771 (2019).
45 Liao, J., Lu, X., Shao, X., Zhu, L. & Fan, X. Uncovering an organ’s molecular architecture at single-cell resolution by spatially resolved transcriptomics. Trends in biotechnology 39, 43-58 (2021).
46 Harrison, S. P. et al. Scalable production of tissue-like vascularized liver organoids from human PSCs. Experimental & Molecular Medicine 55, 2005-2024 (2023).
47 Beghin, A. et al. Automated high-speed 3D imaging of organoid cultures with multi-scale phenotypic quantification. Nature Methods 19, 881-892 (2022).
48 Bozal, S. B. et al. Development of an automated 3D high content cell screening platform for organoid phenotyping. SLAS Discovery 29, 100182 (2024).
49 Liu, H. et al. A droplet microfluidic system to fabricate hybrid capsules enabling stem cell organoid engineering. Advanced science 7, 1903739 (2020).
50 Sanchez Noriega, J. L. et al. Spatially and optically tailored 3D printing for highly miniaturized and integrated microfluidics. Nature communications 12, 5509 (2021).
51 Lamanna, J. et al. Digital microfluidic isolation of single cells for-Omics. Nature communications 11, 5632 (2020).
52 Boya, M. et al. High throughput, label-free isolation of circulating tumor cell clusters in meshed microwells. Nature communications 13, 3385 (2022).
53 Mathur, L. et al. Combi-seq for multiplexed transcriptome-based profiling of drug combinations using deterministic barcoding in single-cell droplets. Nature communications 13, 4450 (2022).
54 Synorg Biotechnology Shenzhen Co. Ltd. Product details., (2025).
55 RJMART, C. Product page for Orgfab‱ Organoid 3D Printer P01. RJMART.cn. , <https://www.rjmart.cn/detail?productId=200437242706&suppId=132526&source=2> (2025).
56 Morse, D. B. et al. Positional influence on cellular transcriptional identity revealed through spatially segmented single-cell transcriptomics. Cell systems 14, 464-481. e467 (2023).
Acknowledgements
The work was supported by the National Key-Area Research and Development Program of China (2024YFA0919800); National Natural Science Foundation of China (32371470 and 82341019); Merck Research Grant, and the Cross-disciplinary Research and Innovation Fund of Tsinghua SIGS (No. JC2022007); Guangdong Basic and Applied Basic Research Foundation (2023B1515120025); Shenzhen Fundamental Research Program (No. JCYJ20240813112004006); Shenzhen Major Science and Technology Projects (KJZD20230923115400001).


