Oct 27, 2025
- Amy Lyden1,
- Gloria Castañeda1,
- Emily Crawford1,
- Saharai Caldera2,
- Jenai Quan1,
- Katherine Travisano1,
- Prince Nijebu1
- 1CZ Biohub;
- 2CZ Biohub, UCSF

Protocol Citation: Amy Lyden, Gloria Castañeda, Emily Crawford, Saharai Caldera, Jenai Quan, Katherine Travisano, Prince Nijebu 2025. FLASH v2.0. protocols.io https://dx.doi.org/10.17504/protocols.io.dm6gpmmm8gzp/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Other
We attempted this protocol but could not get it to work in our workspace
Created: October 25, 2025
Last Modified: October 27, 2025
Protocol Integer ID: 230785
Keywords: crispr, sequencing, antibiotic resistance, targeted crispr, guide rna design help, flash, rna, flash paper, cas9 technology, cas9 library, sequencing library, downstream adaptor ligation, amendable for downstream adaptor ligation, dna, dna fragment, phosphoryl groups of the dna fragment, cas9, targeted sequence, initial dna sample
Abstract
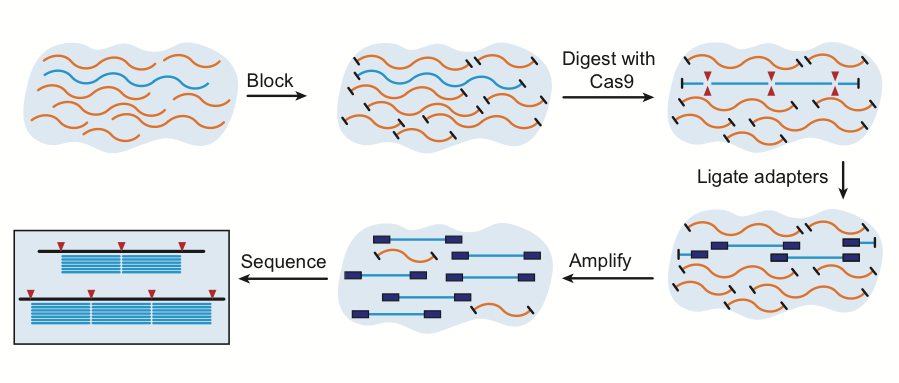
FLASH workflow
FLASH is a crispr-cas9 technology that enriches for targeted sequences in sequencing libraries. The initial DNA sample undergoes a blocking step that removes of the 5’ phosphoryl groups of the DNA fragments, resulting in a product that is not amendable for downstream adaptor ligation or amplification via standard Illumina-based library preparation. The subsequent incorporation of targeted CRISPR-cas9 library exposes the desired regions of interest, allowing them to be processed into a library. For more information on methods and results, please see the FLASH paper. For FLASH guide RNA design help, please see our github.
Guidelines
- This protocol has been used most successfully with starting inputs of DNA ranging from 10 pg – 100 ng. Limited success may be achieved with as little as 100 fg DNA.
- Keep all enzymes on a chilled enzyme block. Immediately before use, allow them to come to room temperature for 5 minutes and then vortex briefly.
- The NEBNext® dA-Tailing Module includes the Klenow fragment and the dA-Tailing buffer. The Klenow fragment allows for the 5’ ends to remain dephosphorylated after the initial blocking step. Do NOT use the NEBNext Ultra II End Prep kit for this step.
- AmpureXP beads or other magnetic beads may be used instead of SPRI beads. Adjust the ratios of beads:sample accordingly to ensure proper removal of unwanted products. Different beads yield varying size selection cut-offs. Refer to this table to adjust ratios according to desired effect.
| SPRI Beads : sample ratio | Step(s) | Desired cut-off/effect | |
| 1.7 : 1 | Post-FLASH cleanup | No size selection; cleanup of unwanted buffers and deactivated cas9 protein from reaction | |
| 1 : 1 | Post-adaptor ligation clean up | Removal of buffers AND Stringent removal of all adaptor dimers | |
| 1: 0.9 | post-indexing Q5 PCR cleanup | Removal of buffers AND Stringent removal of all primer dimers | |
| 1: 1 | post-KAPA amplification cleanup | Removal of buffers AND Stringent removal of all primer dimers |
SPRI bead cut-offs
Materials
MATERIALS
NEBNext dA-Tailing Module - 100 rxnsNew England BiolabsCatalog #E6053L
Sodium Orthovanadate (Vanadate) - 1 mlNew England BiolabsCatalog #P0758S
USER Enzyme - 250 unitsNew England BiolabsCatalog #M5505L
NEBNext Adaptor for IlluminaNew England BiolabsCatalog #E7337 in Kits E7335, E7500, E771
NEBNext Ultra II Ligation Module - 96 rxnsNew England BiolabsCatalog #E7595L
NEBNext Ultra II Q5 Master Mix - 250 rxnsNew England BiolabsCatalog #M0544L
Qubit dsDNA HS Assay kit Thermo Fisher ScientificCatalog #Q32854
Nuclease-free water AmbionCatalog #AM9932
Bioanalyzer chips and reagents (DNA High Sensitivity kit)Agilent Technologies
PCR Thermocycler
TruSeq i7/i5 Indexing Primers - Custom (or NEBNext® Multiplex Oligos for Illumina)New England BiolabsCatalog #E7500L
Proteinase KNew England BiolabsCatalog #P8107S
SPRI beads (homemade) or Ampure XP beads
Kapa HiFi Real-Time Amplification KitKapa BiosystemsCatalog #KK2702
Magnetic rack for PCR strips
rAPID alkaline phosphatase enzyme and bufferMerck MilliporeSigma (Sigma-Aldrich)Catalog #4898133001
cas9 4µM or higher concentration
Dual guide RNAs (4µM - targeted to genes or regions to be depleted - crisprRNA and tracr RNA - quantified by RNA Qubit)
Illumina P5 and P7 primers 5uM combined; P5: 5’ AATGATACGGCGACCACCGAGATCT P7: 5’ CAAGCAGAAGACGGCATACGAGAT
10X Cas9 Activity Buffer (500mM Tris pH 8.0 - 1M NaCl - 100mM MgCL2 - 10mM TCEP)
STEP MATERIALS
rAPID alkaline phosphatase enzyme and bufferMerck MilliporeSigma (Sigma-Aldrich)Catalog #4898133001
Dual guide RNAs (4µM - targeted to genes or regions to be depleted - crisprRNA and tracr RNA - quantified by RNA Qubit)
Proteinase KNew England BiolabsCatalog #P8107S
SPRI beads (homemade) or Ampure XP beads
80% Ethanol
TruSeq i7/i5 Indexing Primers - Custom (or NEBNext® Multiplex Oligos for Illumina)New England BiolabsCatalog #E7500L
SPRI beads (homemade) or Ampure XP beads
80% Ethanol
NEBNext Ultra II Q5 Master Mix - 250 rxnsNew England BiolabsCatalog #M0544L
USER Enzyme - 250 unitsNew England BiolabsCatalog #M5505L
SPRI beads (homemade) or Ampure XP beads
80% Ethanol
Kapa HiFi Real-Time Amplification KitKapa BiosystemsCatalog #KK2702
Illumina P5 and P7 primers 5uM combined; P5: 5’ AATGATACGGCGACCACCGAGATCT P7: 5’ CAAGCAGAAGACGGCATACGAGAT
SPRI beads (homemade) or Ampure XP beads
80% Ethanol
Qubit 1X dsDNA High Sensitivity Assay KitThermo Fisher ScientificCatalog #Q33230
Bioanalyzer chips and reagents (DNA High Sensitivity kit)Agilent Technologies
Sodium Orthovanadate (Vanadate) - 1 mlNew England BiolabsCatalog #P0758S
Klenow Fragment (3'-5' exo-) - 1,000 unitsNew England BiolabsCatalog #M0212L
NEBNext Adaptor for IlluminaNew England Biolabs
NEBNext Ultra II Ligation Module - 96 rxnsNew England BiolabsCatalog #E7595L
cas9 4µM or higher concentration
10X Cas9 Activity Buffer (500mM Tris pH 8.0 - 1M NaCl - 100mM MgCL2 - 10mM TCEP)
Protocol materials
rAPID alkaline phosphatase enzyme and bufferMerck MilliporeSigma (Sigma-Aldrich)Catalog #4898133001
Dual guide RNAs (4µM - targeted to genes or regions to be depleted - crisprRNA and tracr RNA - quantified by RNA Qubit)
cas9 4µM or higher concentration
10X Cas9 Activity Buffer (500mM Tris pH 8.0 - 1M NaCl - 100mM MgCL2 - 10mM TCEP)
Proteinase KNew England BiolabsCatalog #P8107S
SPRI beads (homemade) or Ampure XP beads
Klenow Fragment (3'-5' exo-) - 1,000 unitsNew England BiolabsCatalog #M0212L
NEBNext Adaptor for IlluminaNew England Biolabs
NEBNext Ultra II Ligation Module - 96 rxnsNew England BiolabsCatalog #E7595L
80% Ethanol
TruSeq i7/i5 Indexing Primers - Custom (or NEBNext® Multiplex Oligos for Illumina)New England BiolabsCatalog #E7500L
NEBNext Ultra II Q5 Master Mix - 250 rxnsNew England BiolabsCatalog #M0544L
USER Enzyme - 250 unitsNew England BiolabsCatalog #M5505L
Kapa HiFi Real-Time Amplification KitKapa BiosystemsCatalog #KK2702
Illumina P5 and P7 primers 5uM combined; P5: 5’ AATGATACGGCGACCACCGAGATCT P7: 5’ CAAGCAGAAGACGGCATACGAGAT
Qubit 1X dsDNA High Sensitivity Assay KitThermo Fisher ScientificCatalog #Q33230
Bioanalyzer chips and reagents (DNA High Sensitivity kit)Agilent Technologies
Sodium Orthovanadate (Vanadate) - 1 mlNew England BiolabsCatalog #P0758S
Troubleshooting
Before start
Ensure that you are working in a PCR hood in a pre-PCR space if you are working with metagenomic samples or in a PCR hood if you are working with isolate samples. FLASH is very sensitive to environmental contamination.
Please refer to the guidelines section in this protocol if you are using Ampure beads or other SPRI beads to ensure you use the correct cut-offs.
For mixing of sample prior to PCR, avoid vortexing to keep DNA intact, and instead mix by pipetting and tapping sides of tube.
Dephosphorylation
Normalize your cDNA or gDNA to anywhere between 10pg-100ng. For most samples, we recommend 5-10ng input.
Prepare a reaction for each cDNA or gDNA sample and mix well. Add the components in the order specified below. You can make a master mix (MM) of the rapid alkaline phosphatase (RAP) buffer and enzyme. If you are using the RAP MM, mix 3 µL of MM with each sample. Mix thoroughly with a pipette or by tapping to avoid shearing.
rAPID alkaline phosphatase enzyme and bufferMerck MilliporeSigma (Sigma-Aldrich)Catalog #4898133001
| Component | 1X | |
| DNA, 10pg - 100ng | x μL | |
| rAPid Alkaline Phosphatase Buffer | 2 μL | |
| rAPid Alkaline Phosphatase | 1 μL | |
| H2O | up to 20 μL |
RAP MM.
Incubate at 37 °C for 00:30:00 with heated lid OFF.
Add 1 µL sodium orthovanadate (competitive inhibitor of phosphatases) to the quench the reaction and mix well with a pipette or by tapping.
Sodium Orthovanadate (Vanadate) - 1 mlNew England BiolabsCatalog #P0758S
Cas9 Treatment
Prepare dual-guide RNAs by annealing crRNA and tracrRNA at an equimolar amount at 95 °C for 00:00:30 then allowing the mixture to cool to room temperature on the bench. (Note. If crRNA and tracrRNA have been previously annealed and stored, it is recommended to re-anneal the dual-guide RNAs under the above conditions)
Dual guide RNAs (4µM - targeted to genes or regions to be depleted - crisprRNA and tracr RNA - quantified by RNA Qubit)
If your starting stock of Cas9 is more than 4μM, you must dilute your stock of Cas9 to 4μM by using 1X Cas9 activity buffer.
cas9 4µM or higher concentration
10X Cas9 Activity Buffer (500mM Tris pH 8.0 - 1M NaCl - 100mM MgCL2 - 10mM TCEP)
Make a Cas9 master mix as described below. Add the components in the order specified to prevent precipitation.
| Component | 1X | ____ X | |
| 10x Cas9 Activity buffer | 3 μL | ____ μL | |
| Cas9 4μM* | 2.5 μL | ____ μL | |
| dgRNAs 4μM** | 3 μL | ____ μL | |
| H2O | 0.5 μL | ____ μL | |
| Total | 9 μL | ____ μL |
Cas9 MM
*Remember to dilute your Cas9 stock to 4uM if not already at 4uM. Use 1x Cas9 activity buffer to dilute your Cas9 enzyme if you do need to dilute your stock.
**Remember to dilute your dgRNAs to 4uM using water if needed.
Note: Most experiments for the 2018 FLASH paper were performed at this Cas9 concentration. However, we have demonstrated that lower concentrations work equally well on bacterial isolate DNA. Consult the manuscript for more details.
Add 9 µL of the master mix to each of your 21 µL blocked DNA samples.
Mix well by pipetting or tapping the PCR tubes and incubate the reaction at 37 °C for 02:00:00 .
Deactivate the Cas9 by adding 1 µL of Proteinase K to each of your sample tubes, mixing again by pipetting or tapping. Incubate at 37 °C for 00:15:00 .
Proteinase KNew England BiolabsCatalog #P8107S
SPRI Clean-up at 1.7X
Equilibrate clean SPRI beads to room temperature and vortex well to mix.
SPRI beads (homemade) or Ampure XP beads
Add beads equivalent to 1.7X the sample volume to each sample tube (for 31 µL of sample, add 53 µL beads).
Mix well by pipetting or tapping the tubes. Pulse-spin in a picofuge for no more than 2 seconds.
Incubate for 00:05:00 at room temperature, then put the tubes on the magnetic
rack. Allow beads to separate on the magnet for 3-5 minutes, or until the supernatant is clear.
Keeping the tubes on the magnet, carefully remove and discard the supernatant.
Add 200 µL 80 % volume ethanol (prepared fresh). Incubate beads for 00:01:00 and then remove the ethanol
80% Ethanol
Repeat the above ethanol wash step.
Allow the beads to air dry for 00:05:00 . Do not overdry. Dry beads should appear matte (rather than glossy), but should not have a cracked appearance. Overdried beads, as indicated by a cracked appearance, may not resuspend or elute well.
Remove tubes from magnet and resuspend in 53 µL nuclease-free H2O by pipetting up and down.
Continue to resuspend by tapping the tubes and then spin down briefly in a picofuge.
Allow 00:02:00 for DNA to elute from beads, then transfer tubes back to magnet
Allow the beads to separate for at least 2 minutes.
Collect 50.4 µL of supernatant to clean PCR tubes.
dA tailing
Prepare the following mixture for each sample. You can prepare the dA-tailing buffer and Klenow fragment ahead of time as a MM. If you prepared dA-tailing MM, add 9.6uL of MM to each sample.
| Component | 1X | |
| FLASHed sample | 50.4 μL | |
| dA-Tailing buffer | 6 μL | |
| Klenow fragment | 3.6 μL | |
| Total | 60 μL |
Klenow Fragment (3'-5' exo-) - 1,000 unitsNew England BiolabsCatalog #M0212L
Mix well by pipetting up and down several times with a P200 set to 40uL.
Incubate all tubes at 37 °C for 00:30:00 with heated lid OFF
Cool all tubes to 4 °C and proceed with the next part as soon as possible.
Adaptor Ligation
Note
! The reagents used in this step are very viscous and must be mixed well before using.
! Do NOT make a master mix for this step, although the Ligation Master Mix and Ligation Enhancer may be mixed up to 4 hours before and kept at 4°C.
Prepare the following mixture for each sample. Alternatively, if you prepared an adaptor MM with Ligation MM and Ligation Enhancer, add 31uL of MM to each sample and then add 2.5uL of adaptor to each sample.
| Component | 1X | |
| dA-tailed sample from part IV | 60 μL | |
| NEB Ultra II Ligation Master Mix | 30 μL | |
| NEBNext Ligation Enhancer | 1 μL | |
| NEBNext Adaptor 1:100 or 1:300 dilution* | 2.5 μL | |
| Total | 93.5 μL |
Note: Adaptor dilution of 1:100 works for an initial DNA input up to 100 ng. A 1:300 dilution is recommended for an initial DNA input of under 10 ng. The subsequent cleanup step should remove all extra adaptor and adaptor dimers.
NEBNext Adaptor for IlluminaNew England Biolabs
NEBNext Ultra II Ligation Module - 96 rxnsNew England BiolabsCatalog #E7595L
Prepare the above mixture and mix well by pipetting up and down several times with a P200 set to 50 µL .
Incubate at 20 °C for 00:15:00 in a thermocycler with the heated lid OFF.
SPRI Clean Up 1X + Addition of TruSeq Indexing Primers
Thaw a TruSeq i5/i7 barcode plate or other TruSeq primers, and choose barcodes for each sample. Take note of plate color/barcodes to be used. DO NOT use the same barcode for more than one sample in a sequencing run. Dual unique TruSeq barcodes are preferable.
TruSeq i7/i5 Indexing Primers - Custom (or NEBNext® Multiplex Oligos for Illumina)New England BiolabsCatalog #E7500L
Equilibrate clean SPRI beads to room temperature and vortex well to mix.
SPRI beads (homemade) or Ampure XP beads
Add beads equivalent to 1X the sample volume to each sample tube (for 93.5 µL of sample, add 93.5 µL beads).
Mix well by pipetting or tapping the tubes. Pulse-spin in a picofuge for no more than 2 seconds.
Incubate for 00:15:00 at room temperature, then put the tubes on the magnetic rack. Allow beads to separate on the magnet for 3-5 minutes, or until the supernatant is clear.
Note
Longer incubation time during SPRI addtion is because it may be helpful in removing unwanted fragments/adapter dimers
Keeping the tubes on the magnet, carefully remove and discard the supernatant.
Add 200 µL 80 % volume ethanol (prepared fresh). Incubate beads for 00:01:00 and then remove the ethanol
80% Ethanol
Repeat the above ethanol wash step.
Allow the beads to air dry for 00:05:00 . Do not overdry. Dry beads should appear matte (rather than glossy), but should not have a cracked appearance. Overdried beads may not resuspend or elute well. Ensure the beads are fully dry before eluting.
Remove tubes from magnet and resuspend in 17 µL nuclease-free H2O.
Resuspend well by tapping the tubes and spin down briefly in a picofuge.
Allow 00:02:00 for DNA to elute from beads, then transfer tubes back to magnet
Allow the beads to separate for at least 2 minutes.
Collect 15 µL of elution.
Mix 10 µL of the elution with the appropriate TruSeq indexing primer barcodes as planned.
Note
STOPPING POINT: If necessary, samples may be stored at -20C.
USER Enzyme + Indexing PCR
Spin down plate or tube strips briefly to collect liquid to bottom of the well.
Prepare the following mixture for each sample. You can make a master mix of USER enzyme and Q5. If doing this, add 28µL of MM to your 25µL sample.
NEBNext Ultra II Q5 Master Mix - 250 rxnsNew England BiolabsCatalog #M0544L
USER Enzyme - 250 unitsNew England BiolabsCatalog #M5505L
| Component | 1X | |
| Ligated sample + TruSeq indexing primers from part VI | 25 μL | |
| NEBNext Ultra II 2X Q5 PCR Master Mix | 25 μL | |
| NEB USER Enzyme | 3 uL | |
| Total | 53 μL |
User/Q5
Set up the following cycling conditions for USER enzyme cutting and indexing PCR in a post-PCR room:
| Temperature | Time | Cycles | |
| 37°C | 15mins | 1 | |
| 98°C | 30 sec | 1 | |
| 98°C | 10 sec | 12 cycles | |
| 65°C | 75 sec | ||
| 65°C | 5 min | 1 | |
| 4°C | - | - |
Set the lid heat to ON at 105°C.
SPRI clean-up 0.9X
Equilibrate clean SPRI beads to room temperature and vortex well to mix.
SPRI beads (homemade) or Ampure XP beads
Add beads equivalent to 1:0.9x the sample volume to each sample tube (for 53 µL of sample, add 47.7 µL beads).
Mix well by pipetting or tapping the tubes. Pulse-spin in a picofuge for no more than 2 seconds.
Incubate for 00:05:00 at room temperature, then put the tubes on the magnetic rack. Allow beads to separate on the magnet for 3-5 minutes, or until the supernatant is clear.
Keeping the tubes on the magnet, carefully remove and discard the supernatant.
Add 200 µL 80 % volume ethanol (prepared fresh). Incubate beads for 00:01:00 and then remove the ethanol
80% Ethanol
Repeat the above ethanol wash step.
Allow the beads to air dry for 00:05:00 . Do not overdry. Dry beads should appear matte (rather than glossy), but should not have a cracked appearance. Overdried beads may not resuspend or elute well.
Remove tubes from magnet and resuspend in 53 µL nuclease-free H2O.
Resuspend well by tapping the tubes and spin down briefly in a picofuge.
Allow 00:02:00 for DNA to elute from beads, then transfer tubes back to magnet
Allow the beads to separate for at least 2 minutes.
Collect 23 µL of supernatant to clean PCR tubes.
KAPA Fluorescence-guided Amplification
Using optical PCR strip tubes which are separated from each other so that they can be removed from the thermocycler one at a time, add KAPA amplification MM and Illumina P7 and P5 (5sol-20 and 5sol-21) primers at 5µM to your samples as below. You can make a master mix of KAPA master mix and primers.
Kapa HiFi Real-Time Amplification KitKapa BiosystemsCatalog #KK2702
Illumina P5 and P7 primers 5uM combined; P5: 5’ AATGATACGGCGACCACCGAGATCT P7: 5’ CAAGCAGAAGACGGCATACGAGAT
| Component | 1X | |
| Amplified and indexed DNA | 23 μL | |
| Kapa amplification master mix | 25 μL | |
| Illumina P5 and P7 5uM primers | 2 μL | |
| Total | 50 μL |
KAPA amplification
Add 50 µL of STD 2 to a clean optical PCR tube.
Cap all tubes with optical caps. Do not write on the caps.
Place your labeled samples in the RT-PCR thermocycler.
Set up the following PCR conditions:
| Temperature | Time | Cycles | |
| 98°C | 45 sec | 1 | |
| 98°C | 15 sec | 20 | |
| 60°C | 30 sec | ||
| 72°C | 1 min 30 sec | ||
| Plate read | |||
| 72°C | 30 sec |
Thermocycling Conditions for KAPA Amplification
Run the program, and watch until your sample either:
1. Crosses the standard (STD) 2 threshold
2. Starts to plataeu
Then pull your sample out DURING THE 72C 30sec INCUBATION after the plate read. It is critical to pull it out during this step, and not when it is denaturing or annealing.
If your sample still has not reached the STD 2 by 20 cycles, let the program finish.
Alternatively, if you have many samples, you can simply apply 10 cycles to all, and evaluate if you need further amplification later.
You may want to take note of the unique number of cycles each sample needed to reach the STD 2 threshold.
SPRI Clean-up 1X
Equilibrate clean SPRI beads to room temperature and vortex well to mix.
SPRI beads (homemade) or Ampure XP beads
Add beads equivalent to 1X the sample volume to each sample tube (for 50 µL of sample, add 50 µL beads).
Mix well by pipetting or tapping the tubes. Pulse-spin in a picofuge for no more than 2 seconds.
Incubate for 00:05:00 at room temperature, then put the tubes on the magnetic rack. Allow beads to separate on the magnet for 3-5 minutes, or until the supernatant is clear.
Keeping the tubes on the magnet, carefully remove and discard the supernatant.
Add 200 µL 80 % volume ethanol (prepared fresh). Incubate beads for 00:01:00 and then remove the ethanol
80% Ethanol
Repeat the above ethanol wash step.
Allow the beads to air dry for 00:05:00 . Do not overdry. Dry beads should appear matte (rather than glossy) but should not have a cracked appearance. Overdried beads may not resuspend or elute well.
Remove tubes from magnet and resuspend in 27 µL nuclease-free H2O.
Resuspend well by tapping the tubes and spin down briefly in a picofuge.
Allow00:02:00 for DNA to elute from beads, then transfer tubes back to magnet
Allow the beads to separate for at least 2 minutes.
Collect 25 µL of supernatant to clean PCR tubes.
Library Analysis
Quantify by HS DNA Qubit
Qubit 1X dsDNA High Sensitivity Assay KitThermo Fisher ScientificCatalog #Q33230
Run a fragment analysis, such as with the HS DNA Bioanalyzer. You are expecting a fragment trace that is characterized by sharp peaks in the 250-650 range. A successful FLASHed sample trace looks approximately like this: (lower marker in green, upper marker in purple)
Bioanalyzer chips and reagents (DNA High Sensitivity kit)Agilent Technologies
Expected result
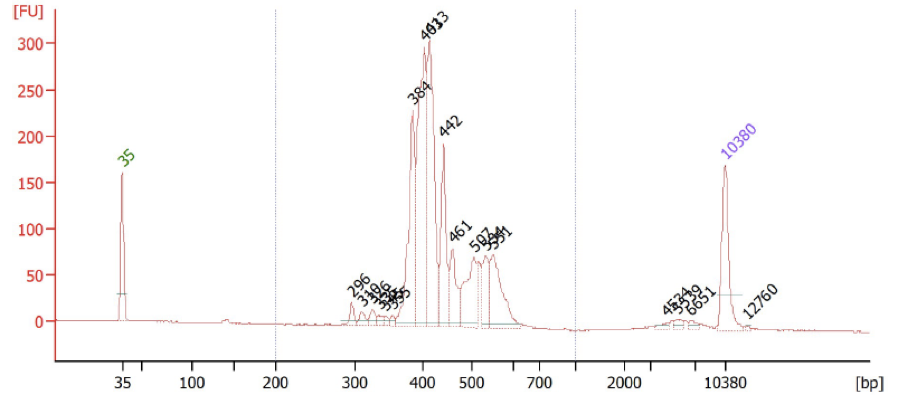
FLASH Bioanalyzer Trace
If there is a large spike at ~138 bp, this is indicative of adaptor dimer in the sample. We recommend: 1) Additional SPRI clean-ups with a sample:bead volume ratio of 1X (as many as necessary for removal of dimers), or 2) size selection to 250-650bp with the BluePippin on a 2% gel.
If pooling multiple samples, use the concentration of DNA between 250-650bp for normalization.
When satisfied with the quality of your pooled/individual library, proceed with quantitative PCR and sequencing.
