Sep 04, 2018
Extracting plasmid DNA from Agrobacterium with the Bioline ISOLATE II Plasmid Mini Kit
- James Lloyd1,
- Ryan Lister1
- 1University of Western Australia

Protocol Citation: James Lloyd, Ryan Lister 2018. Extracting plasmid DNA from Agrobacterium with the Bioline ISOLATE II Plasmid Mini Kit. protocols.io https://dx.doi.org/10.17504/protocols.io.s8fehtn
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: September 03, 2018
Last Modified: September 04, 2018
Protocol Integer ID: 15335
Keywords: Agrobacterium, plasmid, mini-prep, plasmid dna from agrobacterium, bioline isolate ii plasmid mini kit, extracting plasmid dna, plasmid dna, dna from agrobacterium, correct plasmid, transformed agrobacterium, agrobacterium, dna
Abstract
To extract plasmid DNA from Agrobacterium, to confirm the correct plasmid has been taken-up, this protocol allows you to directly perform a mini-prep on transformed Agrobacterium, digest this DNA and visualize it on a gel. This is without the need of rescuing the plasmid in E. coli or genotyping with PCR. This protocol using the commercially available Bioline ISOLATE II Plasmid Mini Kit, designed for use with E. coli. The standard protocol this is modified from is available here: https://www.bioline.com/au/downloads/dl/file/id/1219/isolate_ii_plasmid_mini_kit_protocol.pdf
Materials
MATERIALS
ISOLATE II Plasmid Mini KitBiolineCatalog #BIO-52057
Troubleshooting
Safety warnings
Please check the MSDS information for the ISOLATE II Plasmid Mini Kit: https://www.bioline.com/au/downloads/dl/file/id/2807/isolate_ii_plasmid_mini_kit_msds.pdf
Before start
Ensure RNase A has been added to Resuspension Buffer P1. Pre-warm the required amount of PW1 to 50-55 ºC before starting the extration. Ensure Ethanol has been added to PW2.
Harvest bacterial cells
Pellet 3 ml of a saturated Agrobacterium LB culture with appropriate antibiotics for 30s at 11,000 x g.
Expected result
Typical pellet after spinning down 3 ml of 24 hour grown Agrobacterium.
Note
I grew Agrobacterium in 5 ml of LB with appropriate antibiotics for 24 hours at 29 ºC.
Spinning the 3 ml down can be done in two steps of 1.5 ml.
Discard supernatant and remove as much liquid as possible.
Lyse cells
Add 250μl Resuspension Buffer P1 and resuspend cell pellet by pipetting up and down.
Add 250µl Lysis Buffer P2. Mix gently by inverting tube 6-8 times. Incubate at room temperature for up to 5 min or until lysate appears clear.
Note: Do not vortex to avoid shearing genomic DNA
00:05:00 Lysis
Add 300µl Neutralization Buffer P3. Mix thoroughly by inverting tube 6-8 times.
Note: Do not vortex to avoid shearing genomic DNA
Clarification of lysate
Centrifuge 10 min at max speed at room temperature.
Note
The max speed on the centrifuged I used was 20817 x g but faster is probably not a bad thing.
The pellet is very glopy (snot-like) so a long, fast spin at this stage is vital to being able to get the supernatant.
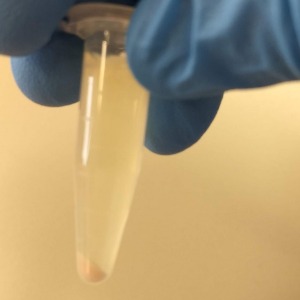
Expected result
This is the very snot-like pellet. Ensure you do not disturb it when you take the supernatant.
Bind DNA
Bind DNA Place ISOLATE II Plasmid Mini Spin Column in a 2ml Collection Tube. Pipette (do not decant) a maximum of 750µl (usually less) of clarified sample supernatant onto column.
Centrifuge 1 min at 11,000 x g and discard flow-through.
Wash silica membrane
Add 500µl Wash Buffer PW1 preheated to 50°C. Centrifuge 1 min at 11,000 x g. Discard flow-through and reuse Collection Tube.
Note
The use of PW1 is considered optional for some E. coli strains but is essential for extracting plasmid from Agrobacterium as it contains many nucleases, which PW1 is used to remove.
I actually warmed it up to 55 ºC, but I am sure the manual's recommended 50 ºC will work just the same.
Add 600μl Wash Buffer PW2 (supplemented with ethanol). Centrifuge 1 min at 11,000 x g. Discard flow-through and reuse Collection Tube.
Dry silica membrane
Centrifuge 2 min at 11,000 x g, to remove residual ethanol. Place ISOLATE II Plasmid Mini Spin Column in a 1.5ml microcentrifuge tube.
Elute DNA
Add 30-50μl Elution Buffer P, nuclease-free water or TE buffer directly onto center of silica membrane. Incubate at room temperature for 1 min.
00:01:00 Elution incubation
Centrifuge 1 min at 11,000 x g.
Restriction enzyme digestion
The DNA is now very dilute so it would be a good idea to use 20 µl in a typical restriction enzyme digest. This is usually much less than you would use from a prep from E. coli.
