Aug 01, 2025
ESPEC-SUIT - A versatile and robust platform to identify and track antigen-specific T-cell receptors in cancer patients
- Isabel Poschke1,2,
- Katharina Lindner1,2,
- Saskia Stange1,2,
- julian.hlawatsch hlawatsch1,2
- 1German Cancer Research Center (DKFZ) Heidelberg;
- 2National Center for Tumor Diseases (NCT) Heidelberg

Protocol Citation: Isabel Poschke, Katharina Lindner, Saskia Stange, julian.hlawatsch hlawatsch 2025. ESPEC-SUIT - A versatile and robust platform to identify and track antigen-specific T-cell receptors in cancer patients. protocols.io https://dx.doi.org/10.17504/protocols.io.rm7vzkq42vx1/v1
Manuscript citation:
ESPEC-SUIT - A versatile
and robust platform to identify and track antigen-specific T-cell receptors in
cancer patients; Journal for Immunotherapy of Cancer - in press
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: February 07, 2025
Last Modified: August 01, 2025
Protocol Integer ID: 119774
Keywords: T cell expansion, TCR sequencing, immunogenicity, TCR discovery, Cytokines, cell receptors in cancer patients espec suit, tcr repertoire analysis, out by tcr repertoire analysis, cancer patients espec suit, tcr candidate, candidate antigen, tcr cloning, epitope specific expansion cultures with subsequent identification, antigen, primed immune response, cell sequencing, immune response, espec, reactive tcr, cell receptor, espec suit, stream tcr, vaccine
Funders Acknowledgements:
German Cancer Research Center; Heidelberg, Germany
Grant ID: _
National Center For Tumor Disesases (NCT); Heidelberg, Germany
Grant ID: NCT IMPlatform
German Cancer Consortium (DKTK)
Grant ID: TCR Discovery Platform
Abstract
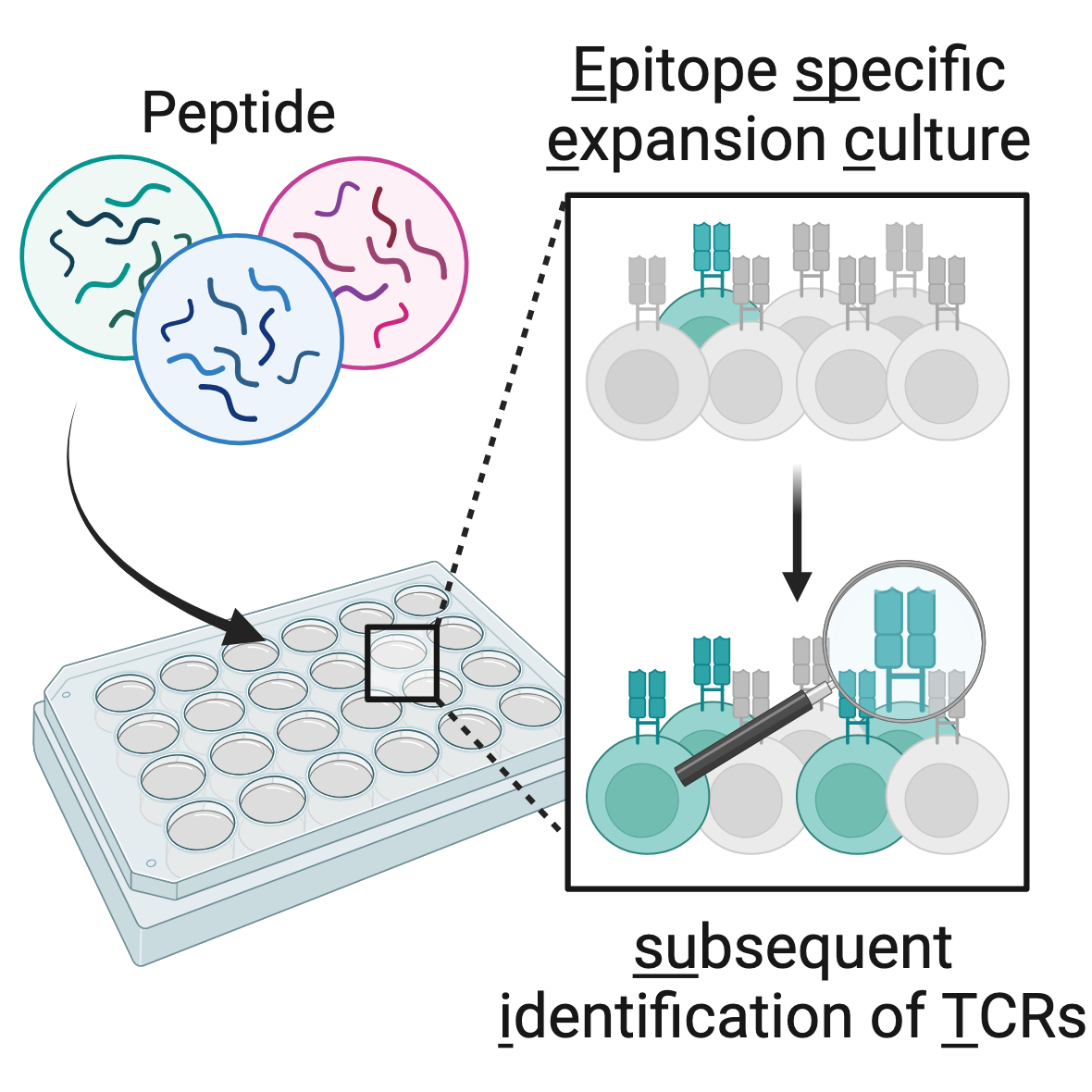
ESPEC SUIT - epitope specific expansion cultures with subsequent identification of (tumor-)specific TCRs:
ESPEC SUIT is a modular platform for identification of antigen-specific T cell clonotypes from peripheral blood. It makes use of in vitro expansion to amplify in vivo primed immune responses, which are then read-out by TCR repertoire analysis, paired with optional functional assays.
Candidate antigen-reactive TCRs can be traced in blood and tissue samples based on their sequence to gain deep insight into e.g. tumor-specific or vaccine-induced T cell responses. We found that >75% of TCR candidates identified in ESPEC SUIT can be functionally validated (if optional single-cell sequencing and TCR cloning, not described herein, are performed).
The supplied protocol describes the in vitro expansion, to be followed up by down-stream TCR repertoire analysis and discovery.
Image Attribution
image copyright: Saskia Stange, image generated with Biorender (www.BioRender.com)
Materials
GENERAL
- pipettes + tips
- serological pipettes + pipetting aid
- centrifuge tubes (conical, sterile) (15 mL, 50 mL)
- 1.5 mL tubes (sterile, safe-lock)
- cell counting equipment
- 24-well plates, cell culture grade
- cryotubes
- controlled rate freezing device
- sterile filters (0.22 µM, protein low-bind)
- DMSO
- EDTA
- Benzonase
- PBS (Ca2+, Mg2+)
OPTIONAL
reagents for magentic-bead based cell enrichment, e.g. CD4+ T-cell isolation kit, human (Miltenyi, 130-096-533)/anti-human CD4 microbeads (Miltenyi, 130-045-101) or CD8+ T-cell isolation kit, human (Miltenyi, 130-096-495)/ anti-human CD8 microbeads (Miltenyi, 130-045-201)
CYTOKINES
| A | B | C | D | E | |
| Working conc (2x) | final conc per well | manufacturer | catalog no. | ||
| IL-7 | 50 ng/mL | 25 ng/mL | Miltenyi | 130-095-362 | |
| IL-15 | 50 ng/mL | 25 ng/mL | Miltenyi | 130-095-764 | |
| IL-2 | 100 IU/mL | 50 IU/mL | Novartis | Proleukin, Ch-B. 502519J |
Cytokine concentrations for ESPEC culture media
CELL CULTURE MEDIUM
- X-Vivo20 (04-448Q, Lonza) with 2% human AB-Serum (e.g. H4522-100ML, Sigma-Aldrich; serum sterile filtered with 0.22 µm protein-low bind filter)
Troubleshooting
Before start
Starting material for this protocol are human peripheral blood mononuclear cells (PBMC), isolated from blood by densitiy gradient centrifugation and cryopreserved. Fresh PBMC may also be used, omitting thawing and resting steps.
THAWING OF CELLS (day 0)
50m
thaw PBMC according to internal protocol. We recommend thawing in pre-warmed lymphocyte medium, such as X-Vivo20 (Lonza), and in presence of human serum
Note
PBMC with known reactivity against the antigen-of-interest, e.g. previously determined by ex-vivo ELISpot, will usually result in the strongest expansion
30m
determine cell count and viability
Expected result
a viability >80% is expected
10m
remove cells for baseline pellet
10m
transfer 1-2 x106 PBMCs to a 1.5 mL tube (DNA-LoBind)
centrifugation:1000 x g, 20°C, 00:05:00
completely remove supernatant, freeze/store pellet at -80°C for subsequent DNA or RNA extraction
RESTING
12h
depending on the cell number, place 2-10 x 106 cells/mL in culture media in a 15 or 50 mL conical tube with a slightly unscrewed lid
Note
to avoid cells setteling too densely at the bottom of the tube, place tabe in a tilted position
incubate PBMC overnight at 37°C, 5% CO2
CULTURE SETUP (day 1)
4h 55m
centrifugation: 430 x g, 20°C, 00:10:00
10m
remove supernatant, resuspend cell pellet in 2 mL culture media without cytokines
5m
determine cell count and viability
5m
optional: if an ex vivo antigen reactivity assay is planned, remove cell number needed for this assay
5m
adjust concentration to 2x106 PBMC/mL
5m
prepare two 24-well plates: one for 'antigen presenting cells' (APC), one for 'effectors', both will have the same layout
5m
add 250 µL culture medium per used well on both plates
required: For each assay, there has to be one culture expanded without antigen ('no peptide').
For each antigen and the 'no peptide' control, at least one well is set up on each plate.
Note
for optimal results, set up 2-4 wells/antigen or condition
Note
if a post-ESPEC readout is planned to verify expansion of T cells with the relevant specificity (QC assay), the number of 'no peptide' stimulated wells increases with the number of antigens used, as it is used to determine the background (unspecific expansion) for all antigens after expansion.
5m
add 250 µL cell suspension/used well on both plates
5m
APC Plate: add 4 µg/mL peptide of interest to each well (total volume: 500 µL --> 2 µg/well)
exception: 'no peptide' wells
5m
incubate both plates at 37°C and 5% CO2 for 4h
4h
after 4h of incubation, thoroughly resuspend wells of 'effectors' plate and transfer to corresponding well of 'APC' plate (--> total volume 1 mL, peptide conc. 2 µg/mL)
Discard effectors plate.
5m
incubate at 37°C and 5% CO2
FEEDING AND SPLITTING OF CULTURES (day 4-11)
10m
feeding of cells is performed on day 4, 7, 9 and 11 by replacing half the medium volume with fresh, cytokine containing medium. No peptide is added!
10m
Carefully remove half the medium volume (500 µL) of each well without disturbing the cells
Replace with 500 µL of fresh cytokine-containing culture medium
Note
see materials section for final cytokine concentrations. Prepare a working solution 2x the cytokine concentration needed/well, to achieve the correct final concentration after exchanging 50% of the medium
Note
Alternatively, splitting of cells is performed if wells have turned orange/yellow, as this indicates strong T cell proliferation. No peptide is added!
--> Carefully resuspend the well in question
--> transfer half the medium volume (500 µL) to a new well
--> fill up both wells (old/new) with 500 µL of fresh medium containing 2x the cytokine concentration needed/well
END OF ESPEC (day 14)
2h 30m
harvest cells by carefully resuspending all wells stimulated with the same peptide and pooling them in one 15 mL tube. Rinse well(s) with medium and add to tube.
10m
centrifugation: 430 x g, 20°C, 00:10:00
10m
remove supernatant and resuspend cells for counting
5m
determine cell count and viability
5m
plan use of cells (see note) for TCR repertoire analysis (required), QC-assay (recommended) and SUIT (if desired)
| A | B | C | |
| culture | viability [%] | total cell no. | |
| 'no peptide' | |||
| 'antigen a' | |||
| 'antigen b' | |||
| ... |
Cells available for down-stream analysis
Note
| A | B | C | |
| Assay | cells needed [x10e6] | comment | |
| TCR repertoire analysis by deep sequencing--> step 22-23 | 0.3-1 | consider pre-enriching CD4/CD8 T cell subsets depending on antigen | |
| QC readout by ELISpot (performed after over night resting in cytokine-free media) --> step 24 | 0.15-1.2 | 25,000 - 125,000 cells/well depending on culture expansion and freq of ex vivo response; plan with duplicates or triplicates, include positive/negative controls; 'no peptide' culture should be tested against all antigens | |
| QC readout by intracellular cytokine stain (performed after over night resting in cytokine-free media) --> step 24 | ~0.9 | 300,000 cells/condition, 1 well/condition, include positive/negative controls, 'no peptide' culture should be tested against all antigens | |
| single cell sequencing for TCR retrieval --> step 23 | 0.3-0.5 | to be frozen, scSeq after evaluation of QC readouts |
Cell amounts needed (values represent general recommendations and will depend on your assay of choice and internal protocols used)
collect cells for TCR repertoire sequencing of antigen-expanded and control stimulated cultures.
in parallel: prepare cells for single cell sequencing (if planned, see step 23) and QC assay (if planned, see step 24)
optional step: Isolate CD4+ oder CD8+ T cell subsets using a method of your choice
Note
We recommend magnetic bead-based cell isolation procedures, which usually give significant enrichment of the cell type of interest, are easy to perform and less stressfull on the cells than flow cytometric cell sorting. The advantage of T cell subset enrichment is improved signal-to-noise in repertoire analysis and more sequence information on the relevant subset, increasing sensitivity.
If antigen restriction is not known, e.g. after stimulating with long peptides with unknown binding to MHC I/II molecules, one can either analyze all T cells, or separate CD4 and CD8 T cells to gain insight into the main responding subset.
1h 30m
Transfer 0.3-1x106 (better 2 x 106) cells to a 1.5 mL tube (DNA-LoBind)
5m
centrifugation: 1000 x g, 20°C, 00:05:00
5m
completely remove the supernatant. Freeze the pellet at -80°C for subsequent DNA/RNA extraction and TCR repertoire sequencing by a method of your choice.
Note
Various methods for TCR repertoire sequencing are available and will require different amounts of input material. We have performed post-ESPEC repertoire analysis using ~0.5 x 106 expanded cells using DNA-based (Immunoseq, Adaptive Biotechnologies) as well as RNA-based (DriverMap AIR, Cellecta) methods. TCRβ sequencing is sufficient for selection of expanded, candidate antigen-reactive clonotypes.
We provide TCRSelect (https://tcrselect.dkfz.de), a tool to explore post-ESPEC repertoire data and select candidates for further analysis.
5m
freeze viable cells for single cell TCR sequencing in DMSO-containing freezing medium.
Note
Single cell TCR sequencing is required to retrieve paired TCRαβ information, which is needed for TCR cloning/recombinant expression. Most platforms will accept <20,000 cells as input. However, since freezing very low cell numbers can lead to loss of viability upon thawing, we recommend to freeze >300,000 cells (optimally 1 x 106) per culture to be analyzed. In most cases, only post-ESPEC samples with clear antigen-specific clonal expansion will undergo single cell TCR sequencing to enable functional validation of candidate antigen-reactive TCRs.
10m
rest cells needed for QC readout at 2 x106/mL in culture media without cytokine or peptide. For resting, cells can be returned to their original well on the 24-well plate or for volumes >500 µL placed in a 15 mL tube (with partically unscrewed lid). Cells are incubated over night at 37°C and 5% CO2. On the next day, perform a relevant QC assay according to your internal protocols.
Note
Consider cell loss during resting when planning the cell number needed for the QC assay
Note
For each culture, T cell reactivity against the antigen-of-interest is assessed.
Relevant controls include: no stimulation; strong, unspecific activation (induced e.g. by PMA/ionomycine, SEB, anti-CD3 beads); optimally: control antigens such as irrelevant peptides or wildtype-variants of neoepitopes)
Note
Recommended cell numbers:
ELISpot as QC readout:
~50,000 cells/well for strongly expanding cultures and/or antigens to which a clearly measurable ex vivo T cell response was detected
~125,000 cells/well for weakly expanding cultures
--> if you are unsure about the strength of the response and require a quantitative readout, consider titrating the cell number or using flow cytometry as readout
Intracellular Cytokine Staining (Flow Cytometry, e.g. IFNγ or TNFα) as QC readout:
- 300,000 - 400,000 cells/condition
5m
