Apr 23, 2025
Enterovirus A71 3C protease small scale expression and purification protocol
- Korvus Wang1,2,3,
- Michael Fairhead1,2,3,
- Eleanor Williams1,2,3
- 1Centre for Medicines Discover;
- 2University of Oxford;
- 3ASAP Discovery Consortium
- ASAP Discovery

Protocol Citation: Korvus Wang, Michael Fairhead, Eleanor Williams 2025. Enterovirus A71 3C protease small scale expression and purification protocol. protocols.io https://dx.doi.org/10.17504/protocols.io.36wgqndzogk5/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: June 14, 2024
Last Modified: April 23, 2025
Protocol Integer ID: 101842
Keywords: 3C proteases, ASAP, CMD, protein purification, Enterovirus A71, Enterovirus, AViDD, enterovirus a71 3c protease small scale expression, enterovirus a71 3c protease, purification of enterovirus a71, enterovirus a71, 3c protease, purification protocol, purification protocol this protocol, purification, protocol details the expression
Funders Acknowledgements:
National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID)
Grant ID: U19AI171399
Disclaimer
Research was supported in part by NIAID of the U.S National Institutes of Health under award number U19AI171399. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Abstract
This protocol details the expression and purification of enterovirus A71 3C protease construct bearing a C-terminal His-tag at small scale (<6L).
Attachments
Guidelines
- Construct / plasmid resource-name: Enterovirus A71 3C protease construct bearing a C-terminal His-tag.
Materials
Plasmid details:
Addgene plasmid #204816
- Vector: pNIC
- Cell line: E. coli Rosetta strain BL21(DE3)-RR
- Tags and additions: C-terminal, non-cleavable hexahistidine tag
- Construct protein sequence: ` MGPSLDFALSLLRRNIRQVQTDQGHFTMLGVRDRLAVLPRHSQPGKTIWVEHKLINILDAVELVDEQGVNLELTLVTLDTNEKFRDITKFIPENISAASDATLVINTEHMPSMFVPVGDVVQYGFLNLSGKPTHRTMMYNFPTKAGQCGGVVTSVGKVIGIHIGGNGRQGFCAGLKRSYFASEQLEHHHHHH
Expression
AIM-TB: TB autoinduction media (Formedium AIMTB0210)
Purification
Chicken hen egg white lysozyme (Merck, 62971)
Benzonase (Merck, 1.01654)
Imidazole (Merck, RDD044)
Ni Sepharose 6 FF resin (Cytiva, 17531801)
Gravity flow column, 2.5cm diameter (Bio Rad, 7372532)
Centrifugal concentrators, 10kDa MWCO (Merck, UFC901008)
On an FPLC system:
SEPAX SEC SRT-100 (Sepax Tech, 215100-21230)
or
HiLoad 16/600 Superdex 75 pg (Cytiva, 28989333)
5mL sample loop
SDS-PAGE sample buffer, gel, and gel tank
Lysis buffer:
| A | B | |
| Hepes (pH 7.5) | 50 mM | |
| NaCl | 500 mM | |
| Glycerol | 5% | |
| Imidazole | 20 mM | |
| TCEP | 0.5 mM | |
| Lysozyme | 0.5 mg/mL | |
| Benzonase | 0.05 mg/mL | |
| MgSO4 | 1mM |
Prepare 100 mL per 1 L E.coli expression
Base buffer:
| A | B | |
| BIcine (pH 8.5) | 10 mM | |
| NaCl | 500 mM | |
| Glycerol | 5% | |
| TCEP | 0.5 mM |
Prepare 2 L per 6 L E.coli expression. Used to prepare the following buffers
Note
Use of bicine buffer at pH8.5 is essential to sample concentration steps. Previous purifications using HEPES buffer at pH 7.4 encountered significant difficulty when concentrating sample for SEC.
Binding buffer: base buffer
Wash buffer: base buffer, add 30mM imidazole
Elution buffer: base buffer, add 300mM imidazole
Gel filtration buffer: same as base buffer
SDS-PAGE: NuPage 4-12%, Bis-Tris protein gel, 26 well (Thermo-Fisher, WG1403BOX)
Run in MES buffer, 200V 35mins.
Protocol materials
Enterovirus A71 3C proteaseaddgeneCatalog #204816
Troubleshooting
Safety warnings
Always wear appropriate PPE for this protocol
Refer to Material Safety Data Sheets for additional safety and handling information.
Abbreviations
CV - column volume, total volume of resin in a column
IMAC - immobilised metal affinity chromatography
A71EV3C - Enterovirus A71 3C protease
Plasmid Transformation
1d
Transform Enterovirus A71 3C proteaseaddgeneCatalog #204816 into BL21(DE3) and store a and store a glycerol stock of this at -80 °C
Note
The EV-A71 3C construct encodes the 3C protease with a non-cleavable C-terminal His tag on a kanamycin resistant plasmid backbone with a T7 promoter.
Protein expression
2d 10h
Scrape off some of the glycerol stock with a sterile loop and use this to inoculate a 50 mL falcon tube containing 10 mL of LB supplemented with 50 ug/mL kanamycin. Grow the starter culture at 37 °C Overnight with 200 rpm shaking.
1d
Use the 10 mL starter culture to inoculate 1 L auto-induction media (see Materials) supplemented with 50 ug/mL kanamycin in a baffled flask. 250 rpm, 37°C
Note
For this protocol typically 2 L of culture is grown for each purification
6h
When the OD600 reaches approximately 4.0, lower the temperature and shaker speed to 200 rpm, 18°C and incubate 24:00:00
1d
Harvest the cells by centrifugation at 5000 x g, 4°C, 00:30:00 . Discard supernatant and store pellet at -80 °C .
30m
Protein Purifcation
2d
Lyse cell pellet
2h 30m
Note
See Materials tab for buffer compositions.
Note
A71EV3C construct protein properties
MW = 21.332 kDa
Extinction coefficient (assume all Cys reduced)=9970 mM-1cm-1
pI = 7.22
Values determined using Expasy ProtParam
Thaw and resuspend the pellet in ~7mL of lysis buffer per g of pellet. Stir gently with a magnetic stir bar at Room temperature for 00:30:00 to allow lysozyme and benzonase to start breaking down cell components.
1h
Lyse by sonication 00:00:02 On 00:00:04 Off for a total 'on' time of 00:15:00 at 35% amplitude to fully rupture the cells. Ensure pellet is 0 °C during sonication to prevent overheating.
15m 6s
Centrifuge the lysed cells 38000 x g, 4°C, 01:00:00 to remove insoluble cell debris, and collect the supernatant 4 °C
1h
Perform IMAC to extract target protein from the lysed cell mixture
Dispense 10 mL Nickle affinity resin (Ni Sepharose 6 FF, Cytiva) into a gravity flow column.
Wash the resin first with ~ 20 µL distilled water to remove the storage solution and then ~ 20 µL binding buffer to equilibrate
10m
Pour the clarified supernatant over the equilibrated resin and allow to flow though. This will allow the His-tagged target protein to bind onto the Ni ions in the resin.
Retain the flow through separately for SDS-PAGE analysis.
Wash the column with 10 µL of wash buffer twice. Allow wash buffer to pass through completely between washes. This is to remove non-specific, weak binding of contaminant proteins from the resin for a cleaner elution.
Collect washes separately for SDS-PAGE analysis.
30m
Elute the protein with 1.5 µL of elution buffer.
20m
Repeat step 7.5 a further 2 times, collecting a total of 3 separate elution fractions. This is to ensure maximum retrieval of protein from the resin.
Measured the A280 values of the elution fractions to estimate the protein content
For example:
E1: A280=5.94
E2: A280=14.77
E3: A280=1.21
Note
High A260 value is observed across all elutions, indicating presence of large amounts of nucleotides. This can be mitigated by including an additional wash step, incubating the resin with base buffer + 1:2000 benzonase and 2mM MgCl2. This method was employed in large scale purifications. See relevant protocol.
20m
Run SDS-PAGE of all samples from total lysis supernatant to final elution. Stain gel with Coomasssie Blue and determine which fractions contain the target protein by finding the band corresponding to the target molecular weight, 21.3 kDa.
Note
The target protein is expected to be present mostly in the elution samples, although small amounts may be found in the flow through and washes.
If that is not the case, then further troubleshooting is required.
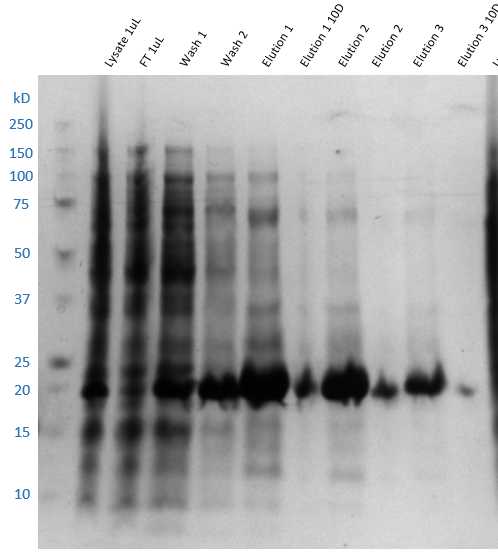
SDS-PAGE analysis of IMAC fractions. The thick protein band observed in all three elutions agree with the calculated molecular weight of A71EV3C protease, 21.3 kDa.
40m
Purify sample further by size exclusion chromatography.
Pool and concentration all elution fractions using a 10 kDa MWCO centrifugal concentrator
as much as possible without protein precipitation. Ideally the final volume should be under 5 mL .
Note
This was not able to be achieved during our purification, possibly because of high DNA concentration as mentioned in section 8.5. The final volume was 21mL, and multiple rounds of SEC was carried out.
Remove any solid aggregates from the sample by centrifugation at 17200 x g, 4°C, 00:10:00 , then immediately draw up the supernatant with a 5mL syringe and a blunt-tip fill needle, taking care not to disturb the pellet.
Note
This is to remove as much solid particles from the injection sample as possible, so as to not clog the in-line filter or frit of the column.
10m
Using an AKTA Pure FPLC system or equivalent:
Inject the sample onto a 5mL sample loop.
Run the sample down Sepax SRT SEC-100 gel filtration column at 7.5mL/min in gel filtration buffer, collecting 1mL fractions in 96 well deep-well blocks.
The column should be pre-equilibrated in SEC buffer.
Note
Here a SEPAX SEC SRT-100 column was used due to availability, however other columns would also be suitable such as a Superdex 75 16/60 (Cytiva)
Run the peak SEC fractions on SDS PAGE to assess purity.
For example:
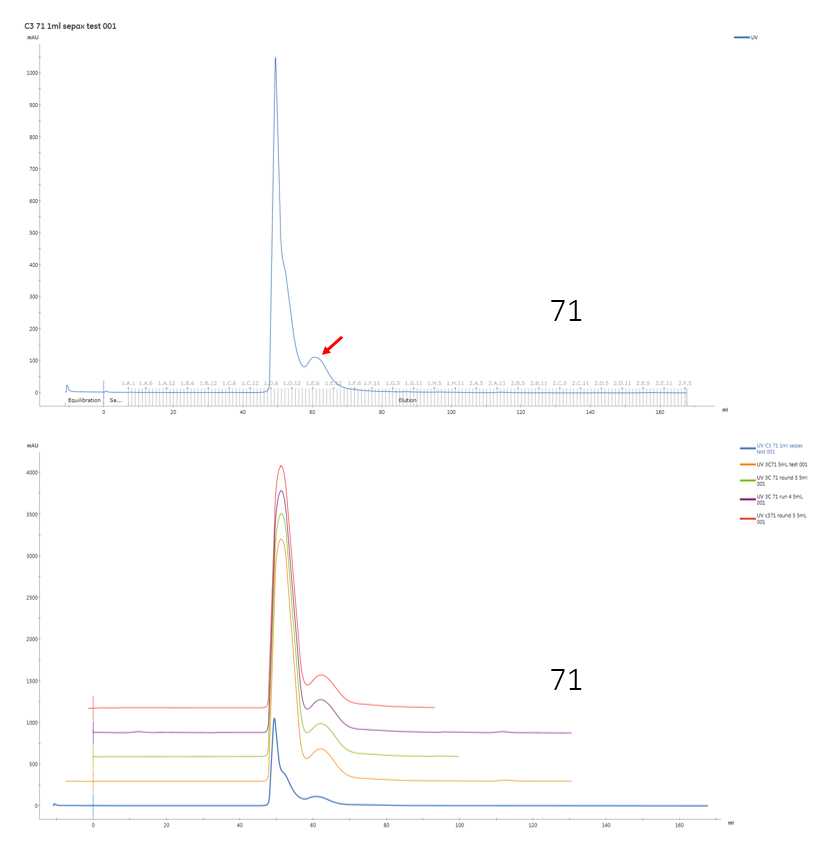
Chromatogram of the SEC run. Top: single chromatogram of A71EV3C SEC. Red arrow indicate the peak containing target protein, as determined by SDS-PAGE. The earlier, higher peak is likely to be from DNA contaminants. Bottom: overlaid chromatograms of repeated SEC runs, illustrating the repeatability of the chromatogram.
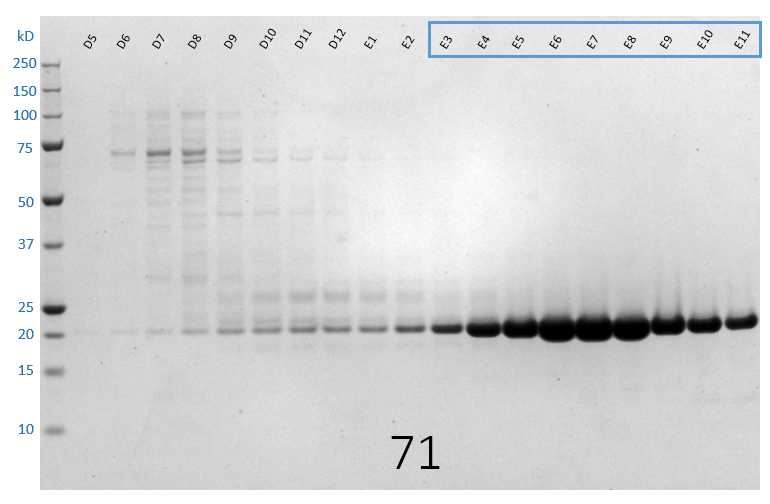
SDS-PAGE analysis of SEC fractions D5-E11 from the first SEC run. Fractions E3-E11 were pooled as they contain majority target protein in comparison to contaminants. Fractions from later runs that contain the same peak were similarly pooled for final sample.
40m
Take the fractions that contain the cleanest target protein and concentrate to around 30 mg/mL using a 10 kDa MWCO centrifugal concentrator
Take 1 µL of the final sample for SDS-PAGE.

SDS-PAGE of the final purified A71EV3CPROB.
2h
Aliquot into appropriate volumes for future usage to minimise freeze/thaw cycles. Flash-freeze in liquid nitrogen, and store at -80 °C until required.
For example:
The final yield from processing 2 L of cells was 128 mg of pure D68 EV 3C protease
NOTE: A71EV3C sample self-cleavage in storage
This A71EV3C construct exhibits self-cleavage of its C-terminal sequence: LEHHHHHH in -80 storage. This is caused by the presence of a 3C cleavage motif near the C-terminal His-tag, and the fact that this is an active construct.

Intact Mass Spectroscopy of sample after around 3 months of -80 storage. Mass shift is observed from the expected 21.331 kDa to 20.267 kDa.

PAWS truncation analysis showing cleavage site

