Jan 26, 2026
Enhanced RNA extraction for diverse tissues
- Damaris Godinez-Vidal1,
- Simon C Groen1
- 1Department of Nematology, University of California Riverside, Riverside, CA 92521.

Protocol Citation: Damaris Godinez-Vidal, Simon C Groen 2026. Enhanced RNA extraction for diverse tissues. protocols.io https://dx.doi.org/10.17504/protocols.io.e6nvwnz5wvmk/v1
Manuscript citation:
Zhou D, Godinez-Vidal D, He J, Teixeira M, Guo J, Wei L, Van Norman JM, and Kaloshian I. 2023. A G-type lectin receptor kinase negatively regulates Arabidopsis immunity against root-knot nematodes. Plant Physiology 193: 721-735. DOI: 10.1093/plphys/kiad253.
Godinez-Vidal D, López-Leal G, Covarrubias AA, and Reyes JL. 2020. Early events leading to water deficit responses in the liverwort Marchantia polymorpha. Environmental and Experimental Botany 178: 104172. DOI: 10.1016/j.envexpbot.2020.104172.
Godinez-Vidal D, Rocha-Sosa M, Sepúlveda-García EB, Lozoya-Gloria E, Rojas-Martínez RI, Guevara L, and Zavaleta-Mejía E. 2013.Transcript accumulation of mevalonic pathway genes and enzymatic activity of HMGCoA-r and EAS in chilli CM-334 infected by Nacobbus aberrans. Plant and Soil 372: 339-348. DOI: 10.1007/s11104-013-1743-8.
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: January 25, 2026
Last Modified: January 26, 2026
Protocol Integer ID: 240862
Keywords: RNA extraction, phenol-chloroform, high-throughput sequencing, arthropod, fungus, insect, nematode, oomycete, plant, rna extraction, allowed rna extraction, standardization of rna extraction, enhanced rna extraction, enhanced rna extraction for diverse tissue, genomic rna free of contamination, total rna recovery, isolating total rna, pure rna, solanum lycopersicum, separate nucleic acid, solanum tuberosum, rna degradation, rna, rna from dna, rna extraction from invertebrate animal, rna from mycelia, obtaining genomic rna, including capsicum annuum, liquid extraction approach, total rna from biological tissue, capsicum annuum, phenolic acid content, cultivated tobacco, nicotiana tabacum, using phenol, various solanaceous species, arabidopsis thaliana, different citrus species, phaseolus vulgari, solubility, plant, mixture of guanidinium thiocyanate
Abstract
RNA extraction is a method mostly employed for isolating total RNA from biological tissues. RNA extraction using phenol-chloroform is a classic pH-dependent liquid-liquid extraction approach that separates RNA from DNA and proteins by creating distinct phases: at acidic pH, RNA remains in the upper aqueous layer, while DNA and proteins migrate to the lower organic layer. This is achieved using a mixture of guanidinium thiocyanate, phenol, and chloroform, followed by isopropanol precipitation to isolate pure RNA. This process relies on differential solubility to effectively denature proteins and separate nucleic acids, though it requires careful handling to prevent RNA degradation. We found that changing the buffer type, in combination with modifications to subsequent washing steps, optimizes and increases total RNA recovery. For plants, our protocol has been used with various solanaceous species, including Capsicum annuum (pepper), Nicotiana benthamiana (Australian wild tobacco), Nicotiana tabacum (cultivated tobacco), Solanum lycopersicum (tomato), and Solanum tuberosum (potato), whose phenolic acid content can be a barrier to obtaining genomic RNA free of contamination. This protocol has also been used with Arabidopsis thaliana (thale cress), Oryza sativa (rice), Phaseolus vulgaris (common bean), Vigna unguiculata (cowpea), Populus spp. (poplars), Marchantia polymorpha (common liverwort), Physcomitrium spp. (earthmosses), and different Citrus species. In addition, the protocol has allowed RNA extraction from invertebrate animals including root-knot nematodes (Meloidogyne spp.), cyst nematodes (Globodera and Heterodera spp.), aphids, mites, and thrips. Lastly, with this protocol we have been able to obtain RNA from mycelia of fungi and oomycetes in different genera. Our protocol allows for standardization of RNA extraction, optimization based on tissue type, and improved repeatability across studies.
Image Attribution
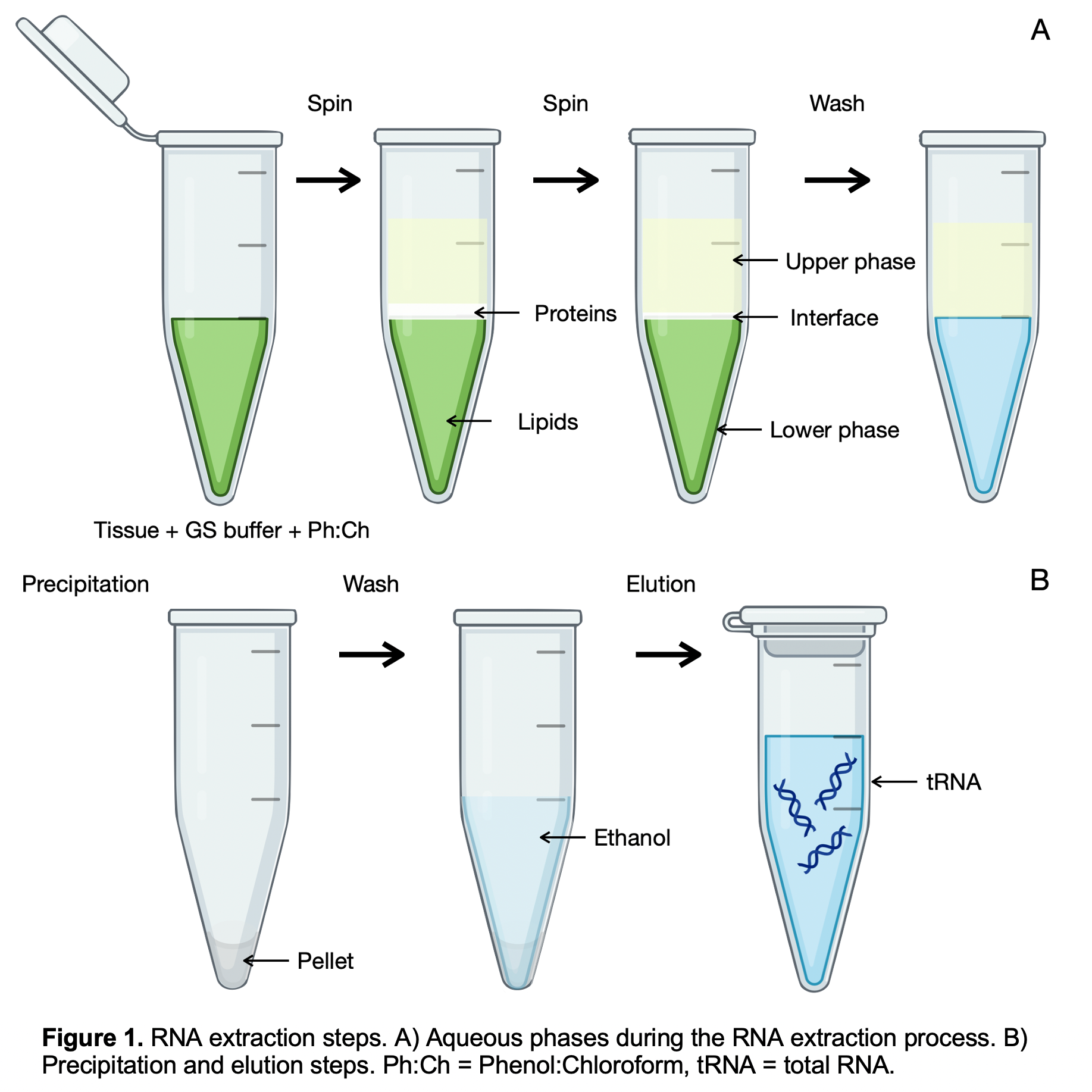
Figure. 1. RNA extraction steps. A) Aqueous phases during the RNA extraction process. B) Precipitation and elution steps. Ph:Ch = Phenol:Chloroform, tRNA = total RNA.
Guidelines
This protocol has been developed from the original method described by Logemann et al. (1987), with modifications for use with different tissues, including plant vegetative and reproductive tissues, plant seeds, fungal mycelia, nematodes, arthropods, and frozen tissue.
Materials
Lab coat (the protocol involves working with toxic solvents)
Goggles (or glasses)
Mortar and pestle
Liquid nitrogen (LN2)
Z6 or GS buffer
Phenol:Chloroform (solution 1:1 vol/vol)
ß-mercaptoethanol
Isopropanol absolute
Ethanol absolute
Sodium acetate at 3 M (pH 5.2)
Ethanol at 70%
DNase- and RNase-free water
Eppendorf Safe-Lock tubes, 1.5 mL
Plastic beaker
Paper towels
Water bath or thermoblock at 65 °C
Refrigerated centrifuge or a standard centrifuge kept in a cold room.
Troubleshooting
Safety warnings
Remember to take precautions before working with detergents and solvents, such as wearing closed-toe shoes, appropriate clothing, protective goggles, and gloves.
Before start
Before beginning, be aware that the materials, a refrigerated centrifuge, a water bath or thermoblock, and ice are available.
Consider selecting the appropriate buffer to use, preparing it in advance, and keeping it at room temperature. Prepare the Z6 buffer if soft tissue (e.g., plant leaves and reproductive tissue) will be used for extraction. This buffer can be made as follows: prepare 38 g guanidine hydrochloride (to obtain 8 M), 0.195 g MES (to obtain 20 mM), 2 mL EDTA (from a 0.5 M stock), and dissolve to a final volume of 50 mL with distilled water. Prepare the GS buffer if hard tissue such as (woody) plant roots, plant tissue with higher levels of phenolic compounds, plant seeds, fungal mycelia, nematodes or arthropods will be used for extraction. This buffer can be made as follows: prepare 30 g guanidine isothiocyanate or guanidine thiocyanate, 0.25 g sodium lauroyl sarcosinate, 2.5 mL sodium citrate (from a 1 M stock), and dissolve to a final volume of 50 mL with distilled water.
Safety warnings
Remember to take precautions before working with detergents and solvents, such as wearing closed-toe shoes, appropriate clothing, protective goggles, and gloves.
Preparation of the laboratory space
Clean around the bench or work area with 70% ethanol and lay paper towels near the work area.
Procedure
Prepare fresh RNA buffer. The type of buffer depends on the type of tissue to be processed. In this procedure, we will refer to the GS buffer.
Use the Z6 buffer for soft plant vegetative and reproductive tissue (e.g., frozen leaves).
Use GS for hard tissue (e.g., [woody] plant roots), plant seeds, fresh fungal mycelia, nematodes, and arthropods.
Add 360 µL of ß-mercaptoethanol to 50 mL of buffer. Once added, cool the buffer before use. If the buffer is not in use, it can be stored at room temperature.
Cool the GS buffer on ice or at 4 °C.
When working with frozen tissue, grind 100 mg of tissue with LN2 to obtain a powder. Add the powder to an Eppendorf tube, then add the cool GS buffer to mix (Notes 1 and 2). If the buffer freezes when added to the tube, KEEP THE SAMPLES ON ICE AT ALL TIMES, until they can be vortexed. Vortex the samples for no more than two seconds.
Keeping the tubes on ice, open them and add 1 volume of phenol:chloroform (1:1 vol/vol).
Vortex the samples for no more than two seconds and return them to the ice. Mix the suspension well through vortexing until it turns white.
Centrifuge the suspension at 12,000 rpm for 10 min at 4 °C (Figure 1A).
Label new Eppendorf tubes and place them on ice.
Recover the upper phase (supernatant) without touching the interphase (white layer) and add it to a new Eppendorf tube.
Add an equal volume of phenol:chloroform (1:1 vol/vol), and centrifuge at 12,000 rpm for 10 min at 4 °C.
Repeat steps 8 and 9 until the white interface disappears or becomes completely transparent (Figure 1A; Note 3).
Label new Eppendorf tubes and place them on ice.
Recover the upper phase without disturbing the interphase (clear layer) and transfer it to a new Eppendorf tube.
Add an equal volume of cold absolute isopropanol, 1/10 of 3 M sodium acetate (pH 5.2; e.g., for 600 µL of volume, add 60 µL of sodium acetate), and incubate for 30 to 40 min at -20 °C (Note 4).
Centrifuge at 14,000 rpm for 15 min at 4 °C.
Wash the pellet with 500 µL of cold absolute ethanol and vortex until it comes off the bottom of the tube (Figure 1B).
Centrifuge at 14,000 rpm for 7 min at 4 °C.
Wash the pellet with 200 µL of absolute ethanol and centrifuge at 14,000 rpm for 7 min at 4 °C.
Decant the ethanol using the plastic beaker and leave the tubes inverted on a paper towel to dry the pellet for no more than 5 min.
Dissolve the pellet by adding nuclease-free water and incubating the tubes at 65 °C for 10 min (Note 5).
Quantify and analyze the total RNA (Note 6).
Notes
1. Add enough powder to fill 100 µL of the Eppendorf tube’s volume. The tissue volume can be increased to 200 µL, but then the GS buffer volume should be adjusted to 500 µL as well. The GS buffer volume cannot exceed 500 µL because a volume equal to the phenol:chloroform must be added later.
2. Always hold the tubes by the top or the cap. This will prevent the tube from opening and will prevent warming of the tube through contact with hands. Avoid touching the part of the tube that contains the sample to prevent RNA degradation.
3. Typically, the number of washes with phenol:chloroform (1:1 vol/vol) required depends on the type of tissue being extracted. Young plant leaf tissue, nematodes, and arthropods, require three washes; (woody) plant roots, plant seeds, and mycelia from fungi or oomycetes usually require four washes. Be careful not to touch the interface to avoid contaminating the RNA with proteins and/or carbohydrates.
4. If the amount of tissue used for the extraction was less than 30 mg, incubating for 3 hr or longer could help improve the RNA concentration. However, if the amount of tissue used was 100 mg or higher, incubation for more than 60 min could promote unwanted DNA contamination.
5. The amount of water used depends typically on the size of the pellet; however, to concentrate the RNA, add no more than 30 µL. If the pellet is not visible, add 20 µL.
6. Using this method, RNA concentrations in the range of micrograms per microliter are obtained from 100 µL of powdered tissue. Total RNA can be used directly to generate cDNA or can be subjected to a trace DNA cleanup before being used in library preparation for RNA-seq workflows.
Protocol references
Zhou D, Godinez-Vidal D, He J, Teixeira M, Guo J, Wei L, Van Norman JM, and Kaloshian I. 2023. A G-type lectin receptor kinase negatively regulates Arabidopsis immunity against root-knot nematodes. Plant Physiology 193: 721-735. DOI: 10.1093/plphys/kiad253.
Godinez-Vidal D, López-Leal G, Covarrubias AA, and Reyes JL. 2020. Early events leading to water deficit responses in the liverwort Marchantia polymorpha. Environmental and Experimental Botany 178: 104172. DOI: 10.1016/j.envexpbot.2020.104172.
Godinez-Vidal D, Rocha-Sosa M, Sepúlveda-García EB, Lozoya-Gloria E, Rojas-Martínez RJ, Guevara L, and Zavaleta-Mejía E. 2013.Transcript accumulation of mevalonic pathway genes and enzymatic activity of HMGCoA-r and EAS in chilli CM-334 infected by Nacobbus aberrans. Plant and Soil 372: 339-348. DOI: 10.1007/s11104-013-1743-8.
Longemann J, Schell J, and Willmitzer L. 1987. Improved method for the isolation of RNA from plant tissues. Analytical Biochemistry 163: 16-20. DOI: 10.1016/0003-2697(87)90086-8.
