Jan 14, 2026
DNAcrypt-AI protocol for generating and encrypting a secret key
- Marvin De los Santos1,
- Chikie Lynn1
- 1ChordexBio

Protocol Citation: Marvin De los Santos, Chikie Lynn 2026. DNAcrypt-AI protocol for generating and encrypting a secret key. protocols.io https://dx.doi.org/10.17504/protocols.io.14egn12k6v5d/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: January 10, 2026
Last Modified: January 14, 2026
Protocol Integer ID: 238375
Keywords: DNAcrypt-AI, FAS2rDNA, Covary, ChordexBio, DNA-based cryptography, human genome, k-mer encoding, sequence-based encryption, secret key generation, DNA sequence hashing, programmable biology, bioinformatics security, synthetic DNA computing, sequence reconstruction, machine learning–assisted encryption, cloud-based computational workflow, Jupyter Notebook, Google Colab, secret key dnacrypt, throughput dna sequence reconstitution pipeline, dnacrypt, using dnacrypt, digital credentials as genome coordinate, readable genome metadata representation, dna, genome vocabulary, encryption data, genome coordinate, sequence intelligence model, secret key, length secret key, decryption, plaintext key, based alphanumeric encoding, inspired cryptographic framework, alphanumeric encoding, cryptographic framework, expected secret key, password recovery, fas2rdna, pipeline, ai protocol, alphanumeric character, sequence, password, digital credential
Disclaimer
This protocol is licensed under CC-BY. DNAcrypt-AI software referenced here is subject to its own separate license terms. Please refer to the license notice at https://github.com/mahvin92/DNAcrypt-AI for your review. Note that this protocol was tested using the Colab implementation of a Jupyter notebook of DNAcrypt-AI. Additional validation may be required when applying this workflow to a different computing infrastructure. DNAcrypt-AI is provided for research use only and has not been validated for personal, clinical, diagnostic, and enterprise use. The software is provided "as is", without warranty of any kind, express or implied. For more information, please visit https://dnacryptai.chordexbio.com/.
Abstract
DNAcrypt-AI is a human genome-inspired cryptographic framework that leverages k-mer-based alphanumeric encoding, genome vocabulary, sequence reconstruction, and sequence-informed machine learning to generate and encrypt digital credentials as genome coordinates. This protocol describes the procedure for generating a fixed-length secret key (16 characters) and encrypting it using DNAcrypt-AI. Unlike passwords that may comprise alphanumeric characters and symbols, secret keys are devoid of symbols. The pipeline is implemented on Google Colab and integrates a high-throughput DNA sequence reconstitution pipeline (FAS2rDNA) with a sequence intelligence model (Covary) to randomly generate a plaintext key into an encrypted, non-human-readable genome metadata representation. The protocol generated the expected secret key of 16 characters long and provided an encryption data that can be used for decryption and password recovery.
Troubleshooting
Protocol procedure
Launch DNAcrypt-AI from https://dnacryptai.chordexbio.com/
Alternatively, you can run DNAcrypt-AI using the recommended methods described in https://github.com/mahvin92/DNAcrypt-AI.
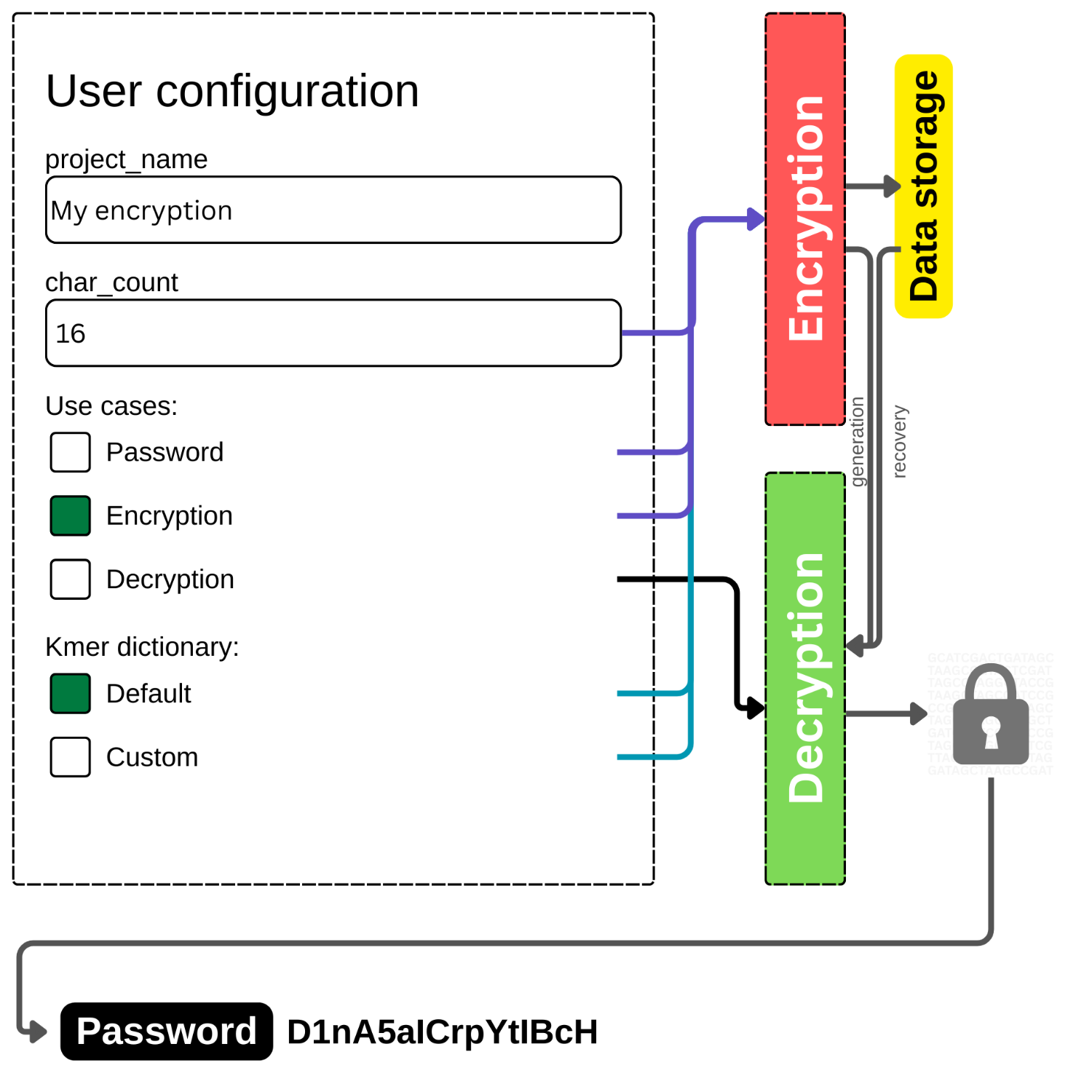
Create a configuration by modifying the user interface, as shown in Figure 1 below.
Figure 1. The interface of user configuration on DNAcrypt-AI, where users can specify the behavior of the cipher.
Type the number 16 in the char_count text box.
Note that users can generate secret keys with length from 6 to 90 characters.
Select the Encryption checkbox, ensuring to uncheck the other options under the Use cases section.
Run DNAcrypt-AI by initiating a runtime, ensuring a GPU (T4 as default) support, if available. Note that DNAcrypt-AI will still work in a non-GPU hosted runtime.
Initiate a runtime:
Click 'Runtime' -> Select 'Run all'
Wait for DNAcrypt-AI to finish running. The status and any error will be displayed below the code cell.
Allow the browser to download the DNAcrypt_metadata.json file. This is your encrypted data and is recommended to be renamed.
Note that if no download is produced, you can fetch the file from the /content/ directory through the File browser.
Save and store your encryption, digitally or physically by printing a copy.
Note that the encryption data is light and about 200kb in size only. When storing the data as a physical copy, ensure to print legibly to allow encoding to a digital copy later on for recovery.
Application note
This protocol is designed for generating secure, non-human-readable secret keys encoded as human genome coordinates. It is suitable for cryptographic demonstrations, secure credential generation, and experimental DNA-based security research.
