May 13, 2025
Version 2
Crystallisation of SARS-CoV-2 MPro in P212121 for compound soaking V.2
Version 1 is forked from Crystallization of SARS-CoV-2 Mpro
- Blake Balcomb1,2,3,
- Peter Marples1,2,3,
- Lizbé Koekemoer4,5,3,
- Charlie Tomlinson1,2,3,
- Daren Fearon1,2,3
- 1Diamond Light Source;
- 2Research Complex at Harwell;
- 3ASAP Discovery Consortium;
- 4Centre of Medicines Discovery;
- 5University of Oxford
- Blake Balcomb: The principle crystallographer on the SARS-CoV-2 Mpro project.;
- ASAP Discovery

Protocol Citation: Blake Balcomb, Peter Marples, Lizbé Koekemoer, Charlie Tomlinson, Daren Fearon 2025. Crystallisation of SARS-CoV-2 MPro in P212121 for compound soaking. protocols.io https://dx.doi.org/10.17504/protocols.io.261ge5emyg47/v2Version created by Mary-Ann Xavier
Manuscript citation:
Melissa L. Boby et al.,
Open science discovery of potent noncovalent SARS-CoV-2 main protease inhibitors.Science382,eabo7201(2023).DOI:10.1126/science.abo7201
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: February 24, 2025
Last Modified: May 13, 2025
Protocol Integer ID: 123294
Keywords: crystallisation, XChem, ASAP, AViDD, Diamond Light Source, i04-1, Mpro, SARS-CoV-2, crystallisation of sar, p212121 for compound, reproducible sar, crystallization protocol, xchem fragment screening, severe acute respiratory syndrome coronavirus, crystallisation, solvent test, strategies for pandemic preparedness, pandemic preparedness, sar, cov mpro
Funders Acknowledgements:
National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID)
Grant ID: Grant ID: U19AI171399
Disclaimer
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Acknowledgements:
Diamond Light Source Ltd, Harwell Science and Innovation Campus, Didcot OX11 0QX, UK
Research Complex at Harwell, Harwell Science and Innovation Campus, Didcot OX11 0FA, UK
Oxford Lab Technologies crystal shifter https://doi.org/10.1107/S2059798320014114
Abstract
The COVID-19 pandemic has highlighted the need to identify novel therapeutic interventions and strategies for pandemic preparedness against Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). This protocol outlines the crystallization protocol and buffer conditions used to obtain reproducible SARS-CoV-2 Mpro crystals suitable for XChem fragment screening. In this new version we added the solvent test, soaking conditions, seeding reference paper and addgene ids used in this experiment.
Materials
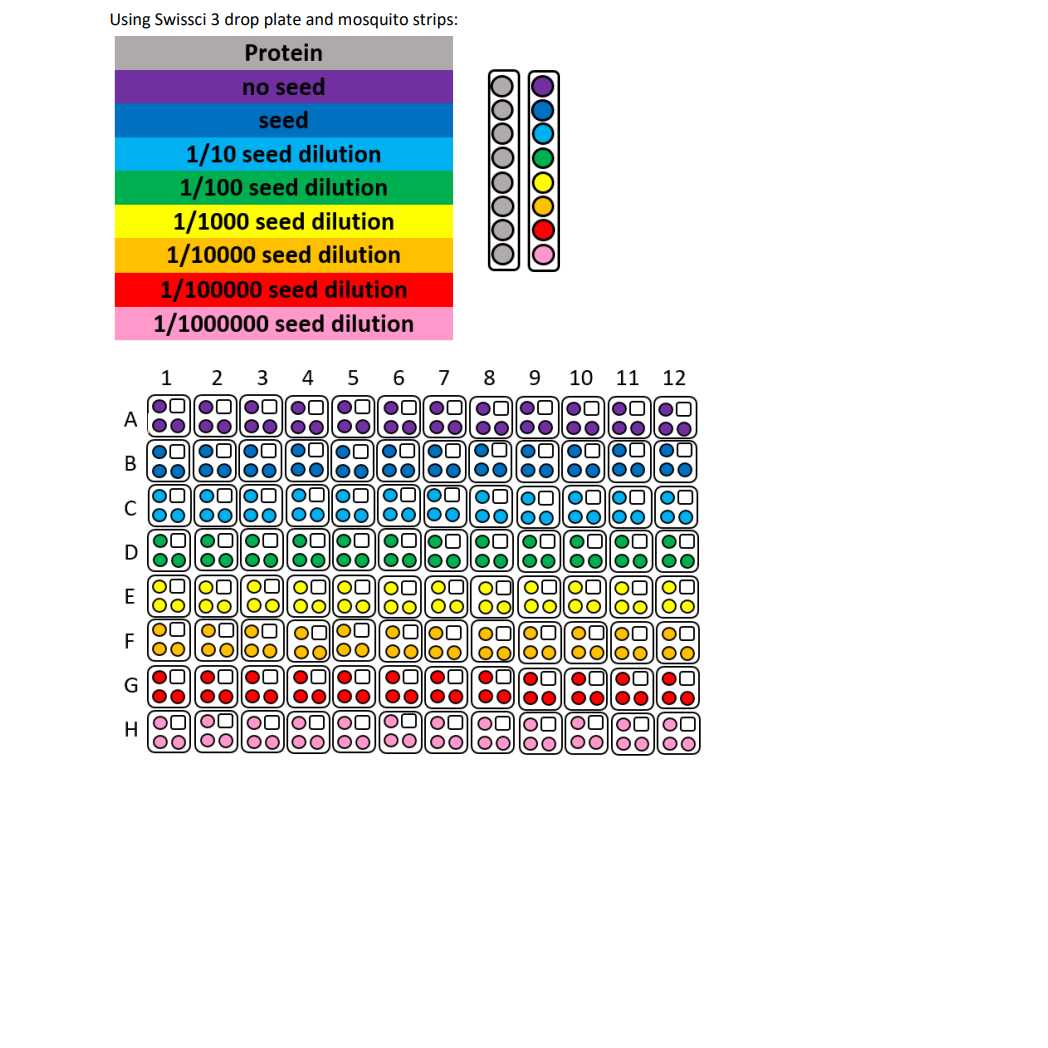
SwissCI 3 lens crystallization plates https://swissci.com/product/3-lens-crystallisation-plate/ Codes:
Midi: UVXPO-3LENS 3W96T-PS 3W96T-UVP
1 Molarity (M) MES adjusted to 6.7 with HCl, Molecular Dimensions, Catalog # MD2-013-PH 6.7
50% w/v PEG 4000, Molecular Dimensions, Catalog # MD2-250-11
99.9% DMSO, Molecular Dimensions, Catalog # MD2-250-159
Purified SARS-CoV-2 Mpro protein (5 mg/mL ) in 10 millimolar (mM) HEPES, 7.5 , 0.5 Molarity (M) NaCl, 5% glycerol, 0.5 millimolar (mM) TCEP
Protocol materials
SARS-CoV-2 isolate: Wuhan-Hu-1 Mpro (3CL protease)addgeneCatalog #228666
SARS-CoV-2 Mpro protease (C145S mutant - catalytically inactive); aliases: 3CLpro, 3C-likeaddgeneCatalog #204769
Troubleshooting
Safety warnings
Follow all handling warning for the chemicals used in the crystalllisation screen composition.
SARS-CoV-2 Mpro expression and purification
The protein used for crystallisation was expressed and purified using the following protocol and plasmid SARS-CoV-2 isolate: Wuhan-Hu-1 Mpro (3CL protease)addgeneCatalog #228666
Protocol
CREATED BY
Korvus Wang
Equipment needed
Formulatrix Rock Imager (or incubator of choice)
Equipment
Mosquito HV
NAME
High Volume 16-Channel Robotic Liquid Handler
TYPE
SPT LabTech
BRAND
3097-01057
SKU
LINK
P100 8 multi-channel pipette
Crystallisation experiment
1d
Preparation of seed stock using imatureSARS-CoV-2 Mpro protease (C145S mutant - catalytically inactive); aliases: 3CLpro, 3C-likeaddgeneCatalog #204769 Mpro and mature protein: Protocol for obtaining SARS-CoV-2 Mpro P212121 crystal form can be found here: DOI: https://doi.org/10.1016/j.jmb.2021.167118.
1: 250 dilution Sample seeds
17m 40s
Protein and buffer requirements:
43.2 µL 5 mg/mL Sample
1.92 mL Crystallisation screen
14.4 µL Sample P212121 crystal form seeds (see step 3), dilution 1:250
Crystallisation screen composition:
0.1 Molarity (M) MES 6.7
11% w/v PEG 4000
5% DMSO
Stock solutions used:
1 Molarity (M) MES adjusted to 6.7 with HCl
50% w/v PEG 4000
99.9% DMSO
Note
The crystallisation screen can be stored in a duran bottle or aliquoted into 96 deep well block for easy dispensing into SwissCI 3 lens plates.
For long term storage keep the crystallisation screen in the fridge at 4°C.
Dispense 20 µL Crystallisation screen into SwissCI 3 lens plate reservoir wells using a 100 µl multi-channel pipette.
Dispense 150 nL Sample to each lens using the SPT mosquito.
Dispense 150 nL Crystallisation screen to each lens using the SPT mosquito.
Dispense 50 nL Seeds P212121 stock to each lens using the SPT mosquito.
Drop ratio: 3:3:1 ratio (150 nL Sample : 150 nL reservoir solution: 50 nl seeds)
Final drop volume: 350 nl
10m
Incubate at 20 °C for 24:00:00 h in Formulatrix Rock Imager.
Imaging Schedule: The first images are taken after 12 h and the imaging schedule follows a Fibonacci sequence of days for further collections.
1d
Crystal form after ~24 h.
Expected result
The crystals reach their maximum size after 48 h.
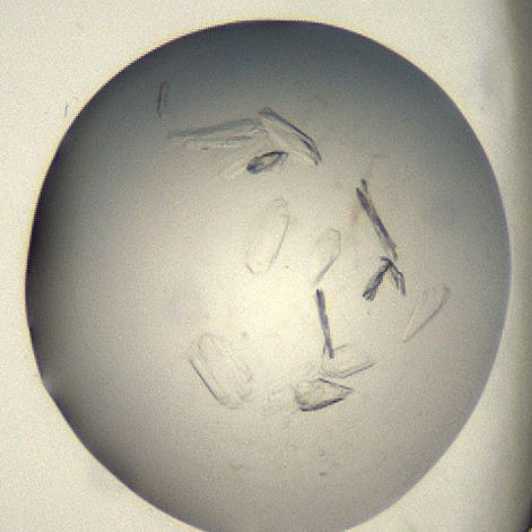
Crystals typically form either as stacked playes or in small clusters containng 4-6 crystals.
Morphology: plates.
Size: ~100 μm in length and ~20 μm in width, depth of the crystals is ~10 μm
Appearance: glass shard.
Average resolution: 1.8 Å
Space group: P212121
Unit cell: 67 Å, 99 Å, 102 Å
90.00°, 90.00°, 90.00°

An example of a drop containing SARS-CoV-2 Mpro protease crystals.
Data collection at Synchrotron
Diamond Light Source
Unattended Data Collection (UDC)
Data Collection Temperature: 100K
Detector: DECTRIS EIGER2 X 9M
Beamline: I04-1
Wavelength: 0.9212 Å
Resolution (Å): 1.78
Beam Size (μm): 60 X 50
Number of images: 3600
Oscillation: 0.10°
Exposure (s): 0.0020
Transmission (%): 100
Flux (ph/s): 9.50e+11
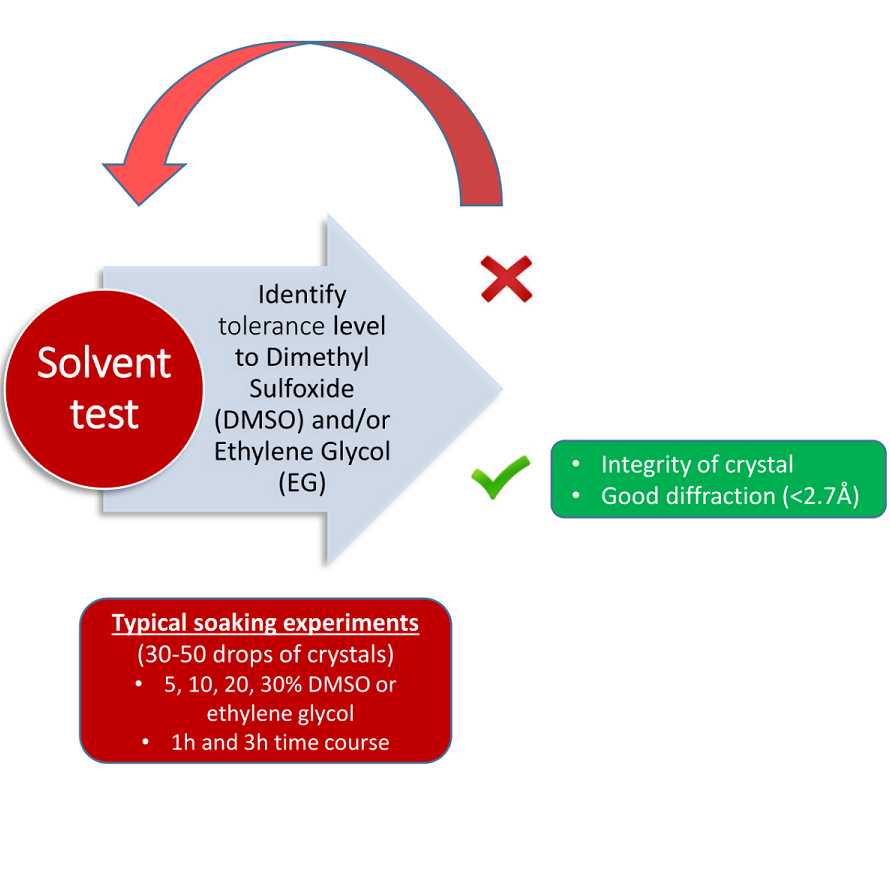
Soaking Tolerance
Crystals can tolerate up to 20% DMSO, 1-2h
Compound Soaking
Crystals can tolerate up to 20% DMSO, 1-2h
Protocol references
For SARS-CoV-2 Main protease in orthorhombic space group
Group deposition SARS-CoV-2 main protease in complex with inhibitors from the COVID Moonshot