Oct 03, 2025
CRISPR/Cas9 knock-in of PINK1-GFP at the endogenous PINK1 locus in U-2 OS cells
- Tom Snelling1,
- Miratul Muqit1
- 1University of Dundee

Protocol Citation: Tom Snelling, Miratul Muqit 2025. CRISPR/Cas9 knock-in of PINK1-GFP at the endogenous PINK1 locus in U-2 OS cells. protocols.io https://dx.doi.org/10.17504/protocols.io.dm6gpm38pgzp/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: August 14, 2025
Last Modified: October 03, 2025
Protocol Integer ID: 224685
Keywords: PINK1, CRISPR, PINK1-GFP, endogenous pink1 gene with pink1, endogenous pink1 gene, gfp at the endogenous pink1 locus, terminus of the endogenous pink1 locus, gfp positive cell, encoding pink1, crispr, expressed pink1, endogenous pink1 locus, cas9 nickase, gfp cdna, in of pink1, pink1, gfp knock, cas9, gene
Abstract
A CRISPR/Cas9-based strategy is presented for generating U-2 OS cells with a PINK1-GFP knock-in at the N-terminus of the endogenous PINK1 locus (replacing the endogenous PINK1 gene with PINK1-GFP cDNA). The method uses Cas9 nickase and paired sgRNAs to introduce a double-stranded break in exon 1 of PINK1, enabling homology-directed repair using a donor plasmid encoding PINK1-GFP cDNA flanked by approximately 500 bp homology arms as the template. Following plasmid transfection, cells undergo a 10 day recovery period to allow depletion of transiently expressed PINK1-GFP. Mitochondrial depolarisation is then induced using antimycin and oligomycin to increase PINK1-GFP expression levels and therefore enhance GFP signal detection. GFP positive cells were isolated by fluorescence-activated cell sorting into 96-well plates using a low threshold gate to capture clones with physiologically relevant expression and were subsequently expanded and validated by immunoblotting.
Guidelines
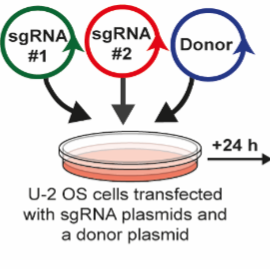
Overview of the workflow
Overview of the CRISPR/Cas9 PINK1-GFP knock-in strategy
Materials
Cell lines
Note: Cells are cultured in an incubator at 37 °C with 5% CO₂
1. Wildtype U-2 OS cells (ATCC HTB-96)
1. DU64062 encoding Cas9 D10A nickase and sgRNA #1 (sense)
2. DU64072 encoding puromycin resistance and sgRNA #2 (antisense)
3. DU64119 encoding PINK1-GFP cDNA flanked by ~500 bp homology arms
Reagents
1. Dulbecco’s Modified Eagle Medium (DMEM) (Gibco, catalog number: 11960-085)
2. Fetal bovine serum (FBS)
3. L-glutamine, 200 mM (Gibco, catalog number: 25030-024)
4. Penicillin–Streptomycin, 100X (Gibco, catalog number: 15140-122)
5. Opti-MEM I Reduced Serum Medium (OptiMEM) (Gibco, catalog number: 31985-062)
6. Trypsin-EDTA solution, 0.25% (Gibco, catalog number: 25200-056)
7. Polyethylenimine Max (Polysciences, catalog number: 24765-1)
8. Puromycin dihydrochloride (Sigma-Aldrich, catalog number: P9620)
9. Antimycin A (Sigma-Aldrich, catalog number: A8674)
10. Oligomycin (Sigma-Aldrich, catalog number: 75351)
11. Porcine gelatin (Sigma-Aldrich, catalog number: G1890)
12. Phosphate-buffered saline (PBS) (Gibco, catalog number: 10010023)
13. DAPI (Biolegend, catalog number: 422801)
Consumables
1. 10 cm Nunc cell culture dishes (Thermo Fisher, catalog number: 150318)
2. 96-well Nunc cell culture plates (Thermo Fisher, catalog number: 167008)
3. 25 mL sterile reagent reservoirs (Thermo Fisher, catalog number: 11405758)
4. 50 mL conical centrifuge tubes (Greiner, catalog number: 227261)
5. 15 mL conical centrifuge tubes (Greiner, catalog number: 188271)
6. Serological pipettes, sterile (Thermo Fisher, catalog number: 10710810)
7. 0.22 µm sterile syringe filter (Millipore or equivalent)
8. 0.45 µm sterile syringe filter (Millipore or equivalent)
9. 1.5 mL sterile microcentrifuge tubes
Recipes
(a) 50 mL of 0.1% (w/v) gelatin solution
1. Weigh 50 mg of porcine gelatin powder into a 50 mL conical centrifuge tube
2. Add 40 mL of sterile distilled water
3. Warm to 37 °C until fully dissolved
4. Bring volume to 50 mL using sterile distilled water
5. Filter sterilise through a 0.22 µm sterile syringe filter
6. Store at 4 °C for up to 1 week
(b) 50 mL of 1 mg/mL PEI max solution
1. Weigh 100 mg of PEI max (linear, 40 kDa) into a sterile bottle
2. Add 80 mL of sterile distilled water
3. Adjust pH to 7.0 using 1 M sodium hydroxide
4. Bring volume to 100 mL with sterile distilled water
5. Filter sterilise through a 0.22 µm sterile syringe filter
6. Aliquot in 1.5 ml sterile microcentrifuge tubes and store at -20 °C
Troubleshooting
Before start
Before starting this procedure, a confluent 10 cm dish of wildtype U-2 OS cells, no higher than passage 30, is required.
Transfection
2w 1d 0h 55m
In the afternoon, aspirate the culture media from a confluent 10 cm dish of cells and replace with 10 mL of PBS using a serological pipette.
Culture media (1 bottle)
| Reagent | Final concentration | Quantity to add | |
| DMEM | Not applicable | 500 mL | |
| FBS | 10% (v/v) | 50 mL | |
| 200 mM L-Glutamine | 2 mM | 5.6 mL | |
| Penicillin-Streptomycin 100X | 1X | 5.6 mL |
Recipe for culture media (1 bottle)
2m
Aspirate the PBS and add 3 mL of trypsin-EDTA solution to the dish. Return the dish to the incubator until the cells have detached, which should take 3-5 min for this cell line.
5m
Add 10 mL of antibiotic-free culture media to the plate and pipette up and down using a serological pipette until a single-cell suspension has been produced. Transfer to a 15 mL canonical centrifuge tube.
Antibiotic-free culture media (1 bottle)
| Reagent | Final concentration | Quantity to add | |
| DMEM | Not applicable | 500 mL | |
| FBS | 10% (v/v) | 50 mL | |
| 200 mM L-Glutamine | 2 mM | 5.6 mL |
Recipe for antibiotic-free culture media (1 bottle)
2m
Remove 20 µL of the cell suspension and dilute with 80 µL of culture media. Mix 20 µL of the diluted cell suspension with 20 µL of trypan blue solution and count the number of cells using standard methods, ensuring that the cell viability is at least 90%.
5m
Plate 4 million wildtype U-2 OS cells into a 10 cm dish and add media up to a total volume of 10 mL.
2m
Ensure that cells are evenly distributed by moving plates in a figure-of-8 motion prior to returning them to the incubator for 18 h at which point the confluency should be 60-70%.
18h
In a 1.5 ml microcentrifuge tube, mix 1 µg of sgRNA #1 (DU64062), 1 µg of sgRNA #2 (DU64072) and 3 µg of the donor template (DU64119).
2m
Add 1 ml of OptiMEM to the microcentrifuge tube and invert five times to mix thoroughly.
2m
Add 20 µl of 1 mg/ml PEI to the diluted plasmid mixture, vortex for 5 seconds and incubate at room temp for 30 min to allow DNA-lipid complexes to form.
30m
Add the DNA-lipid complexes dropwise to the 10 cm dish of U-2 OS cells and return to incubator for 24 h.
1d
Aspirate the culture media from the 10 cm dish and replace with selection media (culture media supplemented with 2 µg/ml puromycin) and return to incubator for 24 h.
1d
Aspirate the culture media from the 10 cm dish and replace with 8 ml of antibiotic-free media and return to incubator for 2 h.
2h
Repeat the transfection procedure by following steps 7-10 again, but do not perform an additional puromycin selection.
1d
Cells were passaged and expanded for an additional 10 days in culture media to allow transient expression of PINK1-GFP from the donor plasmid to diminish. During this expansion, collect the culture media at each passage, filter it through a 0.45 µm filter to remove cells, and store this pre-conditioned media at 4 °C.
1w 3d
Coat five 96-well plates with 0.1% (w/v) gelatin solution:
Transfer 25 mL of the 0.1% (w/v) gelatin solution into a 25 mL sterile reagent reservoir and use a multi-channel pipette to transfer 50 µL of 0.1% (w/v) gelatin solution to wells of five 96-well plates.
5h
Return the 96-well plates to the incubator for 4 h and then aspirate the gelatin solution and air dry for 1 h.
5h
Top up the pre-conditioned media with an additional 20% (v/v) of fresh FBS and transfer to a 25 mL sterile reagent reservoir.
5m
Use a multi-channel pipette to transfer 100 µL of the supplemented pre-conditioned media to each well of the five 96-well plates and return to the incubator overnight.
18h
Sorting and expansion
9w 4d 19h 15m
After 10 days of expansion (step 14), the culture media was aspirated and replaced with culture media supplemented with 5 µM antimycin and 0.63 µM oligomycin and returned to the incubator for 18 h.
18h
The culture media was aspirated and the cells were washed three times with culture media to remove the antimycin and oligomycin and returned to the incubator for 24 h to aid recovery.
1d
Cells were detached using Trypsin-EDTA (see steps 1–2). Culture medium was added to the plate, and the cells were gently pipetted up and down with a serological pipette until a single-cell suspension was obtained. The suspension was then transferred to a 15 mL conical centrifuge tube.
5m
Cells were centrifuged at 800xg for 5 min at room temp and the supernatant then aspirated.
5m
The cell pellet was resuspended in DMEM supplemented with 1% (v/v) FBS and 0.1 µg/mL DAPI.
5m
Fluorescence-activated cell sorting (FACS) was used to deposit single GFP positive cells into the coated 96-well plates containing pre-conditioned medium. The GFP gate was set low to capture cells with lower GFP levels, as cells with the highest GFP signal were predicted to have defective PINK1 processing or potential off-target integration of PINK1-GFP.
GFP gating strategy
1h
Monitor the 96-well plates until individual wells reach confluency, typically 4–6 weeks after FACS.
6w
Expand single-cell colonies from each well into 10 cm culture dishes. Grow the cells until they reach confluency again, typically 2–3 weeks.
3w
Screen the expanded colonies for successful knock-in of PINK1-GFP into the PINK1 locus by performing immunoblotting with antibodies recognising PINK1 and GFP using standard methodologies.
3d
