Nov 11, 2025
Controlled addition of short inosine tails to capture both polyadenylated and non-polyadenylated RNA for direct RNA sequencing on nanopores.
- Jason Talkish1,
- Justin Sim1,
- Jenny Mai Vo1,
- Mark Akeson2,
- Manuel Ares, Jr.1
- 1RNA Center, Genomics Institute, and Department of Molecular, Cell & Developmental Biology, UC Santa Cruz, Santa Cruz, California, USA;
- 2Biomolecular Engineering Department, UC Santa Cruz, Santa Cruz, California, USA
- Ares Lab

External link: http://ribonode.ucsc.edu/index.html
Protocol Citation: Jason Talkish, Justin Sim, Jenny Mai Vo, Mark Akeson, Manuel Ares, Jr. 2025. Controlled addition of short inosine tails to capture both polyadenylated and non-polyadenylated RNA for direct RNA sequencing on nanopores.. protocols.io https://dx.doi.org/10.17504/protocols.io.6qpvrwz4plmk/v1
Manuscript citation:
Vo J, Mulroney L, Quick-Cleveland J, Jain M, Akeson M, Ares M Jr. 2021. Synthesis of modified nucleotide polymers by the poly(U) polymerase Cid1: Application to direct RNA sequencing on nanopores. http://dx.doi.org/10.1101/2021.07.06.451372.
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: November 06, 2025
Last Modified: November 11, 2025
Protocol Integer ID: 231682
Keywords: Inosine tailing, Nanopore, Sequencing, Direct RNA sequencing, Cid1, polyadenylated rna, tailed rna, end of rna, heterogenous inosine tail, rna, individual rna molecule, wide variety of cellular rna, addition of short inosine tail, cellular rna, short inosine tail, inosine tailing, nucleotide during transcription, rna for direct rna, direct rna, cid1 enzyme, incorporated nucleotide, schizosaccharomyce, heterogeneity of rna, modified nucleotide, nanopore, deoxynucleotide end, variety of ncrna, inosine, nanopore technology, transcription, ncrna, histone mrna, oxford nanopore platform, enzyme
Funders Acknowledgements:
Manuel Ares Jr.
Grant ID: 5R35GM145266
Mark Akeson
Grant ID: Oxford Nanopore Technologies royalties
Disclaimer
M. Akeson received reimbursement for travel, accommodation, and conference fees to speak at events organized by Oxford Nanopore Technologies (ONT). M. Akeson holds shares in ONT, is a paid consultant to ONT, has received research funding from ONT, and is an inventor on UC patents licensed to ONT. A patent, as yet unlicensed, has been awarded to the University of California naming Jenny Vo, M. Akeson, and M. Ares Jr. as inventors concerning the use of Cid1 for addition of modified nucleotides.
Abstract
Nanopore technology allows full length sequencing of individual RNA molecules. However, capturing and sequencing all cellular RNAs in a complex mixture is challenging due to heterogeneity of RNA 3’ end structures. The Schizosaccharomyces pombe enzyme Cid1 can bind and add a variety of natural and modified nucleotides to the 3’ end of RNAs (Munoz-Tello et al. 2014; Vo et al. 2021; Wiegand et al. 2025). This approach has been shown to be useful for tailing and capturing a wide variety of cellular RNAs, regardless of their 3’ ends. Inosine tailing is exceptionally promising, since it is not a naturally incorporated nucleotide during transcription and produces a very distinct current trace compared to natural or in vitro produced poly(A) tails. In vitro, Cid1 adds approximately 50 inosines to endogenous poly(A) RNAs. However, non-poly(A) RNAs often acquire much longer, heterogenous inosine tails (Vo et al. 2021). The protocol presented here introduces the addition of dITP in an appropriate ratio to rITP such that the very long I-tails observed on non-polyadenylated RNAs are reduced to an average of ~15-40 nt (Fig 1). The Cid1 enzyme incorporates dITP at a lower rate than rITP, but is unable to add to a deoxynucleotide end; thus, dITP serves as a chain terminator (Fig 1). These inosine-tailed RNAs now carry rI(n)dI ends and can then be used for adapter ligation and sequencing on Oxford Nanopore Platforms to capture both poly(A)+ mRNAs and histone mRNAs, rRNAs and a variety of ncRNAs (Fig 2).
Image Attribution
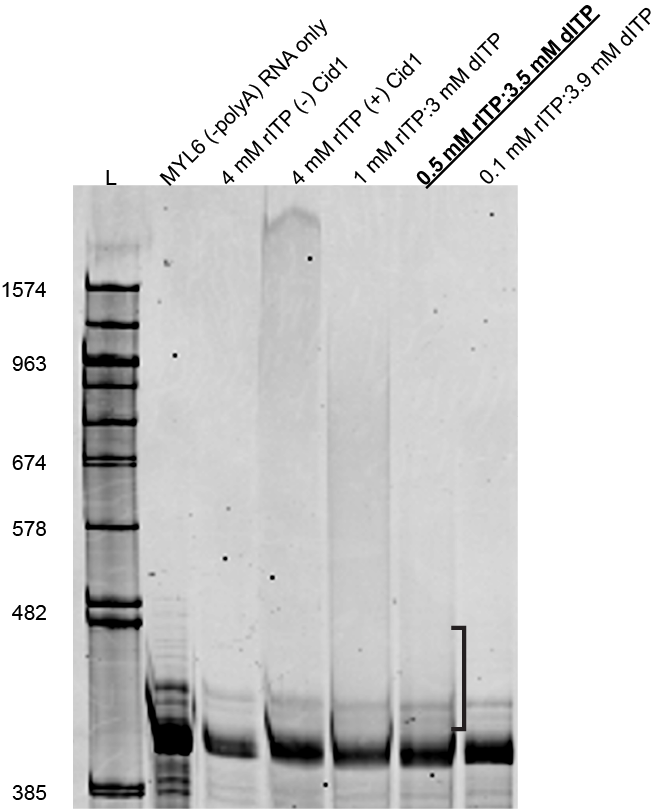
Figure 1: In vitro transcribed MLY6 RNA (without a polyA tail) was incubated with or with Cid1 enzyme in the presence of various ratios of rITP and dITP. As shown in Vo et al. (2021) “rITP only” produces inosine tails much greater than the 1574 nt marker (Lane 3). A 1:7 rITP:dITP ratio produces average tail lengths less than 100 nt (Lane 6). RNAs were resolved on a 6% polyacrylamide gel and stained with Sybr Gold.
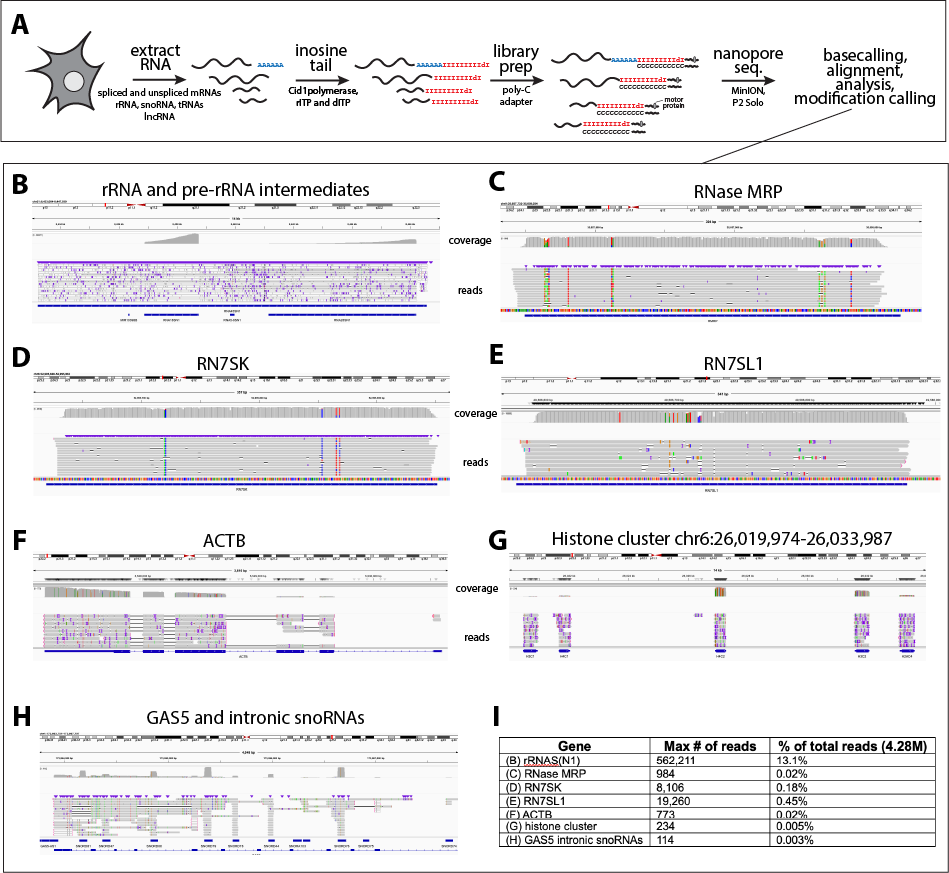
Figure 2: Schematic and example IGV browser images of various classes of RNAs inosine tailed, captured and sequenced using direct RNA sequencing on Oxford Nanopore Technologies P2 Solo. A) Example representation of RNA extraction, inosine tailing, library preparation, nanopore sequencing and analysis. B-H) IGV browser images of various classes and sizes of noncoding, and polyadenylated RNAs I-tailed and captured using the above protocol. I) Summary table of number of reads
Materials
- Total RNA (5 ug) extracted from HEK293 cells
- 10X NEBuffer 2 (NEB B7002S). 1X buffer: 50 mM NaCl, 10 mM Tris-HCl, 10 mM MgCl2, 1 mM DTT, pH 7.9 @ 25°C
- 25 mM MgCl2 (Thermo Scientific R0971)
- 1M Tris-HCl, pH 7.9
- 100 mM (2'-deoxyinosine 5'-triphosphate) dITP solution (Thermo Scientific R1191)
- Inosine 5’-triphosphate trisodium salt (rITP) (Milipore Sigma I0879-50MG)
- Poly(U) polymerase 2U/ul (NEB M0337S)
- RNase-free H2O
- Adapter oligonucleotides (IDT)
- Top oligo (100 uM, H2O): 5PHOS/GGCTTCTTCTTGCTCTTAGGTAGTAGGTTC
- 10C splint (100 uM, H2O): CCTAAGAGCAAGAAGAAGCCCCCCCCCCCC
Troubleshooting
Before start
In heat blocks, or a thermocycler, anneal oligos by incubating at 95 °C for 2 minutes, 65 °C for 10 minutes, 48 °C for 10 minutes, then place at room temperature for 10 minutes. Place on ice. Annealed oligos can be frozen and used multiple times.
Preparing reagents
Extract total RNA using standard Trizol protocols. (Rio et al. 2011)
Quantify RNA using a NanoDrop (Thermo Scientific), and evaluate integrity of the RNA using a Bioanalyzer or TapeStation (Agilent Technologies).
Prepare 30 mM rITP in 60 mM Tris-HCl pH 7.9.
Prepare 30 mM dITP in 60 mM Tris-HCl pH 7.9.
Prepare a 30mM mixture of 1:7 rITP:dITP.
Annealing oligonucleotides for custom adapters
Resuspend oligonucleotides to 100 uM in RNase-free H2O.
Mix 10 ul each of Top oligo and 10C splint. Bring the volume up to 100 ul with RNase-free H2O.
In heat blocks, or a thermocycler, anneal oligos by incubating at 95 °C for 2 minutes, 65 °C for 10 minutes, 48 °C for 10 minutes, then place at room temperature for 10 minutes. Place on ice. Annealed oligos can be frozen and used multiple times.
Inosine tailing reaction setup
Library preparation and sequencing of inosine-tailed RNA on Oxford Nanopore, Minion or P2 Solo
Prepare the sequencing library and sequence on the Minion or P2 Solo using standard Oxford Nanopore Technologies protocols for direct RNA sequencing using 1 ug of inosine-tailed RNA and the following exceptions.
Instead of using the RTA adapter provided by ONT, replace with 1 ul of 10 uM, annealed custom adapter.
Reverse transcription is optional. Sequencing and images in Figure 1 were produced without reverse transcription.
Figures
Figure 1: In vitro transcribed MLY6 RNA (without a poly(A) tail) was incubated with or with Cid1 enzyme in the presence of various ratios of rITP and dITP. As shown in Vo et al. (2021) “rITP only” produces inosine tails much greater than the 1574 nt marker (Lane 3). A 1:7 rITP:dITP ratio produces average tail lengths less than 100 nt (Lane 6). RNAs were resolved on a 6% polyacrylamide gel and stained with Sybr Gold.
Figure 2: Schematic and example IGV browser images of various classes of RNAs inosine tailed, captured and sequenced using direct RNA sequencing on Oxford Nanopore Technologies P2 Solo. A) Example representation of RNA extraction, inosine tailing, library preparation, nanopore sequencing and analysis. B-H) IGV browser images of various classes and sizes of noncoding, and polyadenylated RNAs I-talied and captured using the above protocol. I) Summary table of number of reads shown in browser shots in (B-H) and the percent of total reads of the MinION sequencing run (4.28M reads).
Protocol references
Munoz-Tello P, Gabus C, Thore S. 2014. A critical switch in the enzymatic properties of the Cid1 protein deciphered from its product-bound crystal structure. Nucleic Acids Res 42: 3372–3380.
Rio D, Ares M, Nilsen T. 2011. RNA. Cold Spring Harbor Laboratory Press, New York, NY.
Vo J, Mulroney L, Quick-Cleveland J, Jain M, Akeson M, Ares M Jr. 2021. Synthesis of modified nucleotide polymers by the poly(U) polymerase Cid1: Application to direct RNA sequencing on nanopores. http://dx.doi.org/10.1101/2021.07.06.451372.
Wiegand DJ, Rittichier J, Meyer E, Lee H, Conway NJ, Ahlstedt D, Yurtsever Z, Rainone D, Kuru E, Church GM. 2025. Template-independent enzymatic synthesis of RNA oligonucleotides. Nat Biotechnol 43: 762–772.