Jan 09, 2026
BioID: Identifying Protein-Protein Interactions in Living Cells
- Aleksandar Bartolome1,
- Julia Heiby1,
- Ivonne Heinze1,
- Therese Dau1,
- Alessandro Ori1
- 1FLI Leibniz Institute on Aging

Protocol Citation: Aleksandar Bartolome, Julia Heiby, Ivonne Heinze, Therese Dau, Alessandro Ori 2026. BioID: Identifying Protein-Protein Interactions in Living Cells. protocols.io https://dx.doi.org/10.17504/protocols.io.14egn6326l5d/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: May 23, 2024
Last Modified: January 09, 2026
Protocol Integer ID: 100351
Keywords: promiscuous biotin ligase, biotinylated protein, using streptavidin affinity purification, streptavidin affinity purification, dependent biotin identification, protein interactions within living cell, proximate protein, protein interactions in living cell, identifying protein, protein, expressing hek cell, hek cell, protein interaction, protein of interest, mass spectrometry, bioid, living cell, expression in cell
Disclaimer
DISCLAIMER – FOR INFORMATIONAL PURPOSES ONLY; USE AT YOUR OWN RISK
The protocol content here is for informational purposes only and does not constitute legal, medical, clinical, or safety advice, or otherwise; content added to protocols.io is not peer reviewed and may not have undergone a formal approval of any kind. Information presented in this protocol should not substitute for independent professional judgment, advice, diagnosis, or treatment. Any action you take or refrain from taking using or relying upon the information presented here is strictly at your own risk. You agree that neither the Company nor any of the authors, contributors, administrators, or anyone else associated with protocols.io, can be held responsible for your use of the information contained in or linked to this protocol or any of our Sites/Apps and Services.
Abstract
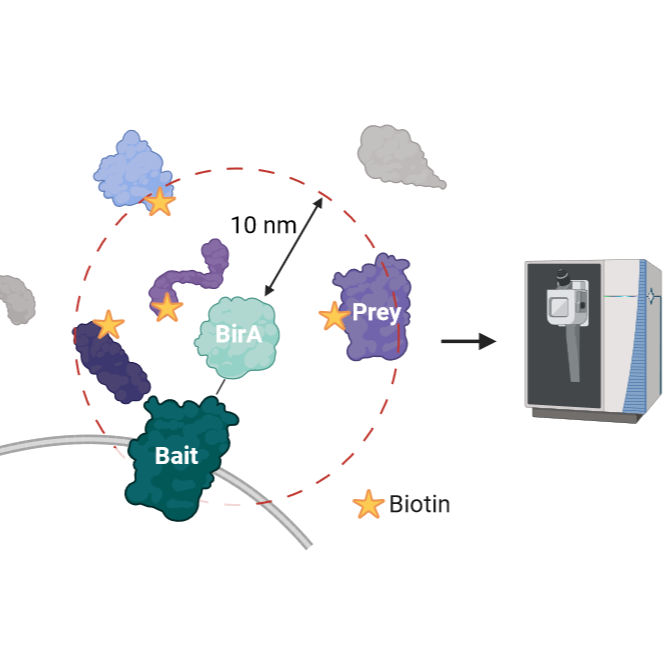
The BioID (proximity-dependent biotin identification) method represents a powerful approach for identifying protein-protein interactions within living cells. This technique utilizes a promiscuous biotin ligase, BirA*, which is fused to a protein of interest (bait). Upon expression in cells, BirA* biotinylates proximate proteins (prey) in the vicinity of the bait, which can then be isolated using streptavidin affinity purification.
Here, we describe in detail how to process stably expressing HEK cells containing BirA*-fusion protein and how to capture the biotinylated proteins which are to be subsequently identified through mass spectrometry (DIA).
Materials
Equipment:
- Lasagna plates: Corning 500cm² Square BioAssay Dish (#431110)
- Teat: Pasteur pipette rubber bulb (Carl Roth, #8404.1)
- 50ml Falcon tubes (Corning, #352070)
- 15ml falcon tube (Corning, #352096)
- Serological Pipettes (Greiner, cellstar)
- Liquid nitrogen
- 1.5ml tubes (Eppendorf, #0030 120.086)
- Ice
- Low-binding pipette tips (Sarsted)
- Pierce Spin Columns Snap Cap (Thermo Fisher Scientific, #69725)
- Macro Spin Columns (Harvard Apparatus, #74-4101)
- pH-indicator strips
- Safety cabinet (Thermo, #Safe S2020 1.2)
- Automated cell counter (Biorad, #TC20)
- Trypan blue solution for cell counting (Sigma, #T8154)
- Counting Slides for cells (Biorad, #145-0011)
- Incubator (Thermo, #BBD 6220, CO2 Incubator), settings: 5% CO2, 95%rH, 37°
- Milli-Q water system (Merck, Advantage A10)
- Bioruptor plus (#B01020001, Diagenode)
- Centrifuge 5810R (#5811000015, Eppendorf)
- Heat-block (#SBH130D3 or #SBH200D3, Stuart)
- Thermo Mixer C (#5382000015, Eppendorf)
- OASIS Oasis 96-well plate vacuum manifold (# 186001831, Waters)
- Waters Oasis HLB μElution plates 30 µg (for maximum 100 µg proteins) (#186001828BA, Waters)
- Macro spin columns C-18 (#74-4101, Harvard Apparatus)
- SpeedVac (Concentrator Plus/Vacofuge Plus, #5305000100, Eppendorf)
- Glass vials and glass inserts for LC-MS (#88909355 vials, #93909134 inserts, #88849362 caps, VDS optilab)
- Orbitrap Exploris 480 Mass Spectrometer (Thermo Fisher Scientific)
Reagents:
- Streptavidin coated Sepharose beads (Merck, #GE17-5113-01)
- Sulfo-NHS-Acetate (Thermo Fisher Scientific, #26777)
- LysC (Wako, #125-05061, sequencing grade)
- Trypsin (for cell culture) (Thermo Fisher Scientific, #25300-062)
- Trypsin (Mass Spectrometry Grade) (Promega, #V511)
- DMEM high glucose 4.5 g/l (Sigma Aldrich, #D6429)
- FBS (Gibco, #10270-106)
- Biotin (Carl Roth, #3822.1)
- Tetracycline (Sigma Aldrich, #87128)
- Aprotinin (Carl Roth, #A162.3)
- Leupeptin (Carl Roth, #CN33.2)
- Turbonuclease (MoBiTec GmbH, #GE-NUC10700-01)
- Trizma base (Carl Roth, #4855.2)
- Ammonium Bicarbonate (Carl Roth, #T871.2)
- HEPES (Sigma Aldrich, #H3375)
- NaCl (Carl Roth, #3957.1)
- EDTA (Carl Roth, #8043.2)
- EGTA (Carl Roth, #3054.1)
- Triton X-100 (Sigma-Aldrich, #3051.3)
- SDS (Sigma Aldrich, #75746)
- Acetonitrile (Biosolve, #0001204102BS)
- Trifluoroacetic acid (Biosolve, #0020234131BS)
- Methanol (Biosolve, #0013684102BS)
- Formic Acid (Carl Roth, #4724.3)
Buffers
- PBS
- Lysis buffer: 50 mM Tris, 150 mM NaCl, 1 mM EDTA, 1 mM EGTA, 1% Triton, 0.1% SDS, 1.5 µM Aprotinin, 10 µM Leupeptin, 250U Turbonuclease
- Acetylation buffer: 10 mM Sulfo-NHS acetate
- Wash buffer: 50 mM AmBic, pH 8.3
- Digest buffer: 0.5 µg LysC in 50 mM AmBic
- Elution buffer: 10% TFA in ACN
- Maintenance buffers: 20% ACN
Abbreviations
ACN - Acetonitrile
AmBic - Ammonium Bicarbonate
DIA – Data Independent Acquisition
DMEM - Dulbecco's Modified Eagle Medium
EDTA – Ethylene Diamine Tetraacetic acid
EGTA - (Ethylene Glycol-bis(β-aminoethylether)-N,N,N′,N′-Tetraacetic acid
FBS - Fetal Bovine Serum
HEPES - 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
LysC - Native endoproteinase from Lysobacter enzymogenes
MetOH – Methanol
PBS - Phosphate-Buffered Saline
RT - Room Temperature
SDS - Sodium Dodecyl Sulfate
TFA - Trifluoroacetic acid
WB - Western Blot
Lasagna - Corning dish for cell culture
Cell Culture
Day1: Seed 8e6 cells per lasagna
Note
1 lasagna is normally enough for one replicate of pull-down experiment
Day 2: Induce expression of BirA construct by adding Tetracycline 1:1000
Note
Tetracycline stock: 1 mg/mL in Ethanol
Day 3: Add 50 micromolar (µM) of Biotin (in water)
Day 4: Collect cells
Collection of cells
Wash cells 2x with RT PBS + Ca/Mg
Add 4 mL Trypsin per lasagna and incubate 00:07:00 at Room temperature or till cells start to detach
7m
Add 2x 20 mL of DMEM with 5% FBS and pipet cells carefully off
Spin 00:05:00 , 500 x g, 4°C
5m
Remove supernatant and resuspend cells in 10 mL cold PBS
Count the cells and prepare tubes with 2e7 containing cells each
Spin 00:05:00 , 500 x g, 4°C
5m
Remove supernatant and snap freeze samples in liquid nitrogen. Store at -20 °C
Preparation of acetylated beads
2m
Equilibrate beads (Streptavidin Sepharose) in PBS by taking 500 µL slurry beads in a new tube
Note
Never vortex or shake the beads too badly to avoid destroying them
Spin 00:01:00 2000 x g
1m
Remove supernatant carefully and add 1 mL PBS
Spin 00:01:00 2000 x g and repeat wash/spin 2x and remove supernatant to have ~400µl left
1m
Add freshly made Sulfo-NHS-acetate to a final concentration of 10mM (400µl beads + 360µl PBS + 40 µL 200 millimolar (mM) Sulfo-NHS-Acetate in PBS)
Incubate 00:30:00 at RT
30m
Add again freshly made (again) Sulfo-NHS-acetate and incubate 00:30:00 at RT
30m
Quench with 1:10 of 1 Molarity (M) Tris 7.5
Wash beads extensively with 1 mL of PBS (at least 2 washes) and spin at 2000 x g for 00:01:00
1m
Remove supernatant and resuspend them in PBS (final vol. 500µl)
Take out the amount that you need
To store the left-over beads, spin them, remove supernatant, wash 2x with 20% ethanol and add 20% ethanol (storage solution) up to the right volume
Pull-down of biotinylated proteins
Take one tube per BirA cell line and thaw on ice
Note
You should have 2e7 cells in total per tube
Prepare lysis buffer
Note
You need more than just the volume for the samples. You need it later for wash steps, too
| A | B | C | D | E | |
| Component | Existing conc. | Final Conc. | Vol. needed in µl | Manufacturer | |
| Tris pH 7.5 | 500mM | 50mM | 950 | Carl Roth, 4855.2 | |
| NaCl | 5M | 150mM | 285 | Carl Roth, 3957.1 | |
| EDTA | 500mM | 1mM | 19 | Carl Roth, 8043.2 | |
| EGTA | 100mM | 1mM | 95 | Carl Roth, 3054.1 | |
| Triton-X100 | 1% | 95 | Carl Roth, 3051.3 | ||
| Aprotinin | 10mg/ml Stock | 1/1000 | 9.5 | Carl Roth A162.3 | |
| Leupeptin | 5ml/ml Stock | 1/1000 | 9.5 | Carl Roth, CN33.2 | |
| Turbonuclease | 250U | 1 | MoBiTec GmbH, GE-NUC10700-01 | ||
| SDS | 20% | 0.10% | 47.5 | ||
| Water | 7988.5 |
In the table above are the amounts needed for one sample
Resuspend the pellet in 4.75 mL of Lysis buffer in a 15ml falcon tube
Lyse the cells by rotating the falcon tube for 01:00:00 , 15 rpm, 4°C
1h
Split each sample into 2x2ml tubes
Sonicate the sample 10x 30sec on/off with the Bioruptor at 4 °C
Note
No visible aggregates should be there, otherwise sonicate more
Spin the samples 00:30:00 , 17000 x g, 4°C -> keep supernatent for further analysis
30m
Equilibrate the (already prepared) beads in the meantime
Rebuffer the beads in PBS (wash 3x with PBS, spin at 2000 x g for00:05:00 , remove supernatant)
5m
Add 80 µL of slurry beads to 1 mL Lysis buffer
Incubate them for 00:30:00 15 rpm, 4°C
30m
Spin 00:05:00 , 200 x g 4 °C
5m
Remove as much of the Lysis buffer as possible
Continue with pull-down of biotinylated proteins
3h 5m
Transfer the supernatants per sample in a fresh 15ml falcon tube
Add equilibrated beads, use some additional supernatant of the sample to transfer all the beads
Incubate 03:00:00 15 rpm, 4°C
3h
Spin 00:05:00 , 2000 x g 4 °C
5m
Remove 4.5ml of supernatant -> keep 50µl of each sample for WB (flow-through)
Transfer the rest to one of the “empty” columns (Pierce Spin Columns Snap Cap)
Rinse falcon tube with500 µL of Lysis buffer to transfer all the beads
Wash beads with 800 µL of Lysis buffer
Wash beads 5x with 600 µL 50 millimolar (mM) AmBic, 8.3
Note
use freshly prepared AmBic
Close column with a plug on the bottom
Transfer the beads to a 2ml tube and spin 2000 x g , 00:05:00 ,4 °C
5m
Remove supernatant to have 200µl left (~700-720µl)
Add 1 µL of LysC (1 µg/µL )
Digest16:00:00 , 37 °C , 500 rpm
16h
Spin 2000 x g , 00:05:00 , Room temperature
5m
Transfer everything to an “empty” column (Pierce Spin Columns Snap Cap)
Elute digested peptides
Add 2x 150 µL of 50 millimolar (mM) AmBic and pipet 5x up and down
Collect all the elutions of digested peptides together in a low-binding tube
Add 2x 150 µL of 80% ACN + 20% TFA and pipet 5x up and down (fast)
Collect all the elutions of digested peptides together in a low-binding tube
For AmBic elutions
3h
Add 0.5 µL of Trypsin (1 µg/µL )
Digest 03:00:00 , 500 rpm, 37°C
3h
Speed-vac the samples
Resuspend in 200 µL OASIS Buffer A, sonicate 00:01:30
1m 30s
Check pH, it should be <3, if not acidify with 10% TFA
For ACN elutions
3h
Speed-vac the samples to near dryness (~50µl)
Add 50 µL of 200 millimolar (mM) HEPES 8.0
Note
Check the pH – it needs to be pH 6-8 for Trypsin to work. If not re-buffer with sodium hydroxide (for example add 3µl of 1N NaOH).
Add 0.5 µL of Trypsin (1 µg/µL )
Digest03:00:00 , 500 rpm, 37°C
3h
Acidify with 10 µL of 10% TFA (check the pH afterwards).
Clean up, for both AmiBic and ACN elutions
3h
Perform MACRO-SPIN clean-up (capacity 30-300µg) (spin always 1000 x g )
Equilibrate column with 500 µL of 100% MetOH. Spin 00:01:00 .
1m
Wash 2x300 µL 5% ACN, 0.1% Formic Acid. Spin 00:01:00 .
1m
Load sample 2x(50-450µl). Spin 00:01:00 .
1m
Wash 4x300 µL 5% ACN, 0.1% Formic Acid. Spin 00:01:00 .
1m
Elute 2x250 µL 50% ACN, 0.1% Formic Acid. Spin 00:01:00 .
1m
SpeedVac to dryness.
Resuspend in 15 µL of MS Buffer A
For AmBic elutions take 10 µL of the sample into a MS vial and inject 5 µL for DIA analysis.
For ACN elutions take 10 µL of the sample into a MS vial and inject 3 µL for DIA analysis.
Note
Please note that the ACN elutions will still contain some background peptides from Streptavidin, thus run them AFTER the AmBic elutions and check for carry over after your runs!!
Store the remaining @ -20°C.
Protocol references
Branon,T. C., Bosch, J. A., Sanchez, A. D., Udeshi, N. D., Svinkina, T., Carr, S. A.,
Feldman, J. L., Perrimon, N., & Ting, A. Y. (2018). Efficient proximity
labeling in living cells and organisms with TurboID. Nature biotechnology,
36(9), 880–887. https://doi.org/10.1038/nbt.4201
May,D. G., & Roux, K. J. (2019). BioID: A Method to Generate a History of
Protein Associations. Methods in molecular biology (Clifton, N.J.), 2008,
83–95. https://doi.org/10.1007/978-1-4939-9537-0_7
Roux,K. J., Kim, D. I., Burke, B., & May, D. G. (2018). BioID: A Screen for
Protein-Protein Interactions. Current protocols in protein science, 91, 19.23.1–19.23.15.
https://doi.org/10.1002/cpps.51
Bartolome, A.; Heiby, J. C.; Fraia, D. D.; Heinze, I.; Knaudt, H.; Späth, E.; Omrani, O.; Minetti, A.; Hofmann, M.; Kirkpatrick, J. M.; Dau, T.; Ori, A. ProteasomeID: Quantitative Mapping of Proteasome Interactomes and Substrates for in Vitro and in Vivo Studies, 2024. https://doi.org/10.7554/elife.93256.1.256.1.
