May 30, 2025
Version 2
A guide to long-range PCR for Nanopore sequencing V.2
- Ricardo De Paoli-Iseppi1,
- Mike Clark1
- 1University of Melbourne

Protocol Citation: Ricardo De Paoli-Iseppi, Mike Clark 2025. A guide to long-range PCR for Nanopore sequencing . protocols.io https://dx.doi.org/10.17504/protocols.io.n2bvj9rdxlk5/v2Version created by Ricardo RDP De Paoli-Iseppi
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working.
Created: May 27, 2025
Last Modified: May 30, 2025
Protocol Integer ID: 219044
Keywords: PCR, Nanopore, long-range, long-read, cDNA, RNA, isoform, range pcr for subsequent nanopore, range pcr for nanopore, subsequent nanopore, multiplexed nanopore, nanopore, efficient sequencing of target, ensuring efficient sequencing, sequencing library, many alternative rna isoform, read amplicon, target gene, rna, amplicon, gene of interest, range pcr, specific to transcript, entire annotated cd, gene
Funders Acknowledgements:
NARSAD Young Investigator Grant
Grant ID: 27184
Australian National Health and Medical Research Council Investigator Grant
Grant ID: GNT1196841
Abstract
This protocol details how to design primers for a target gene and carry-out long-range PCR for subsequent Nanopore long-read amplicon sequencing (LSK114). It outlines the resources, tools and settings we use to amplify the entire annotated CDS of a gene, capturing as many alternative RNA isoforms as possible, and make these ready for a multiplexed Nanopore sequencing library. This protocol should reliably result in amplicons in the 1 - 11 kb range. Amplicons resulting from this protocol should be specific to transcripts of the gene of interest with minimal or no off-targets ensuring efficient sequencing of targets.
Image Attribution
Created in BioRender. De Paoli-Iseppi, R. (2024) https://BioRender.com/r06g873
Guidelines
RNA quality
- For best long-range PCR results for RNA isoform profiling the RNA quality (RIN) should be as high as possible (>7), although we have found RIN >6 works for many genes. The longer the isoforms of interest and thus the amplification length, the more important high RNA quality is.
Capturing RNA isoforms - how many primers do I need?
To ensure capture of known and unannotated RNA isoforms, you may need to design multiple different forward/reverse primers to cover alternate start/end sites. UCSC-GB tracks that can support primer design and selection include:
- Comprehensive Gene Annotation Set from GENCODE: All, non-coding, splice variants.
- Human mRNAs from GenBank.
- NCBI RefSeq genes.
- FANTOM5: CAGE reads.
- Human ESTs.
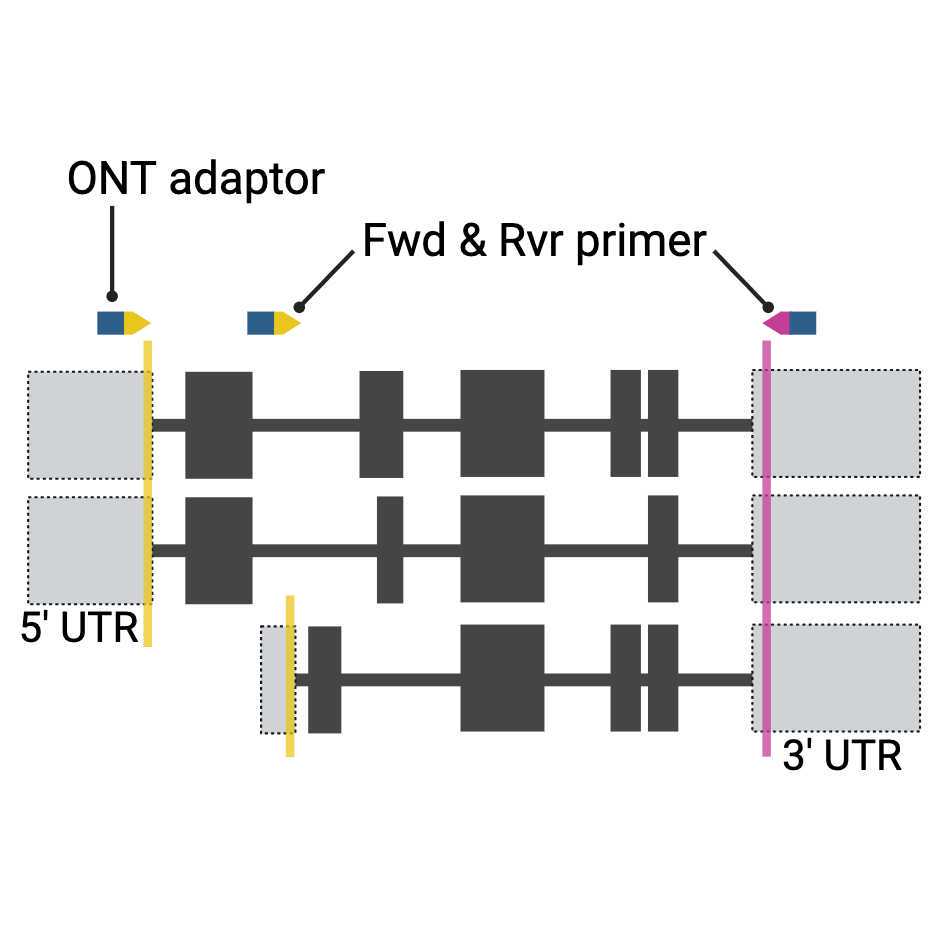
Using the above tracks can provide information and evidence to support design and placement of primers. For example, CAGE tags help identify bona fide transcriptional initiation sites. We suggest placing primers in the 5' and 3' UTR regions close to the ATG and STOP to minimise UTR sequence differences. If these areas are unsuitable for primers the first and last exon can also be used, however isoforms that use alternative exons will be missed. Avoid designing primers across exon-exon boundaries/junctions, as alternative use of either exon may result in amplification failure of specific RNA isoforms. Additionally primers that cross exon-exon boundaries may be able to prime and amplify solely from the 3' exon. This means you don't actually know what 5' exon was present and potentially what isoform was detected.
Note: it is a good idea to test primer pairs individually first to check amplification. E.g. individually test each forward primer with a single reverse: F1-R1 and F2-R1, prior to combing: F1 & F2 with R1.
Taq polymerase & PCR optimisation
There are several DNA polymerases available that are capable of amplifying long products. In some cases it is important to try a different polymerase if one fails to amplify the expected product. Selection of a suitable polymerase can depend on several factors including reaction speed (min/kb), cost, amplicon size and fidelity. In most cases we suggest using test RNA/cDNA first to optimise PCR conditions, considering overall cycles, annealing temperature (test gradient) and 2- or 3-step protocols depending on primer lengths.
Generally, PCR cycles should be kept to a minimum to reduce PCR bias and artifacts, however there needs to be enough PCR product post bead clean-up to ensure reliable ONT barcoding. We commonly barcode 1 ng of PCR product, so the yield required is low, helping keep PCR cycles down.
Materials
We used the following reagents and equipment for Nanopore (ONT) long-range PCR, library preparation and sequencing.
Order primers with ONT Universal Primers (UP) attached:
- Scale (μmole): .025
- Purification: desalt
- Format: liquid/dry – as preferred.
- Concentration if in liquid: 100 micromolar (µM)
- Prepare dilutions for use at 10 micromolar (µM) , (e.g. 90 µL NFW + 10 µL primer stock ).
Options tested for long-range PCR taq polymerase
LongAmp Taq 2X Master Mix - 100 rxnsNew England BiolabsCatalog #M0287S
PrimeSTAR GXL DNA PolymeraseCatalog #R050A
Platinum™ SuperFi II PCR Master MixInvitrogen - Thermo FisherCatalog #12368050
Gel electrophoresis
GeneRuler 1 kb DNA Ladder, ready-to-useThermo FisherCatalog #SM0314
GelGreen™ Nucleic Acid Stain Gel Stain, 10,000X in WaterGold BiotechnologyCatalog #G-745
Library prep and sequencing
Agencourt AMPure XPBeckman CoulterCatalog #A63880
Equipment
MinION
NAME
Sequencer
TYPE
Oxford Nanopore Technologies
BRAND
MinION 1B / MinION 1C
SKU
Equipment
Flongle Flow Cell (R10.4.1)
NAME
Flow cell
TYPE
Oxford Nanopore Technologies
BRAND
FLO-FLG114
SKU
LINK
Equipment
MinION/GridION Flow Cell (R10.4.1)
NAME
Flow cell
TYPE
Oxford Nanopore Technologies
BRAND
FLO-MIN114
SKU
LINK
Protocol materials
RNeasy Lipid Tissue Mini Kit (50)QiagenCatalog #74804
Maxima H Minus Reverse TranscriptaseThermo Fisher ScientificCatalog ##EP0741
GeneRuler 1 kb DNA Ladder, ready-to-useThermo FisherCatalog #SM0314
GelGreen™ Nucleic Acid Stain Gel Stain, 10,000X in WaterGold BiotechnologyCatalog #G-745
Agencourt AMPure XPBeckman CoulterCatalog #A63880
Platinum™ SuperFi II PCR Master MixInvitrogen - Thermo FisherCatalog #12368050
LongAmp Taq 2X Master Mix - 100 rxnsNew England BiolabsCatalog #M0287S
PrimeSTAR GXL DNA PolymeraseCatalog #R050A
Safety warnings
All steps should be carried out by trained personnel in a certified PC2 laboratory and follow established SOPs, RAs and have SDSs available for all reagents used.
Ethics statement
The Human Research Ethics Committee of the University of Melbourne gave ethical approval for this work: #12457 and #28304.
Before start
- QC total RNA before cDNA conversion. RNA quality/integrity should be as high as possible (RIN > 7).
- Read Guidelines for additional primer and PCR design tips.
Retrieve CDS of target isoform - UCSC Genome Browser
Search for a gene of interest in UCSC Genome Browser1 and select the main isoform – usually
highlighted as indicated below:
UCSC Genome Browser screenshot of GRIA1 highlighting the main isoform.
- Details on position, size, coding exon count and strand can be found here.
- Further details including Biotype, Transcript Support Level (TSL) and APPRIS classification can be found under the GENCODE Transcript Annotation.
GRIA1 main isoform details.
GRIA1 main isoform additional Transcript Annotation details.
Select ‘Genomic Sequence’ and use the settings shown below to retrieve the sequence including 5’ and 3’ UTR regions. This sequence can then be copied into Primer3Plus2 and used for primer pair selection.
Settings for main isoform CDS retrieval.
Design primers using Primer3Plus
In Primer3Plus2 (v3.3.0, server: default), copy target sequence into the space and mark the coding region (CAPITALS) with [] to target this region. Press the 'Regions from Seq.' button to apply.
- Primers should be designed in the 5’ and 3’ UTRs (lower case) and close to start and stop codons where possible.
- Exclude UTR areas far from start and stop codons with <> in the sequence.
- Avoid repetitive regions.
- Avoid designing primers across exon-exon boundaries.
General Settings
- Ensure Product Size Ranges includes long PCR products e.g. 1000-15000.
- Primer size: min:18, opt:20, max:27.
- Primer Tm: min:57, opt:60, max:63. (Aim for a max of 2°C difference).
- Primer GC%: min:20, max:80.
- Turn on Mispriming/Repeat Library option to appropriate species (e.g. HUMAN).
Advanced Settings
- Max Poly-X: 3 – 4.
- CG Clamp: 1 – 2. If possible, try primer selection with this set to 2 first and reduce if no primers are found or adjust window selection.
Example settings for Primer3Plus.
Check primers for specificity & off-target amplification
Check that your designed primers are unique by using UCSC tool BLAT3. “BLAT on DNA is designed to quickly find sequences of 95% and greater similarity of length 25 bases or more.”
Copy and paste or upload your primer sequences (min: 20 bp) without the universal primer and click submit e.g.:
>GRIA1_F1
TATGATTGGACCTGGGCTTC
>GRIA1_F2
TGTGCTGCAGTACCCATCTC
BLAT search results for GRIA1 indicating match score, identity, chromosome, strand and start/end positions.
A custom ‘track’ can be made so that you can view primer position in UCSC.
- Click “build a custom track with these results”.
- Click ”Track controls” > Submit and navigate to gene of interest on UCSC GB.
Custom track showing multiple GRIA1 forward primers designed using this protocol.
Perform PCR using UCSC tool: In-silico PCR4.
“In-silico PCR searches a sequence database with a pair of PCR primers.”
- Max Product Size: 10000
- Target: GENCODE Genes
- Copy your forward and reverse primer pair (without universal primers) into the spaces > Submit.
- This will return exact matched ‘amplified’ sequences, check the ENST number against those in UCSC-GB to see which transcripts you aim to capture.
To design additional primers
ONT Universal Primers (UP)
When ordering primers for Nanopore long-read sequencing, add universal sequences for tailing PCR Primers as below. Testing without universal primers first may be necessary as they can reduce efficiency.
The first round of PCR amplification requires tailed primers to be used which carry these sequences:
- 5’ TTTCTGTTGGTGCTGATATTGC-[project-specific forward primer sequence] 3’
- 5’ ACTTGCCTGTCGCTCTATCTTC-[project-specific reverse primer sequence] 3’
Standard example - GRIA1 (2.7 kb)
Using the steps above, we designed primers for Glutamate Ionotropic Receptor AMPA Type Subunit 1 (GRIA1) encompassing a 2787 bp coding region and two supported start sites.
Primers
>GRIA1_F1
TATGATTGGACCTGGGCTTC
20 bp, 59.9 °C , GC:50%
>GRIA1_F2
TGTGCTGCAGTACCCATCTC
20 bp, 59.9 °C , GC:55%
>GRIA1_R1
AAGGGGTCTCCATCTGCTC
19 bp, 59.2 °C , GC:57.9%
Isolate high-quality total RNA from sample following manufacturers' instructions.
Safety information
QIAzol (and equivalents) are toxic. Personnel should be trained in the SOP and appropriate PPE (gloves, eye protection) must be worn when handling this chemical.
RNeasy Lipid Tissue Mini Kit (50)QiagenCatalog #74804
Generate cDNA using 1 µg total RNA and follow manufacturers' instructions.Maxima H Minus Reverse TranscriptaseThermo Fisher ScientificCatalog ##EP0741
Optional: Use test or 'non-critical' samples/cDNA to test the below PCR conditions including anneal temperature, extension and cycles. Cycles should be kept to a minimum to reduce PCR bias.
Prepare 1st round 25 µL PCR reaction for GRIA1 using a standard 3-step PCR approach - 25x cycles with tailed (ONT - UP) gene specific primers. Annealing/extension 60 °C for 00:03:00 .
LongAmp Taq 2X Master Mix - 100 rxnsNew England BiolabsCatalog #M0287S
Check PCR amplicon e.g. gel electrophoresis/TapeStation. Gel: 1.5%, 90V, 01:10:00 .
Example 1st round amplification of GRIA1 in post-mortem human brain tissues using multiple forward (F1, F2) primers. These amplicons are ready for clean-up and barcoding for ONT library preparation. No template control (NTC), Brodmans area (BA), caudate (Caud), cerebellum (Cbm), temporal cortex (Tcx).
GeneRuler 1 kb DNA Ladder, ready-to-useThermo FisherCatalog #SM0314
GelGreen™ Nucleic Acid Stain Gel Stain, 10,000X in WaterGold BiotechnologyCatalog #G-745
Clean-up 1st round PCR: add 0.5X 12.5 µL AMPure XP beads to PCR, follow clean-up procedure and elute in 10 µL NFW.
Agencourt AMPure XPBeckman CoulterCatalog #A63880
Cleaned-up amplicon(s) may now be used for ONT barcoding and multiplexed long-read sequencing following LSK114 library preparation.
Technical example (long) - CSMD1 (10.8 kb)
4h
The schizophrenia risk gene CUB and sushi multiple domains 1 (CSMD1) has the longest CDS we amplified at ~10,838 nt, encompassing 70 coding exons5,6.
Primers
>CSMD1_F1
CCCTCGGGTGATTATTTGG
19 bp, 60.1 °C , GC:52.6%
>CSMD1_R1
CACTGCTTGTCCATCAGAGG
20 bp, 59.4 °C , GC:55%
1h 21m
Prepare 1st round 20 µL PCR reaction for CSMD1 using a 2-step PCR approach - 5x cycles with no ONT Universal Primers (UP). Annealing/extension 72 °C for 00:05:30 .
Platinum™ SuperFi II PCR Master MixInvitrogen - Thermo FisherCatalog #12368050
Clean-up 1st round PCR: add 0.45X 9 µL AMPure XP beads to PCR, follow clean-up procedure and elute in 8 µL NFW.
Agencourt AMPure XPBeckman CoulterCatalog #A63880
Prepare 2nd round 20 µL PCR reaction for CSMD1 (2-step PCR)- 20x cycles with ONT Universal Primers attached. Annealing/extension 72 °C for 00:05:30 .
Platinum™ SuperFi II PCR Master MixInvitrogen - Thermo FisherCatalog #12368050
Check PCR amplicon e.g. gel electrophoresis/TapeStation. Gel: 1.5%, 90V, 01:10:00 .
Primer optimisation result for CSMD1 amplicon. Red box indicates optimal 2-step PCR protocol to result in product incorporated with ONT universal primer tags for subsequent barcoding and library preparation.
GeneRuler 1 kb DNA Ladder, ready-to-useThermo FisherCatalog #SM0314
GelGreen™ Nucleic Acid Stain Gel Stain, 10,000X in WaterGold BiotechnologyCatalog #G-745
Cleaned-up amplicon(s) may now be used for ONT barcoding and multiplexed long-read sequencing following LSK114 library preparation.
Protocol references
1. G. Perez, G. P. Barber, A. Benet-Pages, J. Casper, H. Clawson, M. Diekhans, et al. The UCSC Genome Browser database: 2025 update. Nucleic Acids Res. 2024. Accession Number: 39460617 DOI: 10.1093/nar/gkae974
2. A. Untergasser, H. Nijveen, X. Rao, T. Bisseling, R. Geurts, and J. A. M. Leunissen: Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Research. 2007, 35: W71-W74; doi:10.1093/nar/gkm306
4. A. S. Hinrichs, D. Karolchik, R. Baertsch, G. P. Barber, G. Bejerano, H. Clawson, et al. The UCSC Genome Browser Database: update 2006. Nucleic Acids Research 2006 Vol. 34 Issue suppl_1 Pages D590-D598. DOI: 10.1093/nar/gkj144. https://doi.org/10.1093/nar/gkj144
5. M. B. Clark, T. Wrzesinski, A. B. Garcia, N. A. L. Hall, J. E. Kleinman, T. Hyde, et al. Long-read sequencing reveals the complex splicing profile of the psychiatric risk gene CACNA1C in human brain. Molecular Psychiatry. 2020. Vol. 25 Issue 1 Pages 37-47. DOI: 10.1038/s41380-019-0583-1.
6. R. D. Paoli-Iseppi, S. Joshi, J. Gleeson, Y. D. Joseph Prawer, Y. You, R. Agarwal, et al. Long-read sequencing reveals the RNA isoform repertoire of neuropsychiatric risk genes in human brain. medRxiv. 2024. DOI: 10.1101/2024.02.22.24303189.
Useful links
Acknowledgements
The authors would like to thank the current and previous members of the Clark Lab for their assistance in testing and optimising long-range PCRs.
