Mar 23, 2026
A Centralized Pathology Protocol for Global Cancer Genomics: The Mutographs Project Model
- Behnoush Abedi-Ardekani1,
- Arash Nikmaneshi2,
- Masoud Sotoudeh2,
- Valerie Gaborieau1,
- Farid Azmoudeh-Ardalan3,
- Hiva Saffar4,
- Stefano Serrà5,
- Christine Carreira6,
- Priscilla Chopard1,
- Sophie Ferlicot7,
- Jean-Yves Scoazec8,
- Christophe Lalemand9,
- Elodie Colney9,
- Sophie Guillot9,
- Fatemeh Ghasemi-Kebria10,
- Ana Carolina de Carvalho1,
- Aida Ferreiro-Iglesias1,
- Sandra Perdomo1,
- Paul Brennan1,
- Manuel Salto-Tellez11,12,
- Gholamreza Roshandel13
- 1Genomic Epidemiology Branch, International Agency for Research on Cancer (IARC/WHO), Lyon, France;
- 2Specialized Gastrointestinal and Hepatobiliary specialized center, Tehran, Iran;
- 3Liver Transplantation Research Center, Imam Khomeini Hospital Complex, Tehran University of Medical Sciences, Tehran, Iran;
- 4Department of Pathology, Shariati Hospital, Tehran University of Medical Sciences, Tehran, Iran;
- 5University Health Network, Toronto, Ontario, Canada;
- 6Evidence Synthesis and Classification Branch, International Agency for Research on Cancer (IARC/WHO), Lyon, France;
- 7Service d’Anatomie Pathologique, Assistance Publique-Hôpitaux de Paris, Université Paris-Saclay, Le Kremlin-Bicêtre, France;
- 8Université Paris Saclay, Gustave Roussy, Department of Pathology, Villejuif, France;
- 9Laboratory support, biobanking, and services, International Agency for Research on Cancer (IARC/WHO), Lyon, France;
- 10Golestan Research Center of Gastroenterology and Hepatology, Jorjani Clinical Sciences Research Institute, Golestan University of Medical Sciences (GOUMS), Gorgan, Iran;
- 11Integrated Pathology Unit, The Institute of Cancer Research, London, UK;
- 12Precision Medicine Center, Queen’s University, Belfast, Belfast, UK.;
- 13Golestan Research Center of Gastroenterology and Hepatology, Jorjani Clinical Sciences Research Institute Golestan University of Medical Sciences (GOUMS), Gorgan, Iran

Protocol Citation: Behnoush Abedi-Ardekani, Arash Nikmaneshi, Masoud Sotoudeh, Valerie Gaborieau, Farid Azmoudeh-Ardalan, Hiva Saffar, Stefano Serrà, Christine Carreira, Priscilla Chopard, Sophie Ferlicot, Jean-Yves Scoazec, Christophe Lalemand, Elodie Colney, Sophie Guillot, Fatemeh Ghasemi-Kebria, Ana Carolina de Carvalho, Aida Ferreiro-Iglesias, Sandra Perdomo, Paul Brennan, Manuel Salto-Tellez, Gholamreza Roshandel 2026. A Centralized Pathology Protocol for Global Cancer Genomics: The Mutographs Project Model. protocols.io https://dx.doi.org/10.17504/protocols.io.3byl4p6rzlo5/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: March 20, 2026
Last Modified: March 23, 2026
Protocol Integer ID: 313665
Keywords: centralized pathology protocol for global cancer genomic, global cancer genomic, multicancer genomic initiative, centralized pathology protocol, pathology workflow of the mutographs project, cancer research uk, quality tissue processing in international genomics initiative, stepwise pathology quality control, laser capture microdissection for certain tumor type, quality dna for wg, frozen cancer tissues for pathology, international genomics initiative, macrodissection for tumor, whole genome sequencing, blood dna extraction, pathology workflow, centralized pathology, integrated pathology, quality dna, dna extraction, frozen cancer tissue, certain tumor type, carcinoma, pathology, tumor, viable tumor cell, cancer, laser capture microdissection, percentage of viable tumor cell, differences in cancer incidence, cancer incidence, quality tissue processing, molecular pipeline, mutograph
Funders Acknowledgements:
Cancer Grand Challenges, Cancer Research UK
Grant ID: C98/A24032
Disclaimer
Where authors are identified as personnel of the IARC/WHO, the authors alone are
responsible for the views expressed in this article and they do not necessarily
represent the decisions, policy, or views of the IARC/WHO.
Abstract
We present the pathology workflow of the Mutographs project, a multicancer genomic initiative led by the Sanger Institute and funded by Cancer Research UK, between 2017 and 2023. Mutographs aims to elucidate differences in cancer incidence through whole genome sequencing (WGS) and mutational signature analysis of over 6,000 carcinomas of the esophagus, kidney, colorectum, pancreas, head and neck, and urinary bladder collected from 27 countries. The protocol is designed by the International Agency for Research on Cancer, where centralized pathology and paired tumor-blood DNA extraction was performed. The workflow comprises four main parts. Part 1 defines handling of frozen cancer tissues for pathology and DNA extraction, followed by multi-step pathology quality evaluation and guidance through a structured five-option approach (Part 2), including macrodissection for tumors with percentage of viable tumor cells (PVTC) �3c50% (Part 3) and laser capture microdissection for certain tumor types (Part 4). Rigorous and stepwise pathology quality control led to the overall success rate of 88% in providing high-quality DNA for WGS, while 40.9% of low-quality or non-eligible tissues were excluded. PVTC was successfully enriched in 72% of 1,804 macrodissections performed. Inter-pathologist agreement to report PVTC was substantial (ICC = 0.75, CI: 0.72–0.77). This protocol provides a reproducible fully integrated pathology–molecular pipeline in challenging conditions to ensure high-quality DNA and reliable data interpretation at an unprecedented multinational scale and could be considered as a model for high-quality tissue processing in international genomics initiatives.
Image Attribution
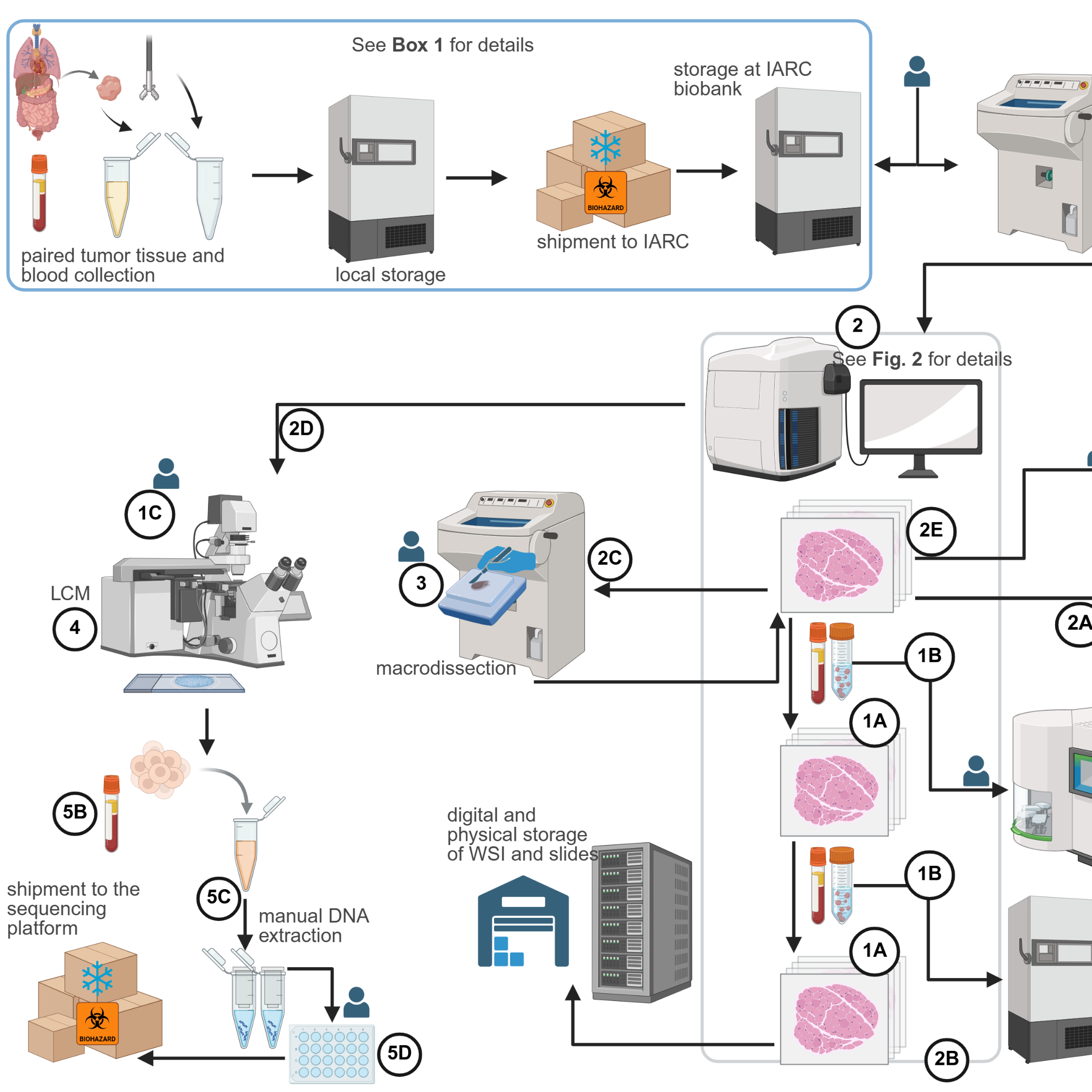
Figure 1: Overview of the protocol workflow
Sample collection and shipment to IARC are
outlined in BOX
1. The protocol comprises five main parts with seven subparts. Part 1
covers embedding frozen tissues in OCT medium. OCT blocks are sectioned and
stained with H&E (1A) for pathology evaluation. Frozen sections are
also prepared for DNA extraction (1B) and LCM (1C), then scanned
to generate WSIs for digital pathology evaluation (Part 2).
Pathologists confirm eligible cancer types, estimate the PVTC, and guide subsequent
steps (2A–2E, Figure 2). Additional WSIs are generated to monitor
PVTC after deep sectioning. Tumors with PVTC <50% undergo macrodissection (Part 3).
Part
4 details LCM, and Part 5 describes automated tumor (5A) and
germline (5B) DNA extraction, manual extraction from LCM-collected cells
(5C), and aliquoting (5D). The workflow is coordinated by
pathologists and a dedicated data manager (“buddy”) who generates weekly case
lists, tracks sample movements, and ensures biobank traceability. All slides
are digitized, annotated, and archived in the IARC image repository, with glass
slides retained for verification. Tissue scrolls for DNA extraction are
collected in duplicate cryotubes: one for extraction and one stored at –80°C as
backup (OCT: Optimal Cutting Temperature; H&E: Hematoxylin & Eosin;
LCM: Laser Capture Microdissection; WSI: Whole Slide Images; PVTC: Percentage
of Viable Tumor cells; IARC: International Agency for Research on Cancer) (Created in https://BioRender.com).
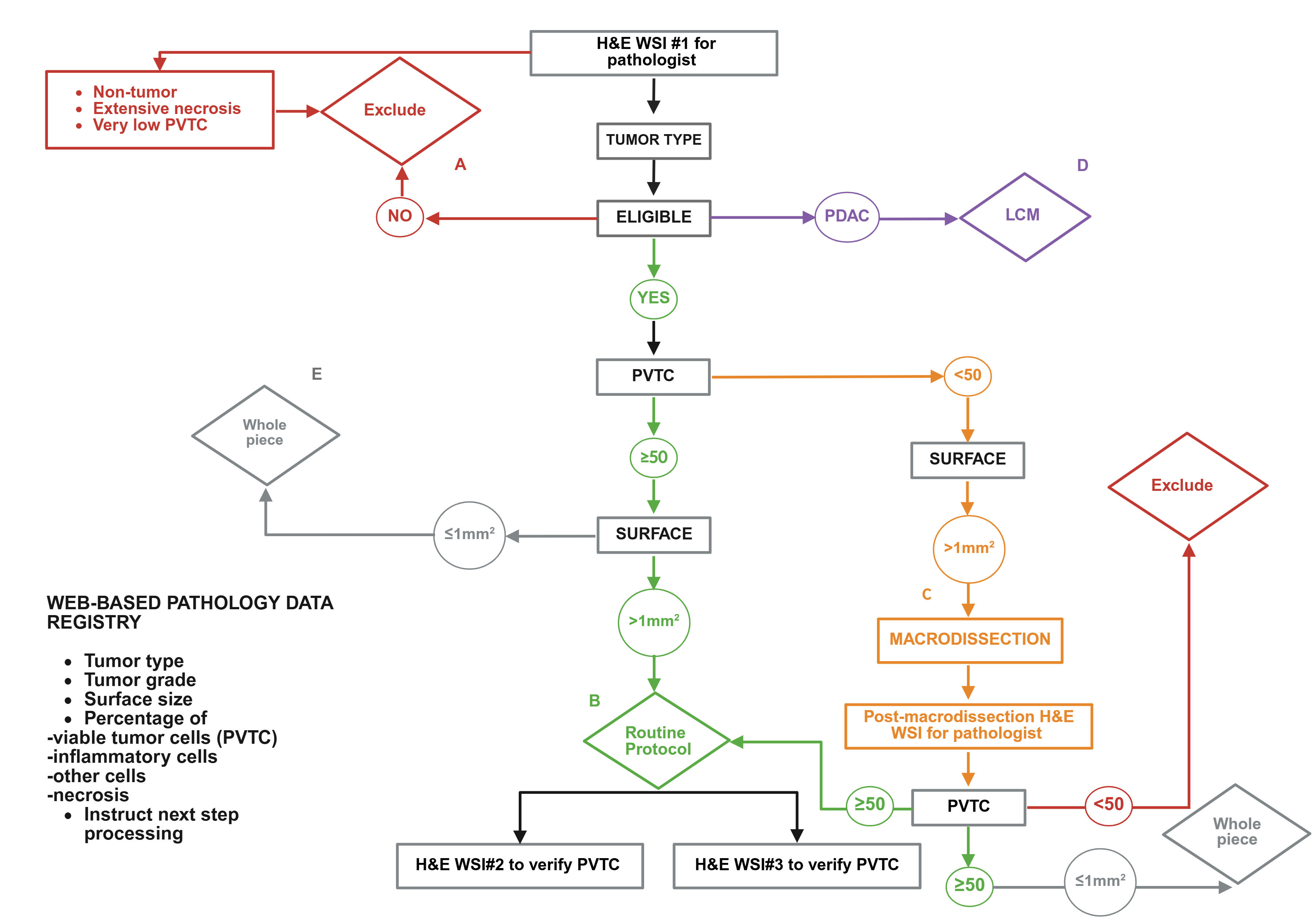
Figure 2: Overview of the multi-step pathology evaluation of
frozen tumor tissues.
The pathology team systematically evaluated
the first H&E-stained WSI. Reports and data registries captured tumor type
and grade, PVTC, inflammatory and stromal cells, and necrosis (total 100%).
Surface area was also measured as a key criterion for decision-making. The next
step was guided by pathology evaluation with five options (A–E). A:
Non-eligible cancers, extensive necrosis, or very low PVTC were excluded.
B:
Eligible cancers with PVTC ≥50% and surface ≥1 mm² entered the routine
protocol, i.e. flank thick sectioning,
collection in two cryotubes, and two additional H&E WSIs to confirm PVTC
maintenance in deep sectioning. C: Tumors with PVTC <50% underwent
macrodissection using pathologist annotated WSIs; post-procedure WSIs verified
success of tumor enrichment and guide next step. D:
PDAC, often with PVTC <50%, were processed by LCM.
E:
Eligible cancers with PVTC ≥50% but surface <1 mm² were processed as whole
piece with no additional pathology work (H&E: Hematoxylin & Eosin;
WSI: Whole Slide Images; PVTC: Percentage of Viable Tumor cells; LCM: Laser
Capture Microdissection; IARC: PDAC: (Pancreatic Ductal Adenocarcinoma) (Created
in https://BioRender.com).
Materials
**Biological materials**
This workflow was successfully applied to the handling of 6,303 frozen tumor tissues and paired blood samples- from Mutographs’ biorepository containing over 11,000 tumor tissues- spanning eight cancer types across seven organs, for WGS. Biological material was collected in accordance with national and international regulations, following signed informed consent from patients enrolled in the project (^^BOX 1^^). Ethical approval was obtained from institutional review boards and the IARC ethics committee (IEC-17-10).
**Reagents**
- OCT (Optimal Cutting Temperature) medium, NEG-50 OCT, Fisher Scientific^®, cat. no. 11912365.
- Ethanol (70%, 95%, 100% v/v), Emprove Expert, Merck^®, cat. no. 1.00983.1000, or VWR International.
- ! CAUTION: Ethanol is flammable. Store in sealed containers away from ignition sources.
- Hematoxylin, Masson’s alum, Merck^®, cat. no. 1.11487.0025.
- Erythrosine B, 0.1% solution, Merck^®, cat. no. 1.15936.0025.
- Xylene, VWR International, cat. no. 28973.363.
- ! CAUTION: Xylene is toxic, flammable, and irritant. Use in a fume hood with adequate ventilation. Store in a flammable cabinet. Wear protective gloves and clothing.
- Paraformaldehyde (PFA), 16% stock and 4% working solutions, Polysciences^®, cat. no. 18814.
- ! CAUTION: PFA is classified as a high-hazard chemical. Handle under a fume hood or in a well-ventilated area. Wear gloves, protective eyewear, and lab coat. Avoid inhalation and skin contact. Store sealed in a cool, dry place.
- Phosphate-buffered saline (PBS), Eurobio^®, cat. no. C3SP80S01-01.
- Mounting medium, PERTEX, Labelling^®, cat. no. PERTEX500.
- DNA Away, Dutscher, cat. no. 038188.
- RNase Away, M-Thermo, Fisher Scientific, cat. no. 11580095.
- Cresyl violet acetate (powder), Aldrich^®, cat. no. 86098-0.
- Ethanol (50% v/v), Emprove Expert, Merck^®, cat. no. 1.00983.1000, or VWR International.
- DNA Away, Dutscher, cat. no. 038188.
**Equipment**
- Deep freezers, –80 °C and –20 °C, any institutional model.
- Cryostat (Institutional model): Microm HM550, Epedia, Thermo Fisher, cat. no. 956454D, equipped with:
o Multiple-size metal embedding cryomolds, Tissue-Tek cryomolds; Sakura Finetek.
o Multiple-size metal specimen chucks, Thermo Fisher, 30 mm.
o Built-in pre-cooled area (freezing stage).
- Digital slide scanner (Institutional model): Aperio GT450, ref. no. 23GT450; Leica Biosystems), or any equivalent scanner.
- Facilities for digital pathology, including computer and monitors.
- Label printer, e.g., Brady Bench i3300, serial no. PIG332134012022.
o NOTE: Used at IARC to print specimen identification barcodes on StainerBondz™ solvent-resistant labels for glass slides. Any institutional approach for secure identification of products is considered.
- Disposable microtome blades (high-profile and low-profile), Fisher Scientific, cat. no. 1291830.
- Sterile disposable surgical scalpels, cat. no. 3752021.
- Fume hood, e.g., Thermo Scientific Herasafe 2030i, cat. no. 51033051/42952033.
- Forceps, medium and small (fine) size, serrated and flat tips, Fisher Scientific, cat. no. 12-651-915, or equivalent.
- Fine paintbrush, e.g., M-M France, cat. no. F/PINCEAUA12.
- Cryotubes, 1.5–2.0 ml, with secure caps, Safelock tubes, Dutscher, cat. no. 033290.
- Superfrost™ Plus adhesion slides, Fisher Scientific, cat. no. 10149870.
- Polyethylene Naphthalate (PEN)-NF membrane slides, Zeiss SAS, cat. no. 415190-9081-000.
- Cryo-resistant deep freezer storage boxes, 136 x 136 x 50 mm, Grossroen, cat. no. 6.080130.
- Cryo-resistant pen.
- Aluminium foil sheets.
- Kimwipes or lint-free tissue, Paredes, cat. no. 484104.
- Staining station slide racks and staining jars, Myreva station, cat. no. SS30, or equivalent.
- Coplin (glass staining) jars, Fisher Scientific, cat. no. 12697696.
- Glass coverslips, 24 x 50 mm, Roth, cat. no. 1871.2.
- Personal protective equipment (PPE): laboratory coat, gloves, and eye protection, VWR International, or any institutional equivalent such as StarLab France.
**Part 4 (Steps 44 to 60)**
- Laser Capture Microdissection device, e.g., Zeiss PALM MicroBeam IV.
- Adhesive cap opaque 500 µl tube, Zeiss, cat. no. 415190-9201-000.
- Deep freezer, –20 °C.
- Coplin (glass staining) jars, Fisher Scientific, cat. no. 12697696.
- Personal protective equipment (PPE): laboratory coat, gloves, and eye protection, VWR International.
Troubleshooting
Safety warnings
**! CAUTION: All experiments involving human tissues must have ethical approvals in accordance with national, institutional, and international regulations. Informed consent must be obtained from all patients.
**! CAUTION: Unfixed tissues should be considered potentially infectious. They must be handled in a containment level 2 laboratory. All users should be screened for appropriate immunizations according to local guidelines. Use personal protective equipment (PPE), including safety glasses, gloves, and laboratory coat.
**! CAUTION: Ethanol is flammable. Store in sealed containers away from ignition sources.
**! CAUTION: Xylene is toxic, flammable, and irritant. Use in a fume hood with adequate ventilation. Store in a flammable cabinet. Wear protective gloves and clothing.
**! CAUTION: PFA is classified as a high-hazard chemical. Handle under a fume hood or in a well-ventilated area. Wear gloves, protective eyewear, and lab coat. Avoid inhalation and skin contact. Store sealed in a cool, dry place.
**! CAUTION: Be careful when handling cryostat blades. To clean from tissue debris, always use a brush. Wear anti-cutting metal gloves if not experienced. Always lock the handwheel of cryostat if not cutting to prevent accidental movement of the blade.
**! CAUTION: To avoid DNA carryover, clean the cryostat after each use per specimen. Remove tissue debris with a brush and wipe all surfaces with DNase/RNase decontamination solution first followed by 70% ethanol. At the end of daily use, run the chemical decontamination or UN irradiation-based on the cryostat model.
**! CAUTION: Fixation and staining should be done in a hood (laminar flow recommended) in a containment Level 2 lab.
**! CAUTION: Place the OCT block on a solid, even surface; cutting a mispositioned block risks hand injury.
**! CAUTION: Never hold the block with your hand. Use forceps and keep hands away from the cutting area.
**! CAUTION: Discard surgical scalpels in sharps container.
Ethics statement
Ethical approval was obtained from institutional review boards and the IARC ethics committee (IEC-17-10).
PROCEDURE OVERVIEW
Part 1: Embedding of frozen tissues in the Optimal Cutting Temperature (OCT) medium and cryosectioning
Remove the cryotubes containing frozen tissues from deep freezer (– 80° or – 20° C) according to the list provided by the data manager.
In Mutographs, a dedicated data manager co-ordinated the circuit of procedures in
regular manner and as a quality control measurement. Institutional and project-based design might act differently.
Verify the identification number of each cryotube. CRITICAL STEP: Always confirm sample ID. Apply the unique ID to all equipment (glass slides, cryotubes, OCT blocks, etc.) and by-products using your institutional sample tracking system.
CRITICAL STEP: Always carry the cryotubes containing tissues from the biobank to a manipulation area, such as pathology laboratory, in a cryobox and in frozen state, by using dry ice (long distance) or ice cubes (short distance). Be careful to avoid unnecessary heat shock. Transfer the cryotubes from the cryobox into the cryostat chamber, pre-cool to -20 °C chamber with maintained – 20° C temperature. Allow the tubes to equilibrium for 1-2 min.
CRITICAL STEP: Process only 1-2 cryotubes at a time. Keep the remaining tubes in the cryobox on dry ice, with the lid firmly closed, to avoid heat chock. Select
cryomold of appropriate size, slightly larger than the specimen, to allow a surrounding OCT border.
Label the cryomold with the specimen ID using a cryo-pen.
Pour a thin layer of OCT to cover the bottom of the cryomold. Allow it to opacify (not fully solidify) for 10-30 sec, creating a flat, level base for tissue placement.
Using pre-cooled forceps, remove the tissue from the cryotube and place it on the OCT base.
CRITICAL STEP: For RNAlater®-preserved tissues, gently blot with a Kimwipe or other lint-free absorbent tissue until no visible liquid remains. Do not wash with PBS, as this promotes undesirable ice crystal formation. Press the tissue gently to ensure even attachment and check that no air pockets remain.
Add OCT to fully cover the tissue, avoiding bubble formation.
Leave the cryomold in the cryostat chamber undisturbed for 15–30 s to allow OCT infiltration and dissipation of air bubbles.
Transfer the cryomold to the freezing stage and allow the OCT to fully solidify (20–50 sec).
Proceed to sectioning
PAUSE POINT: The OCT blocks can be stored at – 80° C if sectioning will not be performed immediately.
A. Frozen sectioning for pathology evaluation
Apply a small drop of fresh OCT onto the flat surface of the chuck and place the OCT block on it (OCT bedding).
ii. Place the chuck with attached OCT in the freezing stage (pre-cooled area). Allow it to freeze for about 5 min.
iii. Slide a pre-cooled disposable blade into the holder. Adjust the angle and secure it with the locking mechanism. CRITICAL STEP: Use a new blade per tissue block. Process one specimen at a time to prevent contamination.
iv. Select tissue section thickness by ‘+’ and ‘-‘ buttons on the left hand side. Adjust the thickness based on the procedure and institutional design. We usually trim at 20-30 µm, cut at 5 µm for histopathology assessment, at 10 µm for flank sectioning, and at 15 µm for LCM.
Trim the block until a flat tissue surface is exposed. CRITICAL STEP: For small biopsies, slow trimming once tissue contrast appears against OCT to avoid losing tissue.
vi. Mount the section onto a pre-labeled, pre-cooled Superfrost™ Plus slide.
Always, and particularly when treating frozen tissues, use Superfrost™ Plus slides to prevent washed-out sections during staining.
vii. Leave the slide on the slide rack inside the cryostat.
viii. After all slides are prepared, remove the slide rack and allow at room temperature for ≥10 min to air dry before staining.
Ensure complete air drying to prevent folding or cracking during staining.
ix. Fix sections in freshly prepared 4% PFA for 10–15 min.
Briefly rinse slides in PBS; drain excess liquid. CAUTION: Fixation and staining should be done in a hood (laminar flow recommended) in a containment Level 2 lab.
xi. Stain sections sequentially: Hematoxylin (5 min), tap water rinse (5 min), Eosin (2–3 min), ethanol series (70%, 95%, 100%, 1–2 changes each), xylene (2 changes, 5 min each).
We recommend avoid rinsing after eosin to prevent washing out the stain.
xii. Mount and coverslip.
xiii. Proceed to scanning and whole-slide image (WSI) creation for Part 2.
B. Frozen sectioning for LCM
Continue from step viii
ii. Change the section thickness by ‘+’ and ‘-‘ buttons on the left hand side to 15 µm. CRITICAL STEP: Avoid thickness >15 µm; laser cutting is inefficient and may burn tissue. CRITICAL STEP: Ensure flat, wrinkle-free section in the center of the membrane; leave 1.5 cm margin for laser. Discard any folded section.
To avoid DNA contamination, PEN slides must be UV crosslinked for 30 min. In out institute, certified DNase/RNase-free PEN slides were used.
iii. Proceed for LCM, if feasible. PAUSE POINT: Unfixed slides can be stored at –80°C until LCM.
Part 2: Stepwise pathology evaluation
Follow from step xiv.
Open the image viewer. o Note: SlidePath was the image viewer software linked to the Leica SCN400 scanner in the Mutographs project, replacing Aperio Image Scope v12.4.6.7001 after SCN400 being out of support.
Open the pathology data registry program, according to the institutional set up
o Note: We developed a specific pathology data registry for Mutographs. The point is to ensure secure and hramonized pathology data registry, by any approved institutional approach.
Tumor type, PVTC, and surface size are key components to guide the workflow. Guiding criteria are designed based on the aims and objectives of each project.
Report and instruct based on options A to D (Figure 2).
If non-malignant tissue, extensive necrosis, ou non-eligible cancer type instruct option A
ii. If eligible cancer type and PVTC ≥50%, instruct option B
iii. If eligible cancer type, PVTC <50%, enrichment of tumor content is feasible, and surface size >1 mm2, instruct option C
iv. If pancreatic ductal adenocarcinoma (PDAC) with PVTC <50%, instruct option D
If eligible cancer type, PVTC ≥50%, but surface is ≤ 1mm2, instruct option E
Annotate the area of interest if instruct for macrodissecton or LCM.
Part 3: Macrodissection
Follow steps 9 to 11 to set up the cryostat.
Verify the case list prepared by the data manager.
Extract the OCT block from -80° C storage. Check the case ID.
Transfer OCT blocks to the pre-cooled cryostat chamber.
Equilibrate the OCT block in the freezing stage for 10–15 min.
Examine corresponding glass slides and WSI; orient pathologist’s annotations on the OCT block. Keep images and slides visible.
Re-check that the case ID matches all materials.
Apply a new disposable scalpel and gradually cut off unwanted areas using shallow cuts first. CAUTION: Place the OCT block on a solid, even surface; cutting a mispositioned block risks hand injury. CAUTION: Never hold the block with your hand. Use forceps and keep hands away from the cutting area.
Use a new scalpel for each sample to avoid DNA contamination.
Deepen cuts after delineating the area with shallow lines.
Allow the dissected OCT block to slightly thaw, remove, and place on a clean cryomold.
Follow steps to embed and freeze
Identify the post-macro OCT block according to institutional tracking.
Proceed to sectioning, post-macro WSI creation, and thick sectioning for DNA extraction (steps A i to A xiv and B i to v). PAUSE POINT: Post-macro block can be stored at –80°C if not immediately sectioned. CAUTION: Discard surgical scalpels in sharps container.
Part 4: Laser Capture Microdissection
Follow steps 1-4 for reagent set up and 12-13 for equipment set up
Check the slide ID based on the list of cases provided by data manager.
Fix slide in 70% ethanol for 2 min.
Dip the slide for 10 sec in 1% cresyl violet acetate working solution.
Fix in 70% ethanol for 1 min.
Dip in 100% ethanol for 30 sec.
Air-dry at room temperature for 5 min.
Proceed to step 10 of equipment setup.
Load stained slide into the slide holder.
Select observation objective (5X or 10X).
Load collection tube (adhesive cap, 500 µL).
Follow pathologist guidance to annotate areas of interest using the software drawing tool.
Acquire images before and after microdissection.
Start laser beam to cut and catapult material into the cap. CRITICAL STEP: Calibrate laser, align tube, avoid air currents. Check alignment regularly. Scan slide at end to confirm collection. Apply additional laser power if needed.
Remove tube, close, seal, label, and store at –20°C until DNA extraction.
Switch off the machine.
Decontaminate and clean equipment.
Protocol references
1. Díaz-Gay, M. et al. Geographic and age-related variations in mutational processes in colorectal cancer. medRxiv (2025). https://doi.org/10.1101/2025.02.13.25322219
2. Moody, S. et al. Mutational signatures in esophageal squamous cell carcinoma from eight countries with varying incidence. Nat. Genet. 53, 1553-1563 (2021). https://doi.org/10.1038/s41588-021-00928-6
3. Senkin, S. et al. Geographic variation of mutagenic exposures in kidney cancer genomes. Nature 629, 910-918 (2024). https://doi.org/10.1038/s41586-024-07368-2
4. Torrens, S. et al. The complexity of tobacco smoke-induced mutagenesis in head and neck cancer. Nat. Genet. (2025). https://doi.org/10.1038/s41588-025-02134-0
5. Perdomo, S. et al. The Mutographs biorepository: A unique genomic resource to study cancer around the world. Cell Genom 4, 100500 (2024). https://doi.org/10.1016/j.xgen.2024.100500