Jun 26, 2025
Version 1
2025 Oxford Nanopore Technologies (ONT) Sequencing Proficiency Testing Exercise V.1
- Narjol Gonzalez-Escalona1,
- Maria Hoffmann2,
- Jayanthi Gangiredla2,
- james.pettengill P2,
- arthur.pightling P2
- 1FDA;
- 2US FDA
- GenomeTrakrTech. support email: [email protected]

Protocol Citation: Narjol Gonzalez-Escalona, Maria Hoffmann, Jayanthi Gangiredla, james.pettengill P, arthur.pightling P 2025. 2025 Oxford Nanopore Technologies (ONT) Sequencing Proficiency Testing Exercise. protocols.io https://dx.doi.org/10.17504/protocols.io.dm6gpq6oplzp/v1
License: This is an open access protocol distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Protocol status: Working
We use this protocol and it's working
Created: May 09, 2025
Last Modified: June 26, 2025
Protocol Integer ID: 218013
Keywords: ONT, long read sequencing, PT exercise, RBK-114, R10.4.1, proficiency testing isolates in june, sequencing proficiency testing exercise, following proficiency testing isolate, ont proficiency testing exercise, applicable to all lffm laboratory, proficiency testing exercise this sop, lffm laboratory, entire proficiency test, participating laboratory, completion of the entire proficiency test, pt strain, sequencing data, fda in accordance, pt exercise data in subsequent analysis, fda sop, fda hfp, pt exercise data, fda, oxford nanopore technology, ont pt exercise, sequencing record, laboratory
Funders Acknowledgements:
FDA Foods Programme
Abstract
This SOP outlines guidelines on how to process the isolates for the 2025 ONT Proficiency Testing exercise.
- This SOP is applicable to all LFFM laboratory participating in the 2025 ONT Proficiency Testing exercise.
The FDA HFP/DFSG/GDAB will ship the following proficiency testing isolates in June 2025.
| A | B | C | |
| Strain ID | isolate_name_alias | Organism | |
| GTVSS-001 | CFSAN000189; 231742; PTC003; PTD003; CFSAN000189_B; CFSAN137036 | Salmonella enterica serovar Bareilly | |
| GTVSS-011 | CFSAN023464; 869225-3c; CFSAN137046 | Listeria monocytogenes | |
| GTVSS-014 | CFSAN023469; 869226-3c; CFSAN137049 | Listeria monocytogenes | |
| GTVSS-019 | CFSAN044836; UNITO-144; CFSAN137054 | Listeria innocua | |
| GTVSS-020 | CFSAN051458; FDA858783-1-52; FNW19M81; FDA00010253; CFSAN137055 | Escherichia coli O121 | |
| GTVSS-021 | CFSAN068773; PR0330_EX_017; CFSAN137056 | Cronobacter sakazakii | |
| GTVSS-016 | CFSAN030807 | Shigella sonnei | |
| GTVSS-025 | CFSAN084952 | Pseudomonas fluorescens |
Completion of the entire proficiency test entails the following:
- LFFM participating laboratory generates sequencing data (fastq files) using the PT strains provided by FDA in accordance to FDA SOPs.
- Submission of sequencing records to the appropriate shared folder (box.com), see protocol.
- By participating in the 2025 ONT Proficiency Testing exercise, LFFM laboratories provide consent to use the PT exercise data in subsequent analysis and manuscript publications. Participants will be acknowledged for their contribution on any publication that might require processing data from the 2025 ONT PT exercise.
- Complete the participation survey at the end of the PT exercise.
Materials
- Slant Cultures Materials and reagents
Trypticase Soy Agar (TSA) (or equivalent)
Trypticase Soy Broth (TSB) (or equivalent)
Brain Heart Infusion (BHI) Agar
Brain Heart Infusion (BHI) Broth
Disposable inoculating loops (1 µL or 5 µL )
Ethanol 70% [Note: freshly prepared]
Permanent marker
Incubator range 25 °C to 37 °C
2. Recommended equipment for ONT sequencing.
| A | B | C | |
| Equipment | Product Number | Manufacturer | |
| Centrifuge (e.g., Eppendorf max speed 15,000rpm) | 15881635 | Fisher Scientific | |
| GridION sequencer | Oxford Nanopore Technologies (ONT) | ||
| Plate centrifuge | Fisher Scientific | ||
| Thermomixer (e.g., Eppendorf F2.0 Model) | 15356551 | Fisher Scientific | |
| PCR machine (e.g., T100 Thermal Cycler) | Any | Any | |
| Magnetic rack Invitrogen DynaMag -2 Magnet | 10723874 | Fisher Scientific | |
| Optional: Hula mixer | 10548425 | Fisher Scientific | |
| Qubit fluorometer | 16223001 | Fisher Scientific | |
| Vortex (e.g., Fisherbrand mini vortexer) | Any | Any |
3. Consumables for ONT sequencing.
| A | B | C | |
| Consumables | Product Number | Manufacturer | |
| Eppendorf DNA lo bind tubes 1.5ml (250) | EP0030108051-250EA | Sigma Aldrich | |
| Eppendorf DNA lo bind tubes 2ml (250) | EP0030108078 | Sigma Aldrich | |
| 96-well PCR plate semi-skirted (10) | AB0900 | ThermoFisher Scientific | |
| Adhesive PCR Plate Seals (100) | AB0558 | ThermoFisher Scientific | |
| Qubit Assay Tubes (500) | 12037609 | Fisher Scientific | |
| R10.4.1 Flowcell | FLO-MIN114 | ONT |
4. Reagents for ONT sequencing.
| A | B | C | |
| Reagents | Product Number | Manufacturer | |
| Nuclease-Free H2O | W4502-10x50ml | Sigma Aldrich | |
| Qubit 1x dsDNA HS Assay Kit (100) | Q33230 | Fisher Scientific | |
| Rapid Barcoding Kit 24 V14 or Rapid Barcoding Kit 96 V14 | SQK-RBK114.24 or SQK-RBK114.96 | Oxford Nanopore Technologies (ONT) | |
| Ethanol, absolute (e.g., Fisher Bioreagents) | 16606002 | Fisher Scientific | |
| Optional: Bovine Serum Albumin (BSA) (50 mg/ml) | AM2616 | ThermoFisher Scientific | |
| Optional: Wizard HMW DNA Extraction Kit | A2920 | Promega | |
| Optional: Monarch HMW DNA Extraction Kit | T3060S or T3050L | New England Biolabs | |
| Optional: Nanobinds HMW | 102-301-900 | PacBio | |
| Optional: Flow cell wash kit | EXP-WSH004 | Oxford Nanopore Technologies (ONT) |
Protocol materials
Ethanol 70% [Note: freshly prepared]
Safety warnings
Biological Safety Warning: Escherichia/Shigella, Salmonella, and Listeria strains are considered Level 2 biological agents by the U.S. Department of Health and Human Services. Use appropriate precautions when handling the vial or culture. Carry out laboratory work in a biological safety cabinet when applicable to ensure aseptic conditions and personal safety.
Before start
There are four sections in this protocol:
Section 1: Culture preparation of isolates.
Section 2: DNA Extraction
Section 3: Library Preparation and Sequencing
Section 4: Quality control and preliminary data analysis of Sequencing Data
Section 5: Data Transfer
Culture Preparation
2d 12h
Salmonella, Shigella, Cronobacter sakazakii, and Escherichia from slant cultures:
Day 1
Label the agar plate and broth tube (organism name, sample number, date, and initials), and document the isolate number for your records.
Wipe the outside of the slant tube with Ethanol 70% [Note: freshly prepared] (be careful not to wipe the strain label).
Open the slant tube and gently touch a disposable 1-5 µL loop to a small amount of growth.
Streak the TSA agar plate for colony isolation (i.e., Quadrant or T-Streak).
Incubate the inverted TSA plate (agar side up , aerobically) at 37 °C for Overnight or 18:00:00 .
18h
Day 2
Purity Check (Plate): colonies should appear similar in size, shape, and color.
After Purity check, pick an isolated colony and streak it onto a fresh TSA agar plate. Incubate (Aerobic) at 37 °C Overnight or 24:00:00 .
1d
Day 3
Use the growth from this plate to make DNA templates of the PT strains.
Listeria monocytogenes and L. innocua slant cultures:
Day 1
Label the agar plate and broth tube (organism name, sample number, date, and initials), and document the isolate number for your records.
Wipe the outside of the slant tube with Ethanol 70% [Note: freshly prepared] (be careful not to wipe the strain label).
Open the slant tube and gently touch a disposable 1-5 µL loop to a small amount of growth.
Streak the BHI agar plate for colony isolation (i.e., Quadrant or T-Streak).
Incubate the inverted BHI agar plate (agar side up , aerobically) at 37 °C for Overnight or 18:00:00 .
18h
Day 2
Purity Check (Plate): colonies should appear similar in size, shape, and color.
After Purity check, pick an isolated colony and streak it onto a fresh BHI agar plate. Incubate (Aerobic) at 37 °C Overnight or 24:00:00 .
Day 3
Use the growth from this BHI plate to make DNA templates of the Listeria PT strains.
Pseudomonas fluorescens from slant cultures:
Follow the same procedure than for Salmonella.
DNA Extraction
2m 5s
**- Each isolate for the proficiency testing exercise shall be processed as any routine isolate according to laboratory guidelines.
Perform DNA extraction according to lab’s normal workflow described on GenomeTrakr and/or PulseNet SOPs.
(1) Gram negative bacteria (i.e. Escherichia coli, Klebsiella pneumoniae, Pseudomonas aeruginosa, Serratia marcescens, Enterococcus faecalis, Bacillus subtilis)
a. DNA is extracted using the Maxwell RSC cultured cells DNA kit with a Maxwell RSC instrument (Promega, Madison, WI) following the manufacturer’s protocols for Gram-negative bacteria with additional RNase treatment. The protocol for the Maxwell RSC cultured cells DNA kit is in annex 1.
b. DNA concentration is determined by Qubit 4 Fluorometer (Invitrogen, Carlsbad, CA) according to manufacturer’s instructions.
(1) Gram positive bacteria (i.e. Staphylococcus aureus, Staphylococcus epidermidis, Clostridium botulinum, etc)
a. pre-lysis treatment of the cells to rupture the wall (according to your own protocols for Gram positive bacteria)
b. Continue as described for Gram negative bacteria above.
Place the extracted DNA at 4-8 °C °C until needed. This step is required to avoid further shearing from freezing and thawing.
DNA quantification (Qubit HS Assay kit and Qubit fluorometer)
a. Using 1x dsDNA high sensitivity Qubit reagents, aliquot 198 µL per sample and 2 x 190 µL per standard.
b. Add 10 µL of each standard to 190 µL of Qubit reagent.
c. Add 2 µL of DNA extract to 198 µL of Qubit reagent.
d. Vortex for 00:00:05 then incubate at Room temperature for 00:02:00 2min.
e. Read on the Qubit.
f. Expected yield 50-120 ng/µl (optimum input concentration for library preparation is 200 ng total) - Yield will vary by organism >11 ng/µl ng/µl is sufficient for 200 ng in 18 µL barcoding reactions.
2m 5s
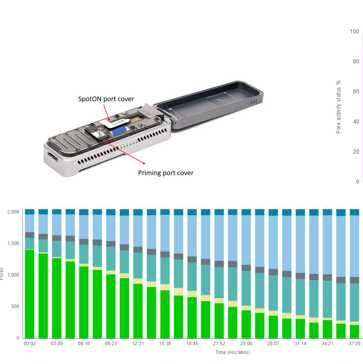
Library Preparation and Sequencing
** The sequencing run for the PT exercise shall be loaded as any routine run following established loading requirements.
** Control reference strains shall be included in the run when there is not enough isolates to fill a sequencing run.
Perform library preparation according to lab’s normal workflow described on FDA Human Foods Program ONT SOPs.
Protocol
CREATED BY
Narjol Gonzalez-Escalona
Sequencing with ONT does not require a sample sheet but you might be to do if desire as below:
| A | B | |
| barcode | alias | |
| barcode13 | sample01 | |
| barcode14 | sample02 | |
| barcode15 | sample03 | |
| barcode16 | sample04 | |
| barcode17 | sample05 | |
| barcode18 | sample06 | |
| barcode19 | sample07 | |
| barcode20 | sample08 | |
| barcode21 | sample09 | |
| barcode22 | sample10 | |
| barcode23 | sample11 | |
| barcode24 | sample12 |
you can use this format to perform analysis using the epi2me lab software from ONT
Strain_ID or Sample_Name: Include in these fields the strain identifiers as it is indicated in Table 1, *do not modify these IDs*. You will also find this identifier in the tube of the slant culture.
| A | B | C | |
| Strain ID | isolate_name_alias | Organism | |
| GTVSS-001 | CFSAN000189; 231742; PTC003; PTD003; CFSAN000189_B; CFSAN137036 | Salmonella enterica serovar Bareilly | |
| GTVSS-011 | CFSAN023464; 869225-3c; CFSAN137046 | Listeria monocytogenes | |
| GTVSS-014 | CFSAN023469; 869226-3c; CFSAN137049 | Listeria monocytogenes | |
| GTVSS-019 | CFSAN044836; UNITO-144; CFSAN137054 | Listeria innocua | |
| GTVSS-020 | CFSAN051458; FDA858783-1-52; FNW19M81; FDA00010253; CFSAN137055 | Escherichia coli | |
| GTVSS-021 | CFSAN068773; PR0330_EX_017; CFSAN137056 | Cronobacter sakazakii | |
| GTVSS-016 | CFSAN030807 | Shigella sonnei | |
| GTVSS-025 | CFSAN084952 | Pseudomonas fluorescens |
Quality of Sequencing Data
Perform quality control and preliminary data analysis of Sequencing Data according to lab’s normal workflow described below:
Protocol
CREATED BY
Narjol Gonzalez-Escalona
Data Transfer
Data transfer
After checking the quality of your records, transfer the data with acceptable quality according to GenomeTrakr guidelines via the HHS Box platform.
Sharing a sequence data via the HHS Box platform
A folder will be generated within the HHS Box platform to which the uploader(s) will be invited to the platform where they can then upload the fastqs sequence data.
Below is an example of the Box interface through which participants will upload sequence data. Participants can either drag and drop the required files or folder with the files or do so via the Upload button.
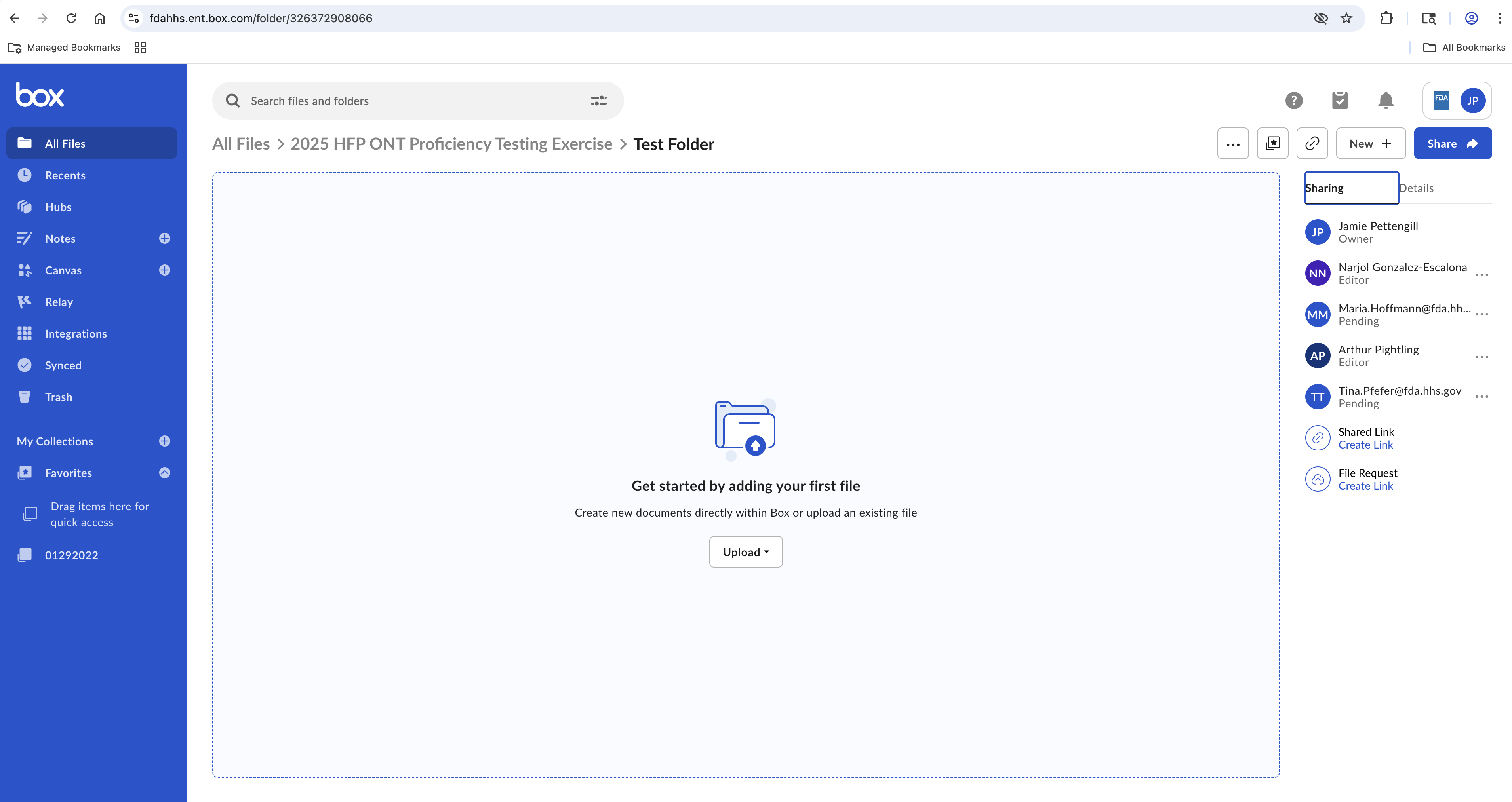
Figure 1. Example of the Box interface through which participants will upload files.
